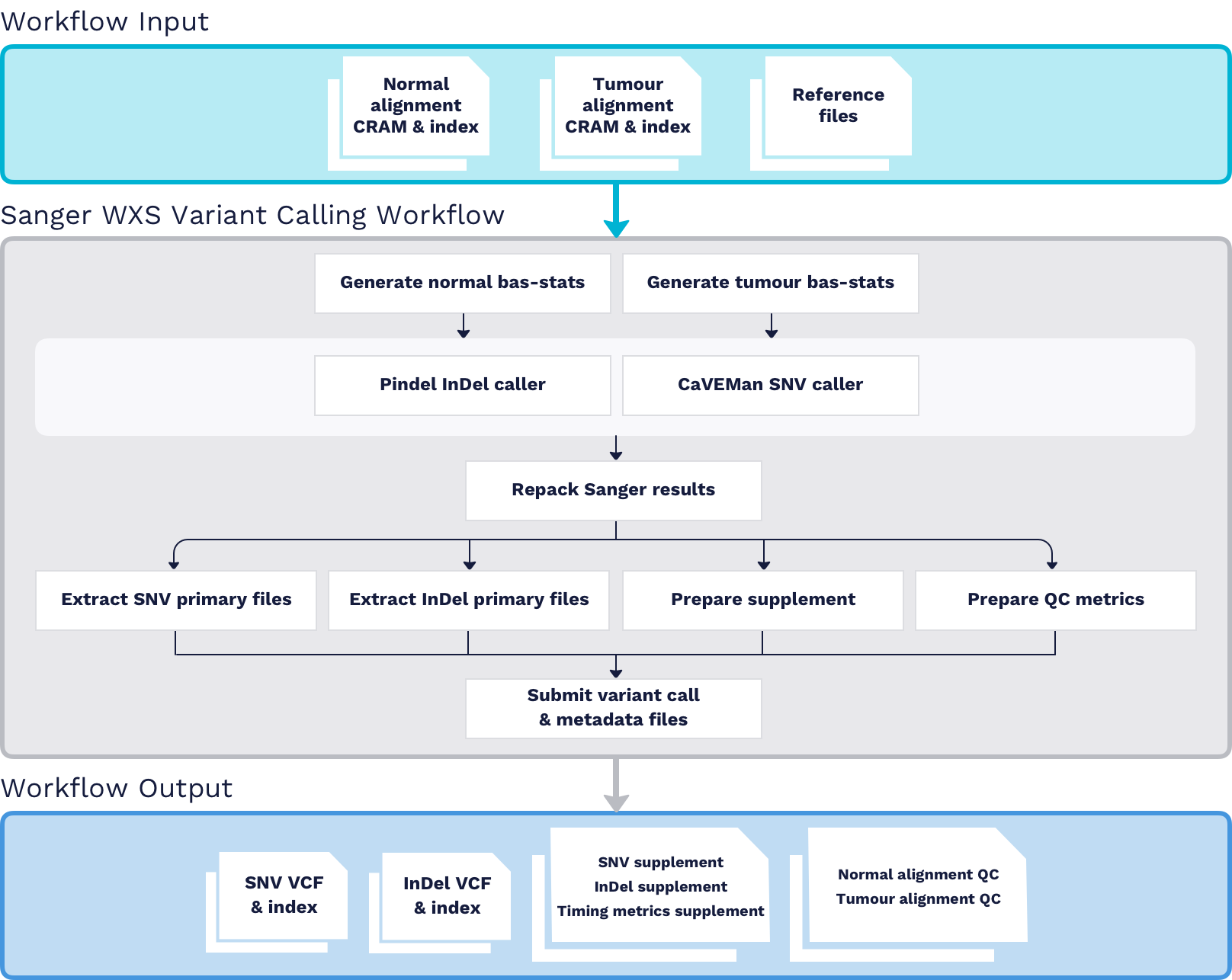
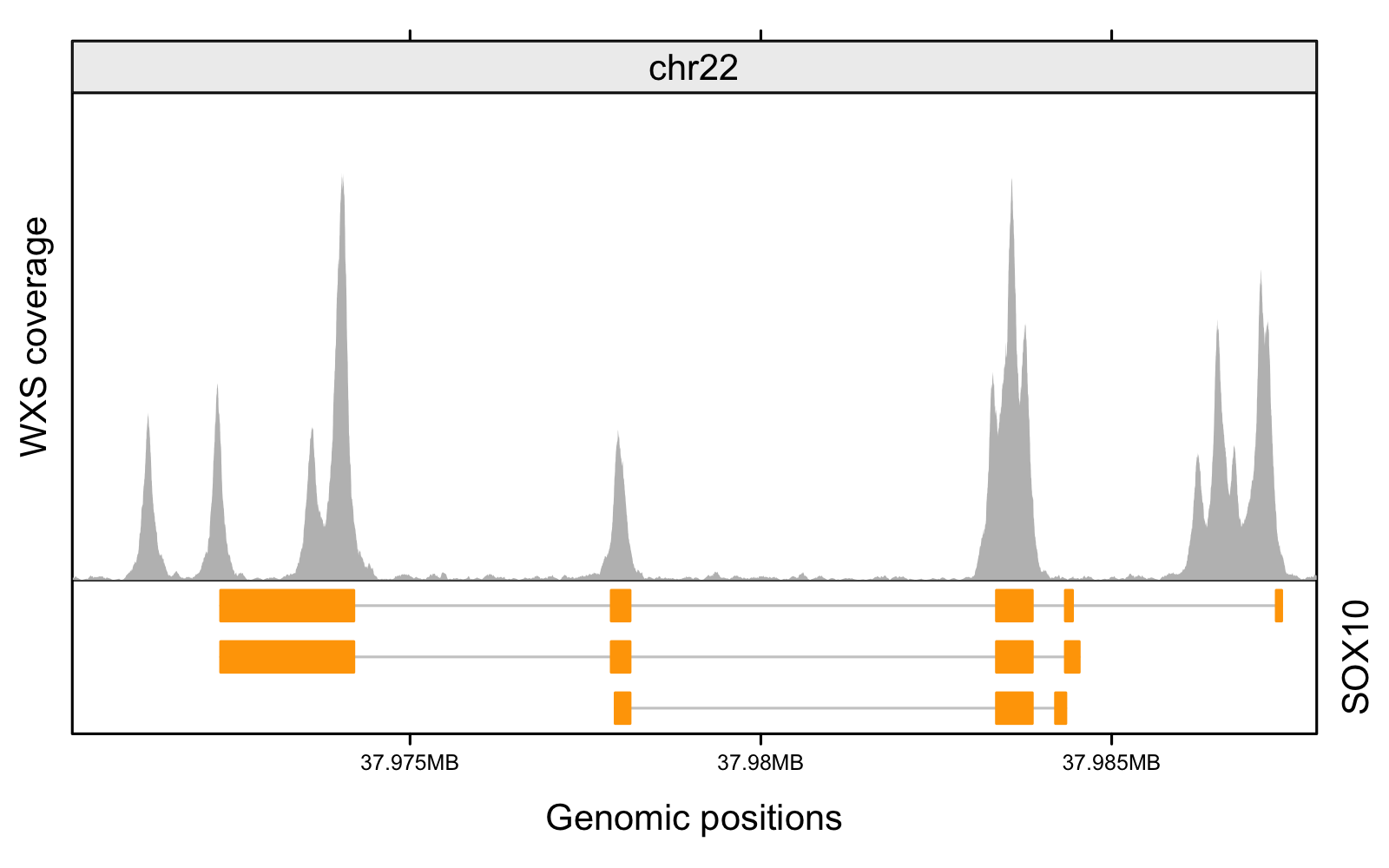
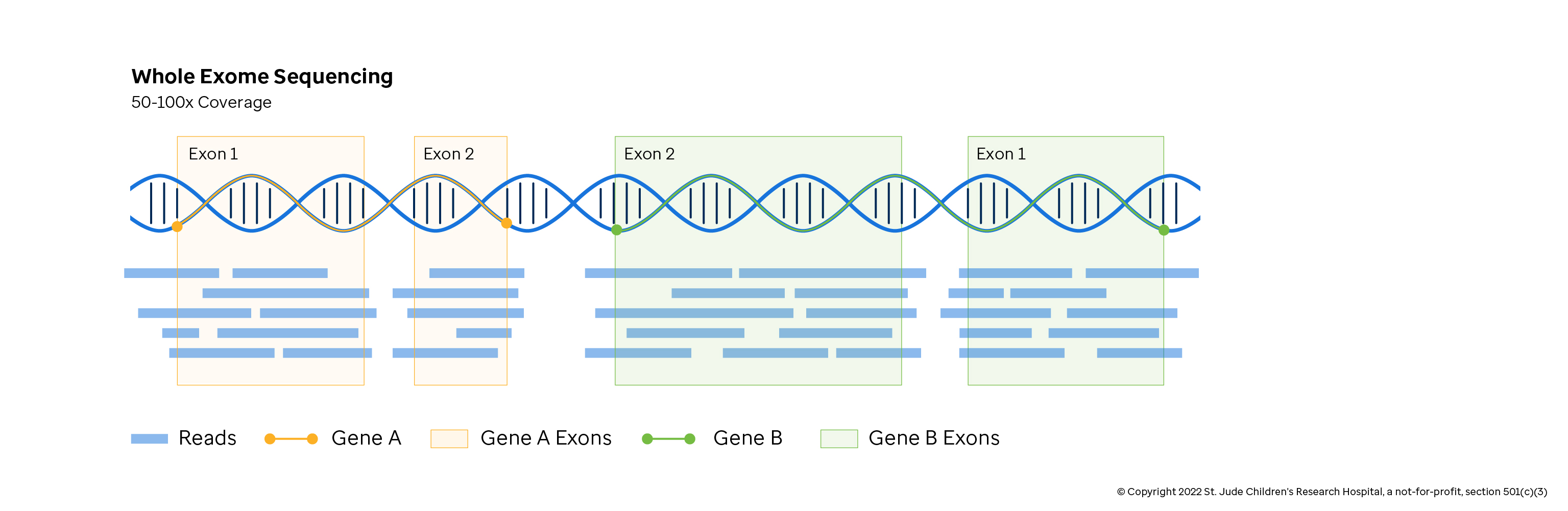
Preview on Skymap project: extracting allelic read count and expression profiles of >400,000 sequencing run into simple omic matrices – Brian Y Tsui
