UMI-Gen: a UMI-based reads simulator for variant calling evaluation in paired-end sequencing NGS libraries | bioRxiv
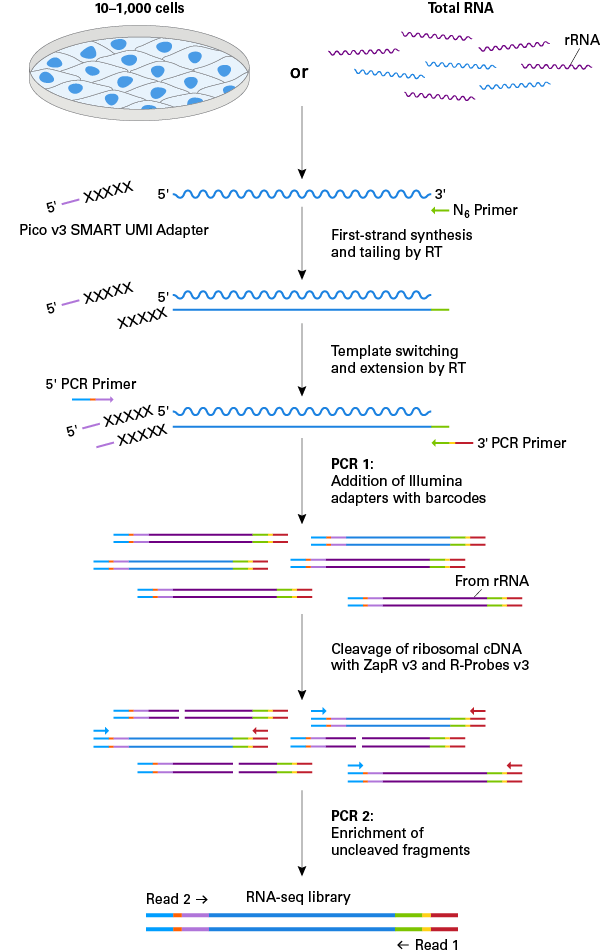
