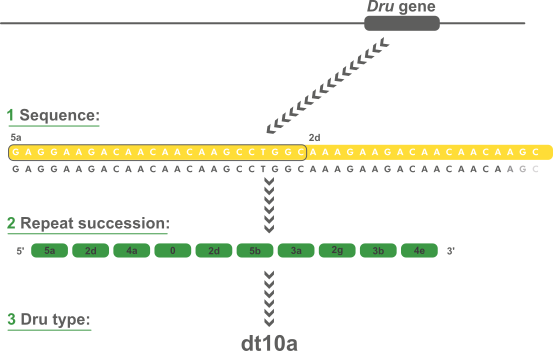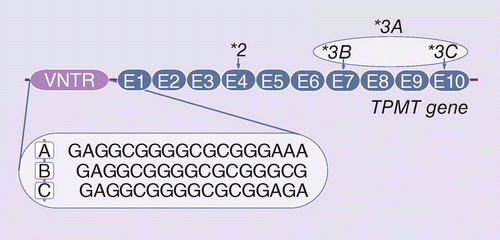
Novel motif of variable number of tandem repeats in TPMT promoter region and evolutionary association of variable number of tandem repeats with TPMT*3 alleles | Pharmacogenomics

Identification of Variable-Number Tandem-Repeat (VNTR) Sequences in Acinetobacter baumannii and Interlaboratory Validation of an Optimized Multiple-Locus VNTR Analysis Typing Scheme | Journal of Clinical Microbiology

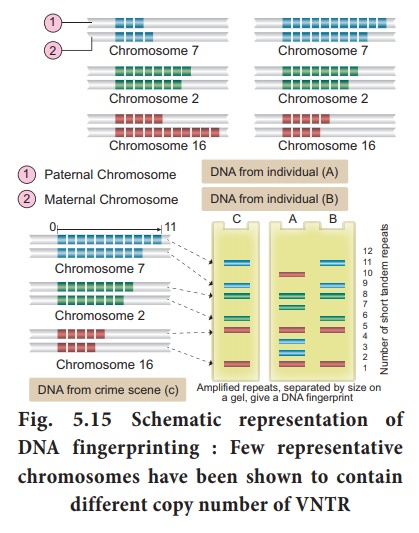
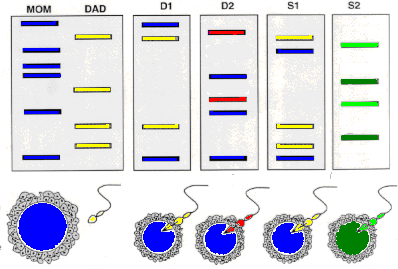
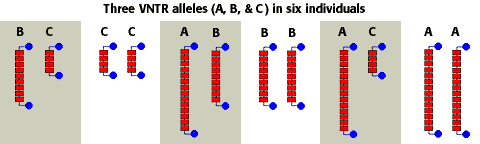
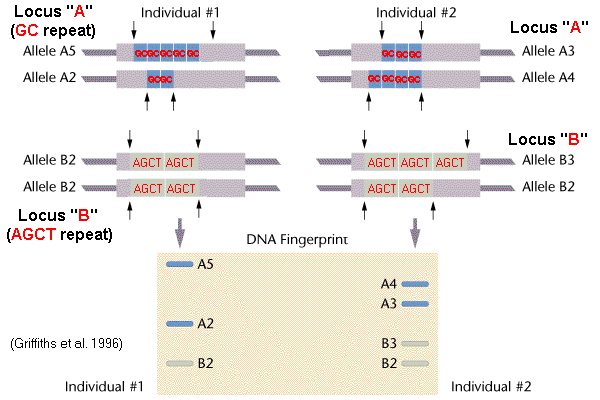
molecular genetics - Use of restriction endonucleases to analyse Variable Number Tandem Repeats (VNTRs) - Biology Stack Exchange
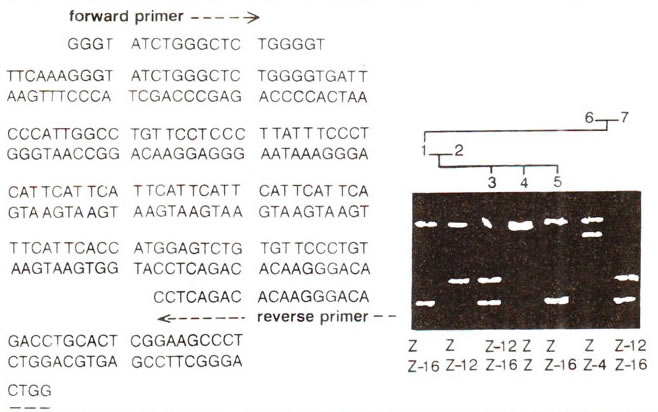
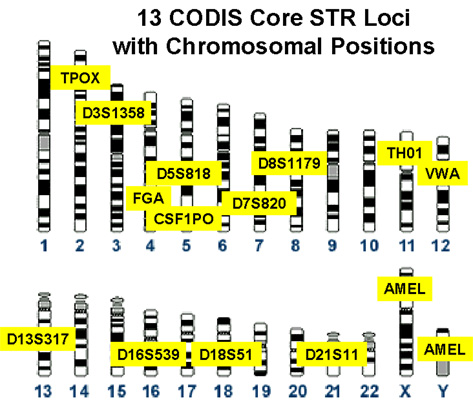
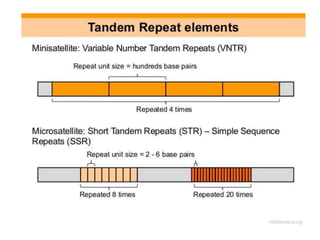
Minisatellites (VNTRs) and Microsatellites (SSRs) | Genetic Engineering and Biotechnology Restriction Maps and Molecular Genetic Maps