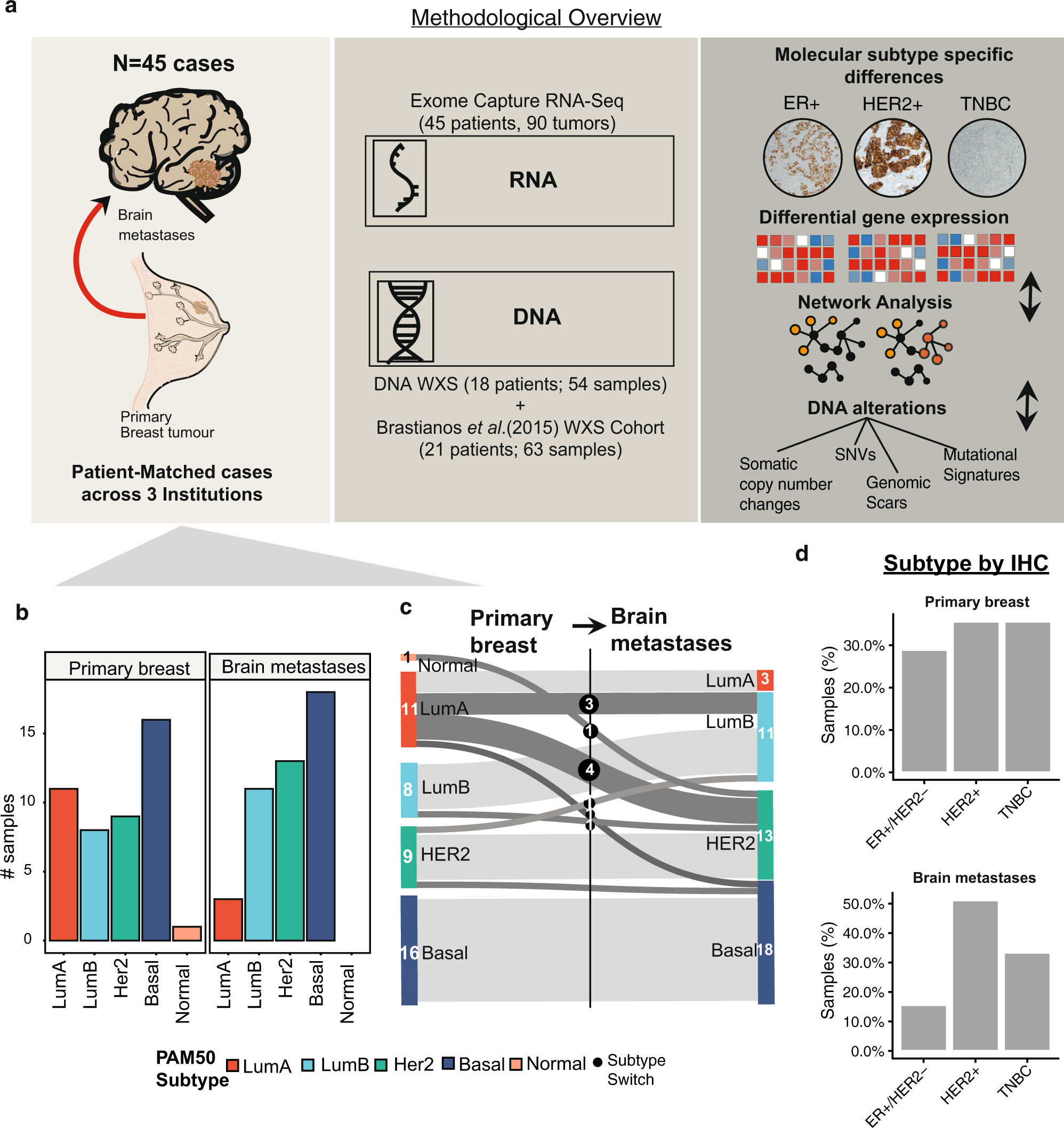
Mapping molecular subtype specific alterations in breast cancer brain metastases identifies clinically relevant vulnerabilities | Nature Communications