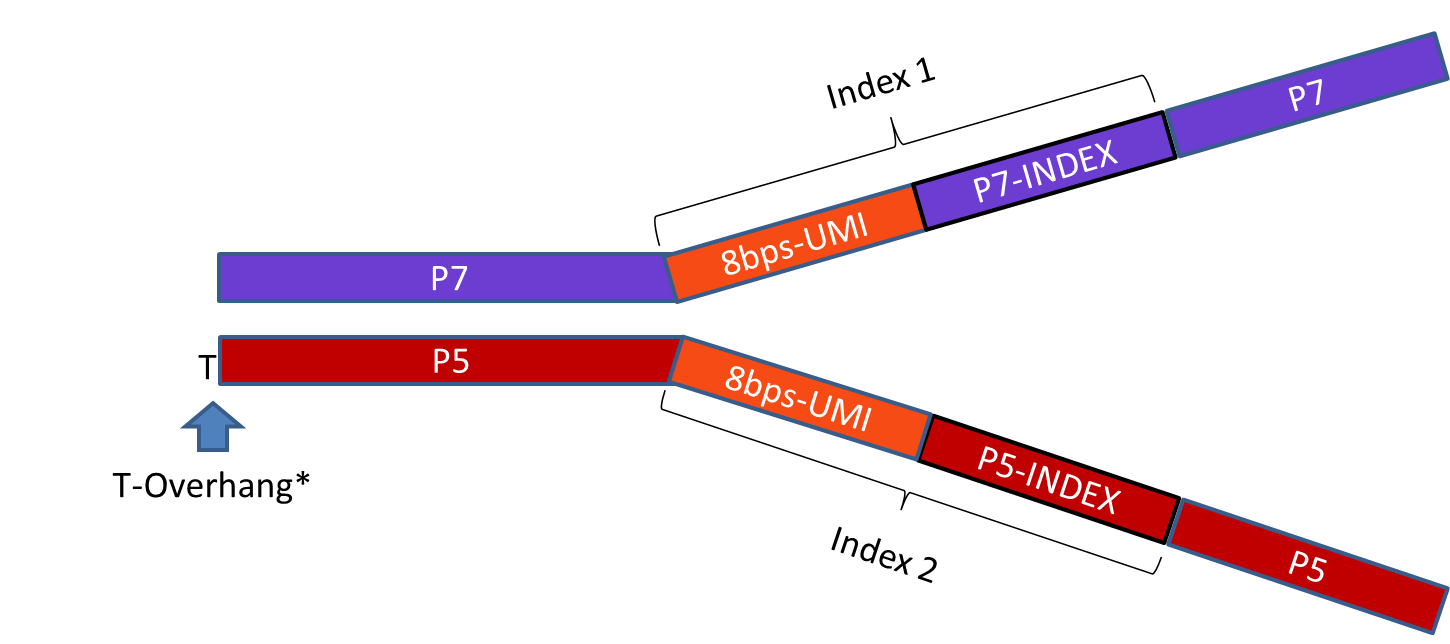
Generate UMI-labeled, stranded libraries for whole-transcriptome analyses, compatible with Illumina NGS platforms.

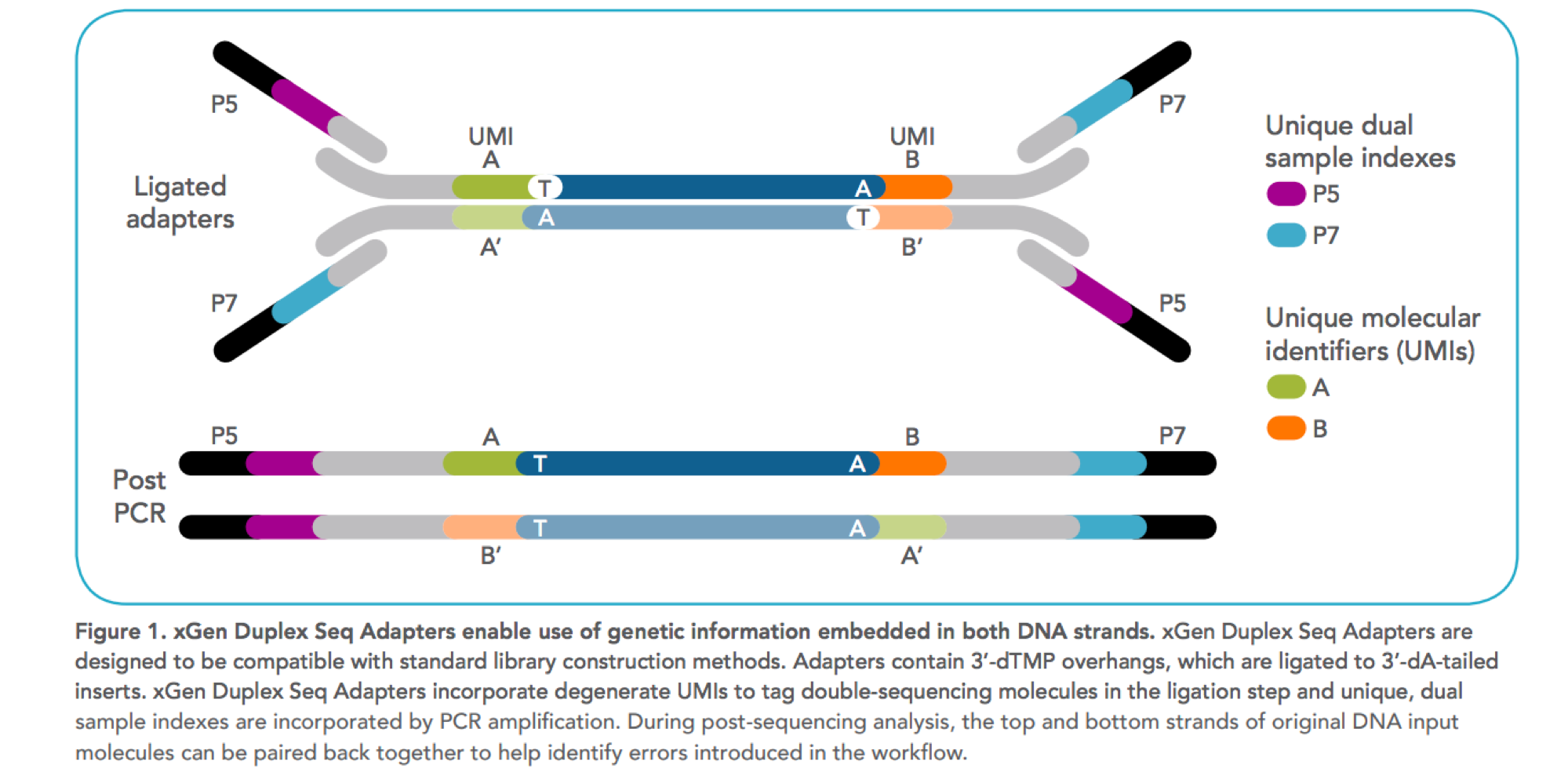