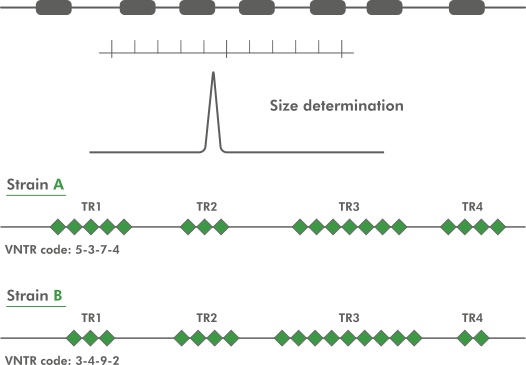
Variable number tandem repeats – Their emerging role in sickness and health - Jack NG Marshall, Ana Illera Lopez, Abigail L Pfaff, Sulev Koks, John P Quinn, Vivien J Bubb, 2021
![MIRU-profiler: a rapid tool for determination of 24-loci MIRU-VNTR profiles from assembled genomes of Mycobacterium tuberculosis [PeerJ] MIRU-profiler: a rapid tool for determination of 24-loci MIRU-VNTR profiles from assembled genomes of Mycobacterium tuberculosis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2018/5090/1/fig-1-full.png)
MIRU-profiler: a rapid tool for determination of 24-loci MIRU-VNTR profiles from assembled genomes of Mycobacterium tuberculosis [PeerJ]
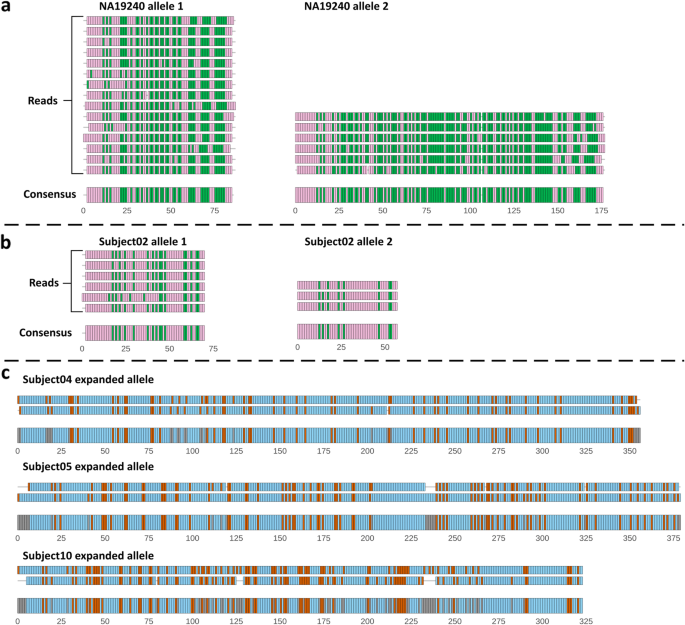
NanoSatellite: accurate characterization of expanded tandem repeat length and sequence through whole genome long-read sequencing on PromethION | Genome Biology | Full Text

YB-1 and CTCF Differentially Regulate the 5-HTT Polymorphic Intron 2 Enhancer Which Predisposes to a Variety of Neurological Disorders | Journal of Neuroscience

VNTR prediction on sequence characteristics using long-read annotation and validation by short-read pileup | bioRxiv