Extracting allelic read counts from 250,000 human sequencing runs in Sequence Read Archive 1. Introduction The reduction of sequ

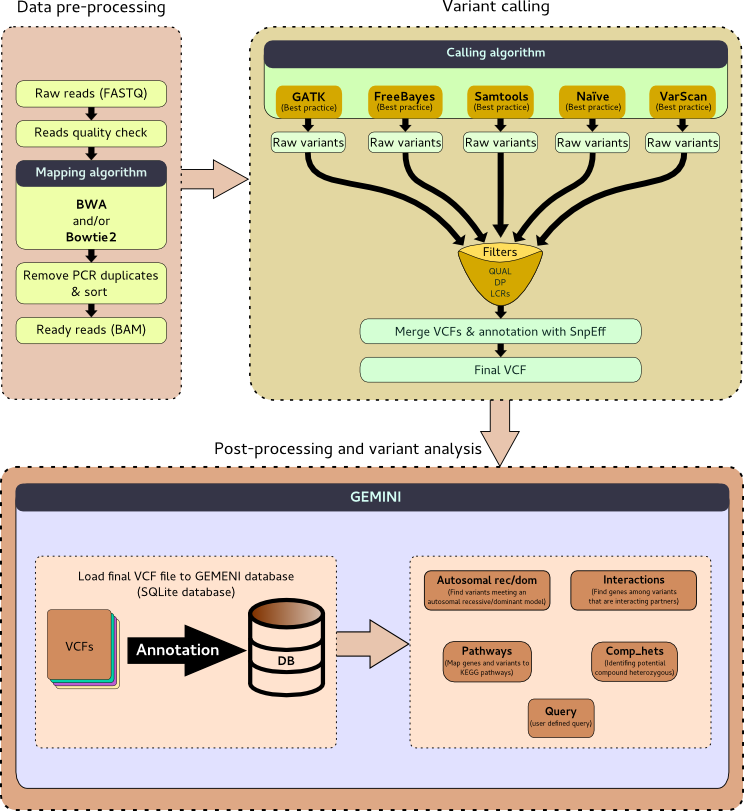
MuPeXI flow chart. Pre-processing: Raw sequencing data (WXS/WGS and... | Download Scientific Diagram