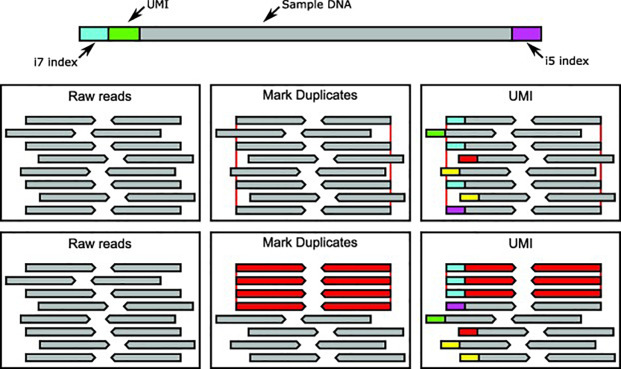
Incorporation of unique molecular identifiers (UMIs) in TruSeq adapters improves the accuracy of quantitative sequencing | RNA-Seq Blog

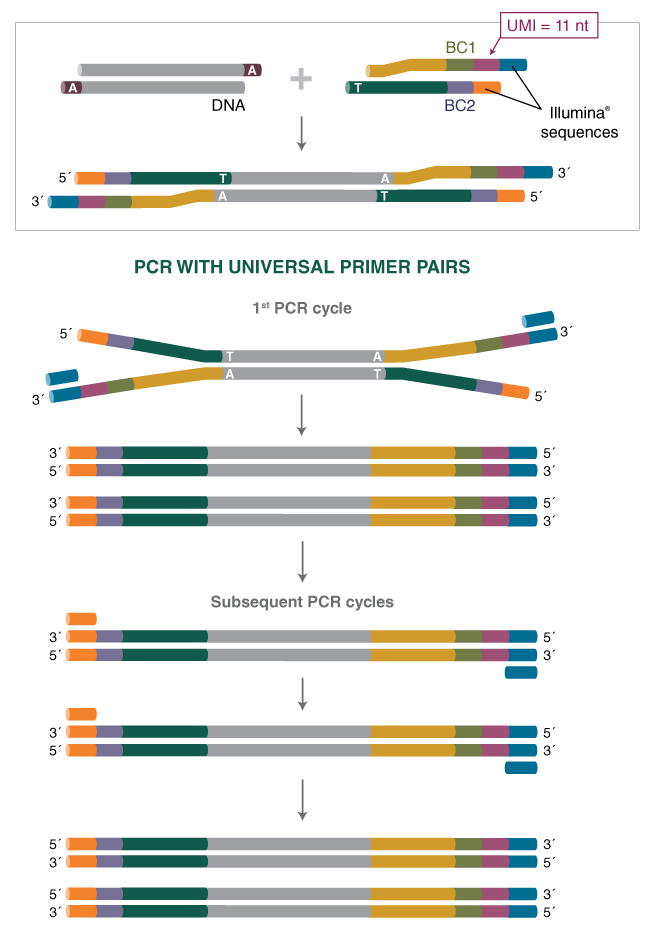
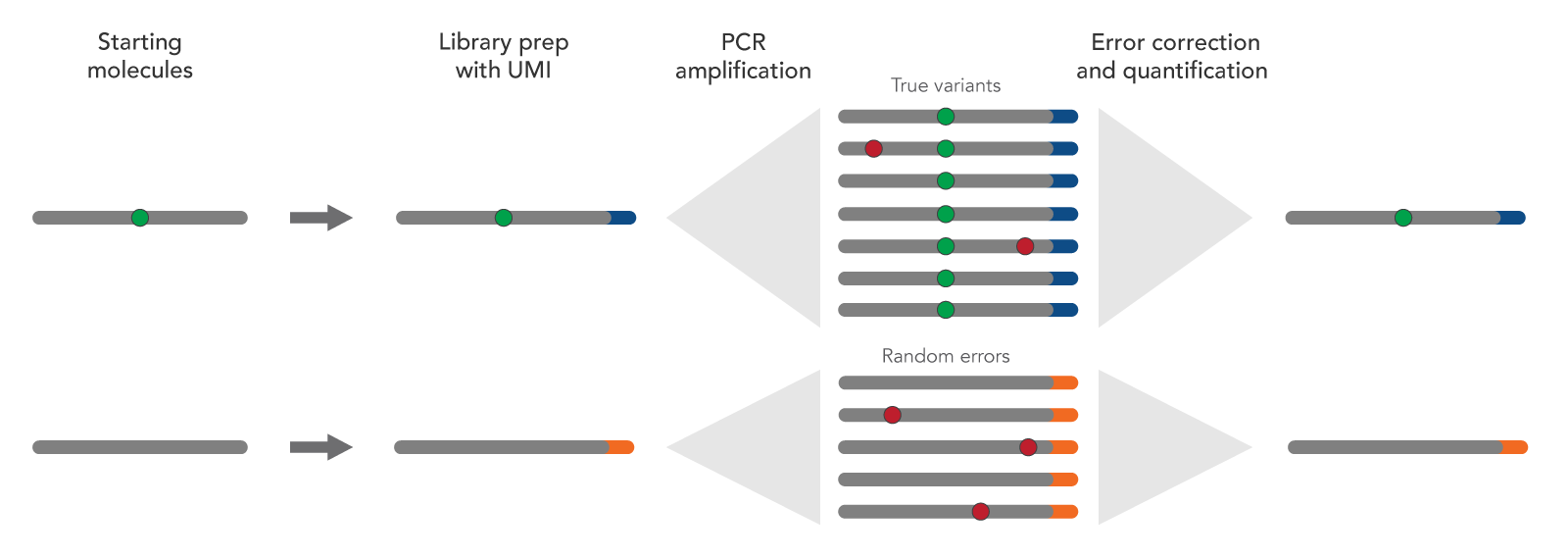
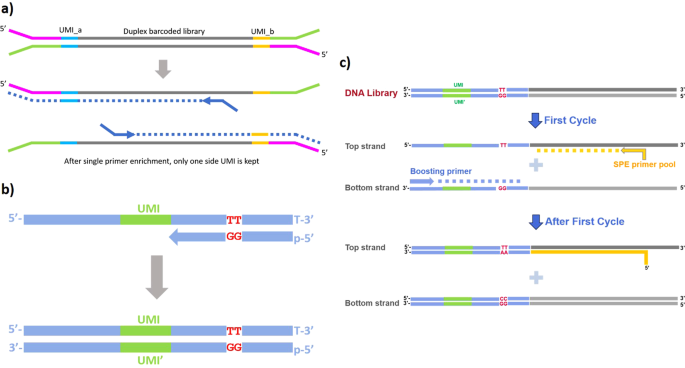
MuA-based Molecular Indexing for Rare Mutation