Targeted or whole genome sequencing of formalin fixed tissue samples: potential applications in cancer genomics
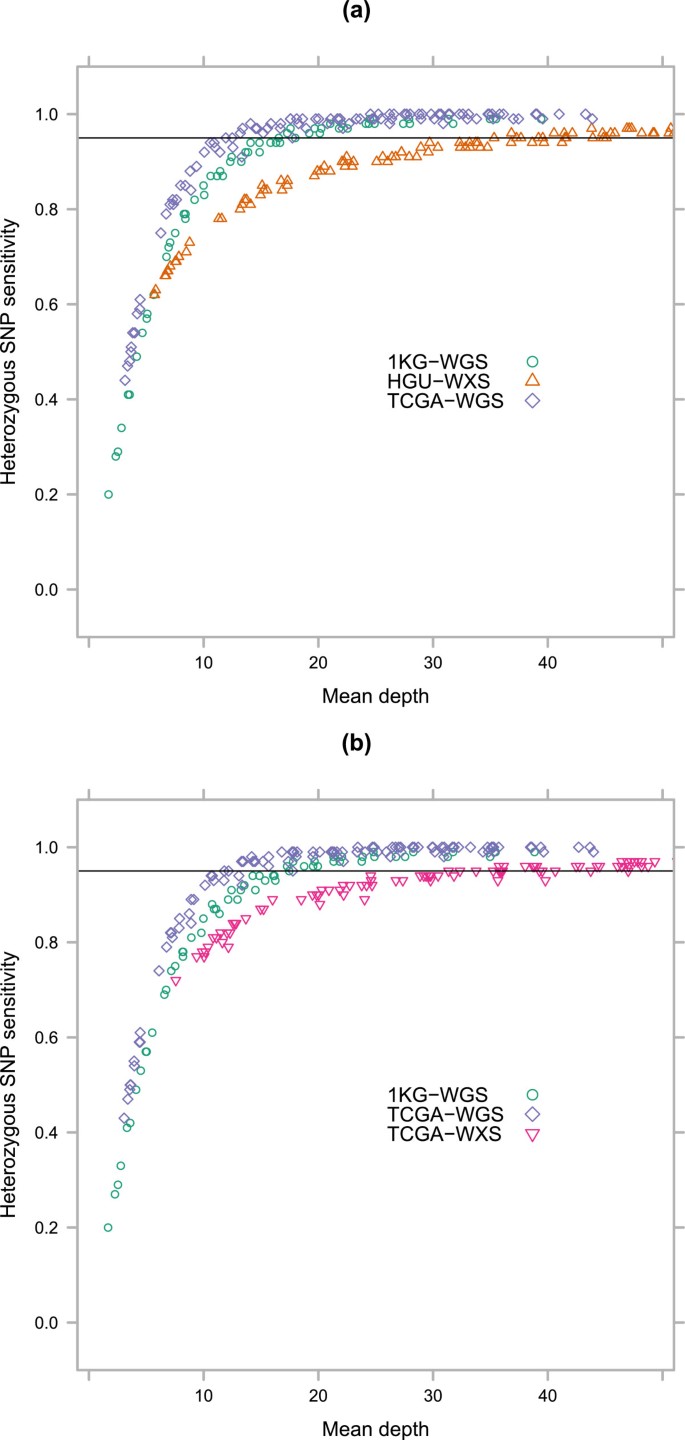
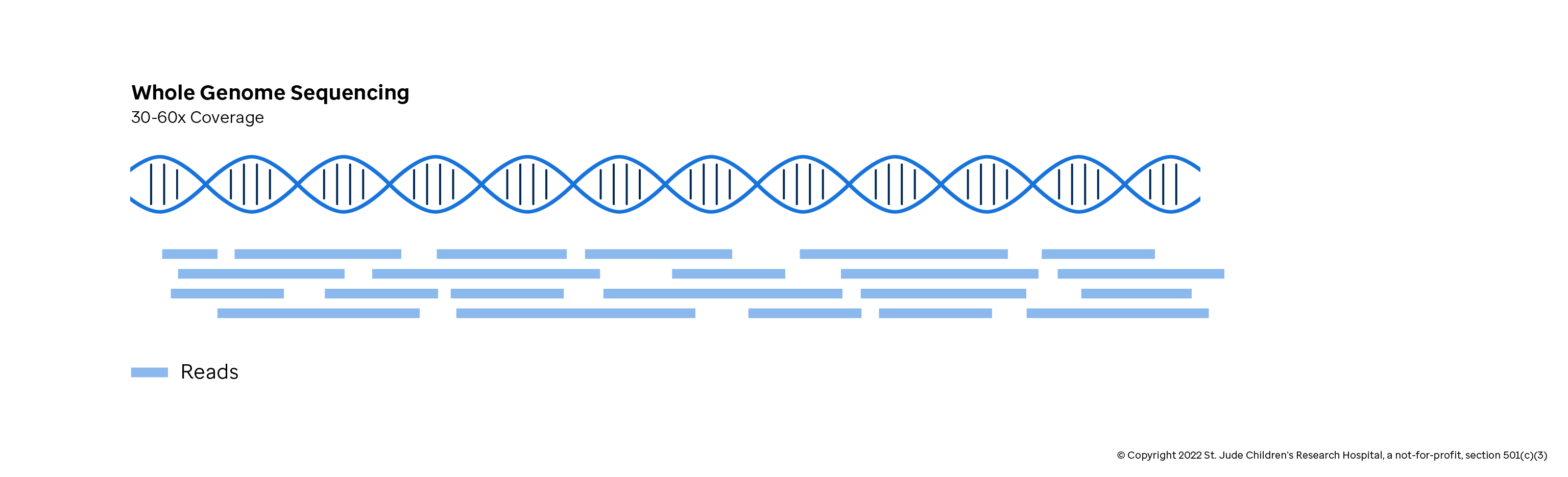
Variant detection sensitivity and biases in whole genome and