Detection by Next-Generation Sequencing - ScienceDirect

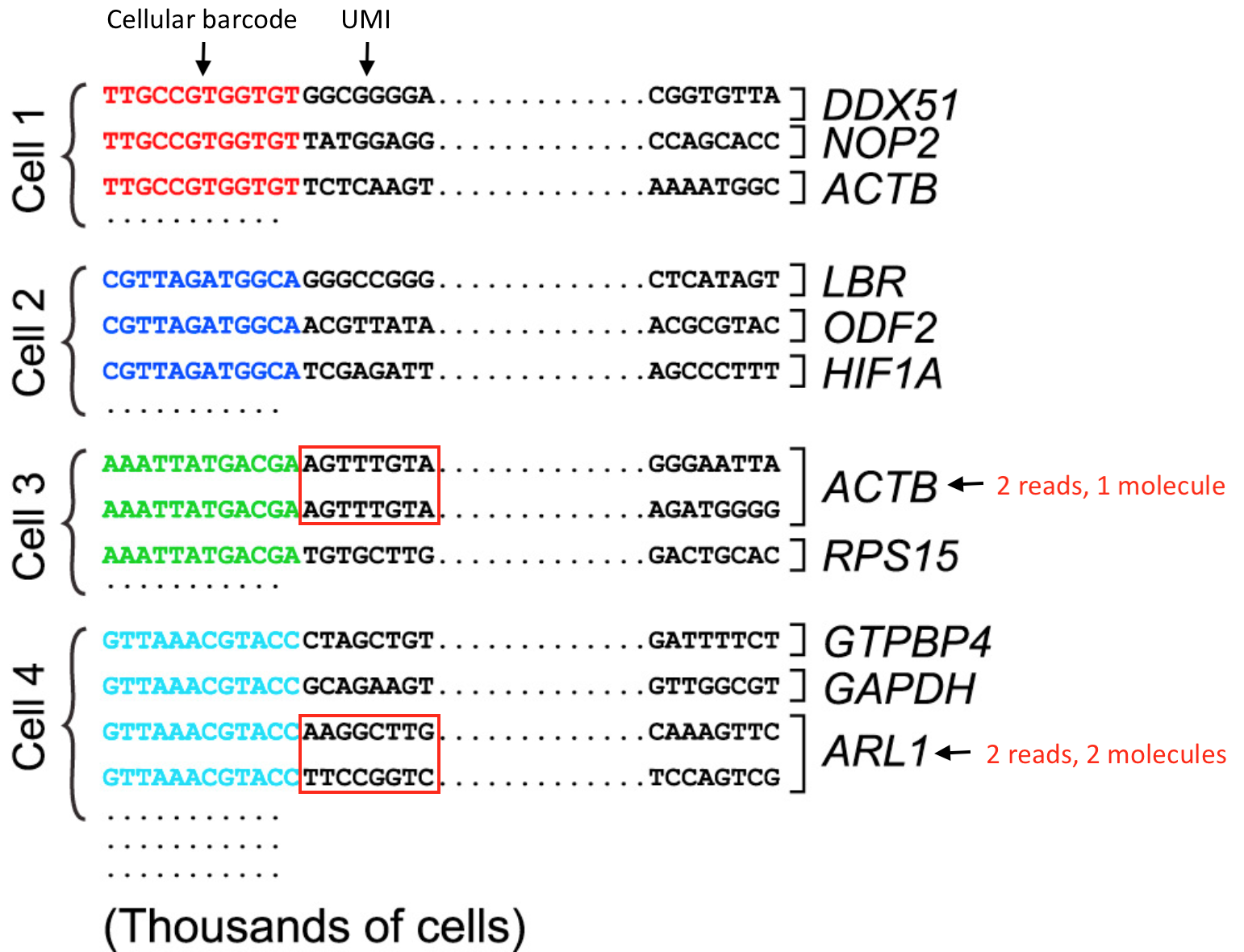
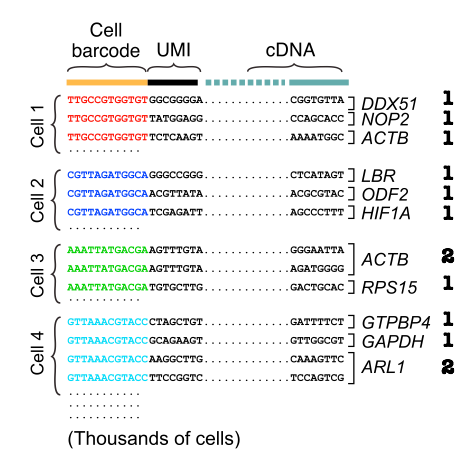
UMI incorporation into RNA-seq. a Overall workflow. Schematic of a read... | Download Scientific Diagram
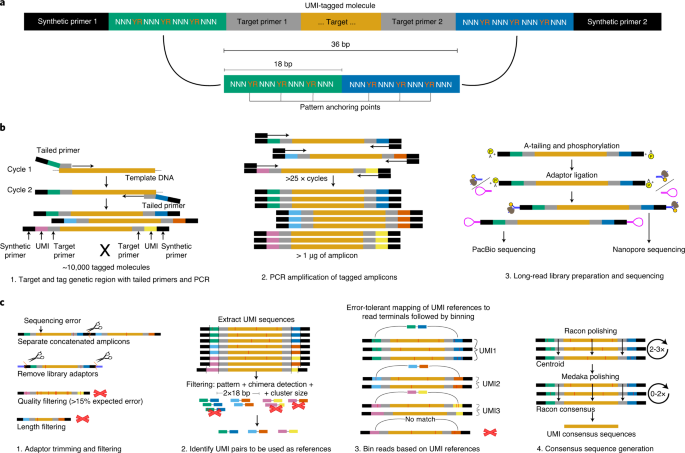
High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods

IsoSeek – unbiased and UMI-informed sequencing of cell-free miRNAs at single-nucleotide resolution | RNA-Seq Blog
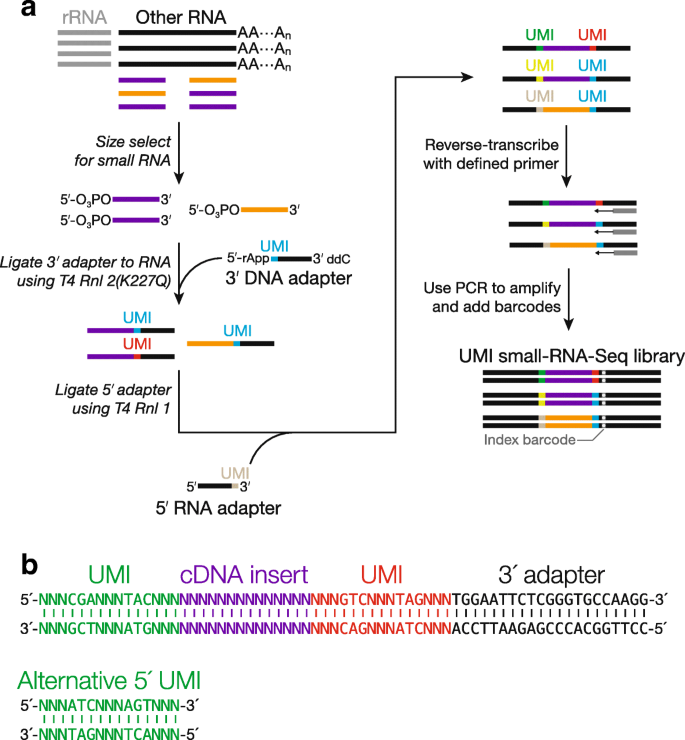
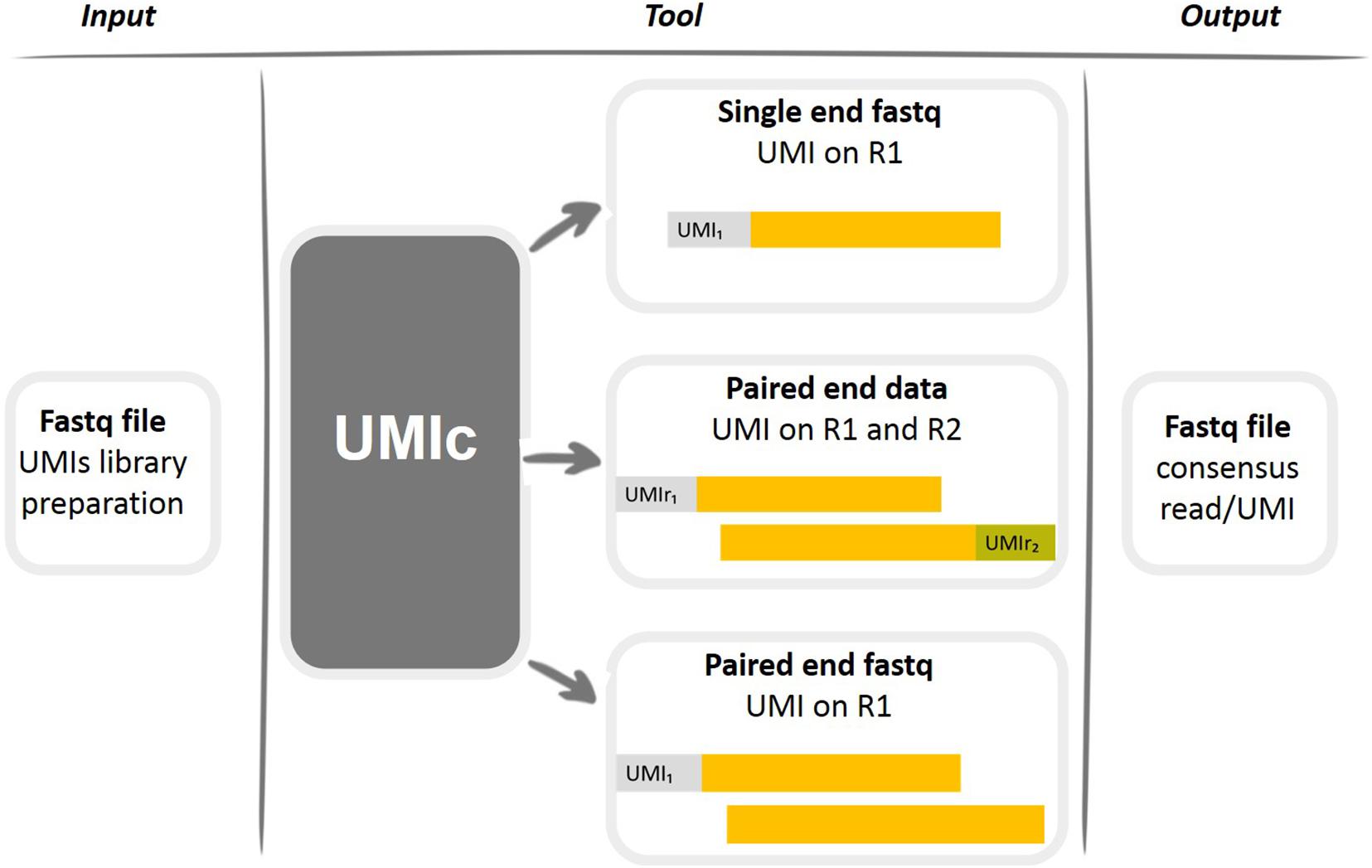