exome sequencing | BMC Bioinformatics | Full Text

Analytical Validation of a Computational Method for Pharmacogenetic Genotyping from Clinical Whole Exome Sequencing - ScienceDirect
Extracting allelic read counts from 250,000 human sequencing runs in Sequence Read Archive 1. Introduction The reduction of sequ
Workflow of the search for variants through filtering and analysis. The... | Download Scientific Diagram

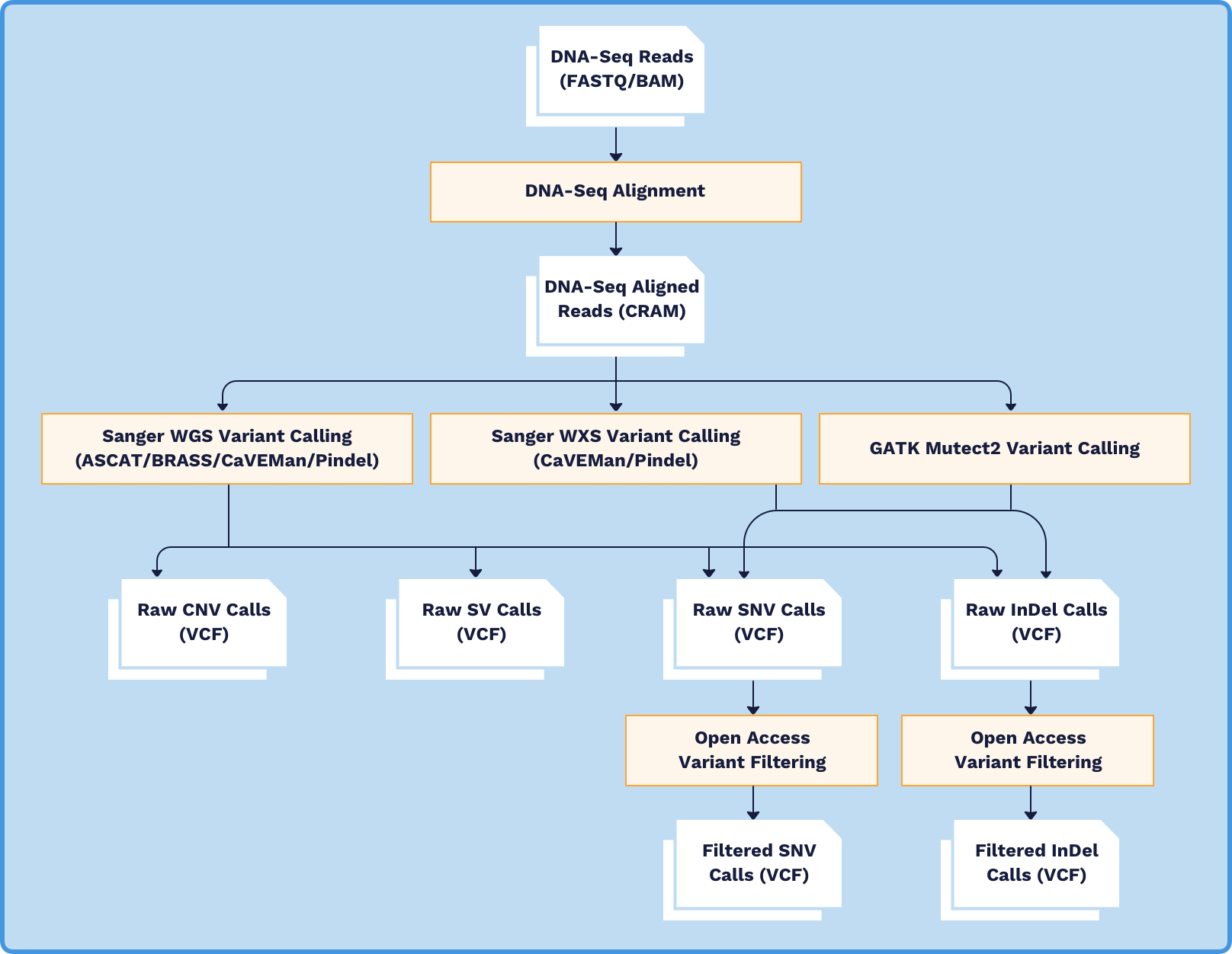
NCI GDC on Twitter: "Find WXS, RNA-seq, and targeted sequencing data from NCI's Exceptional Responders Initiative in the GDC: https://t.co/iL8c6j9Dx8 https://t.co/FDzKf32W56" / Twitter
Figure S6. Comparison of receptor-derived reads provided by WXS and... | Download Scientific Diagram