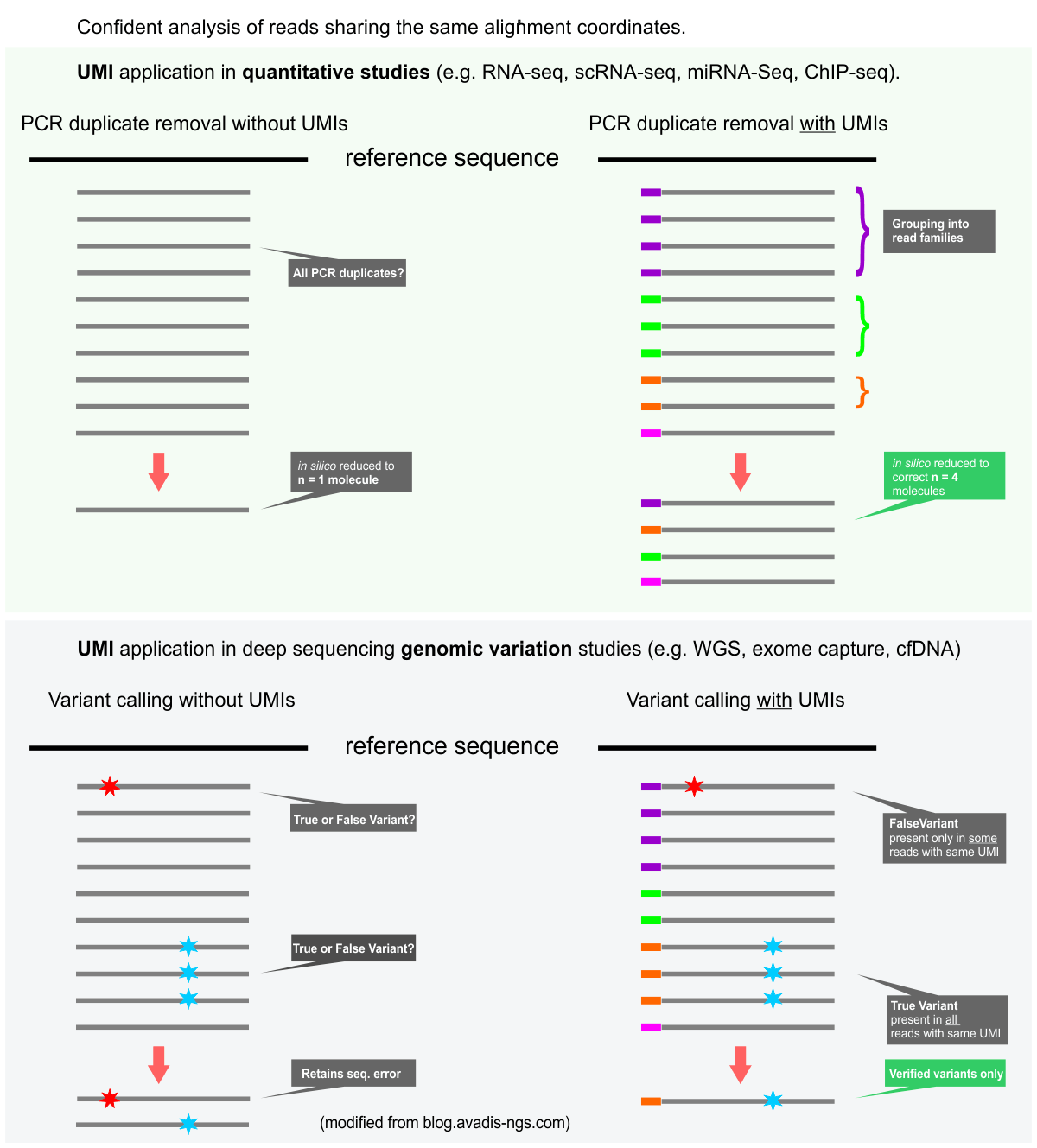
Elimination of PCR duplicates in RNA-seq and small RNA-seq using unique molecular identifiers | BMC Genomics | Full Text