Mads Albertsen on Twitter: "Great to see our paper on high quality (Q40++) long-read amplicons on @nanopore or @PacBio using UMI's published. Started as a blogpost and spearheaded by @SorenKarst and @RyanZiels.

IsoSeek – unbiased and UMI-informed sequencing of cell-free miRNAs at single-nucleotide resolution | RNA-Seq Blog

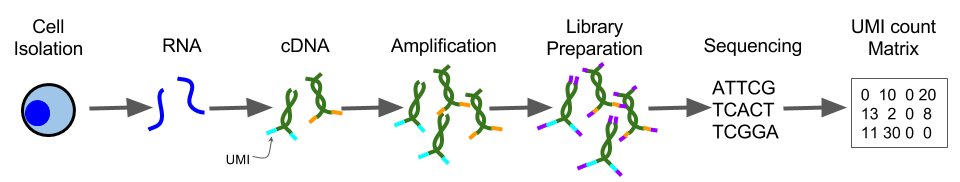
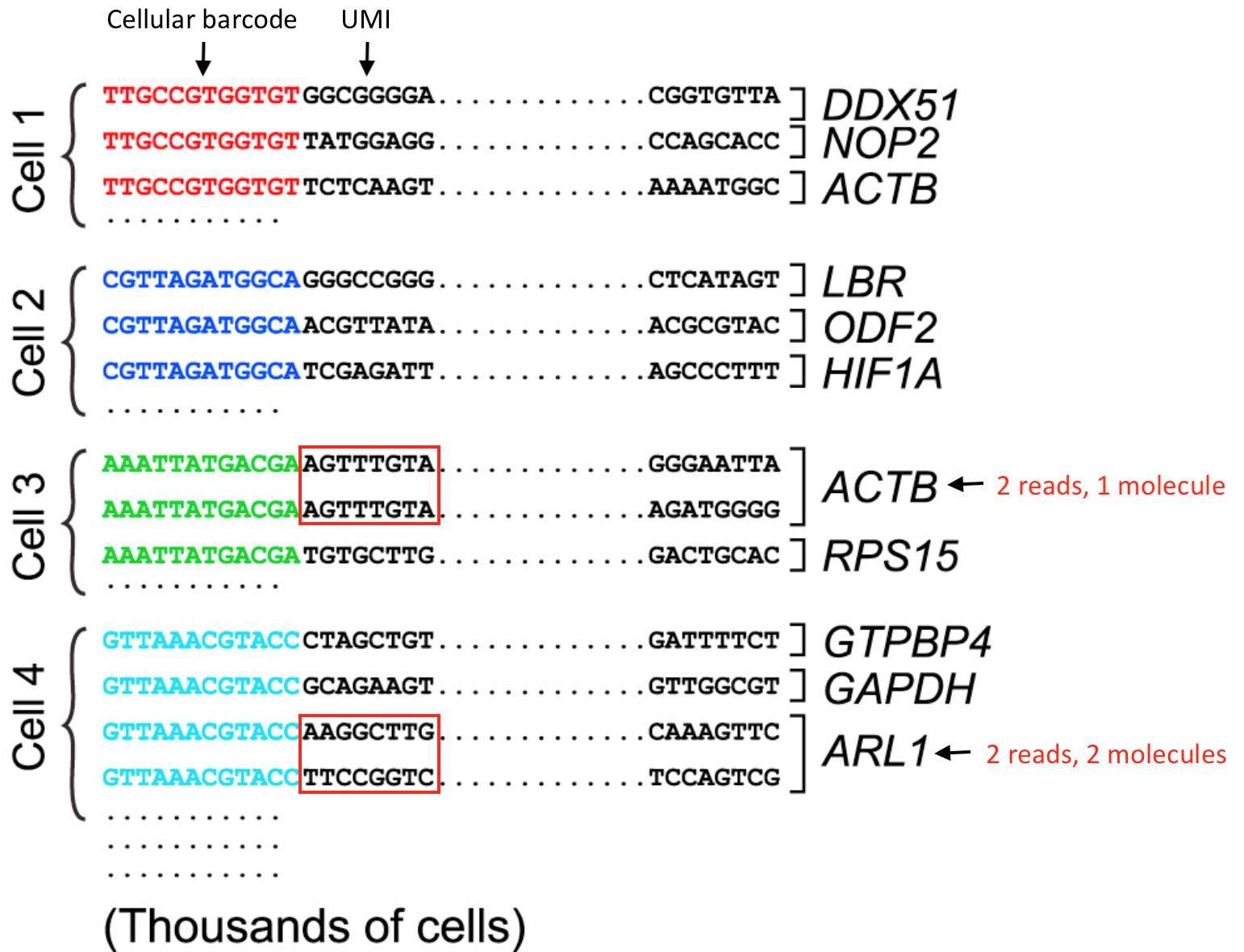
UMI incorporation into RNA-seq. a Overall workflow. Schematic of a read... | Download Scientific Diagram
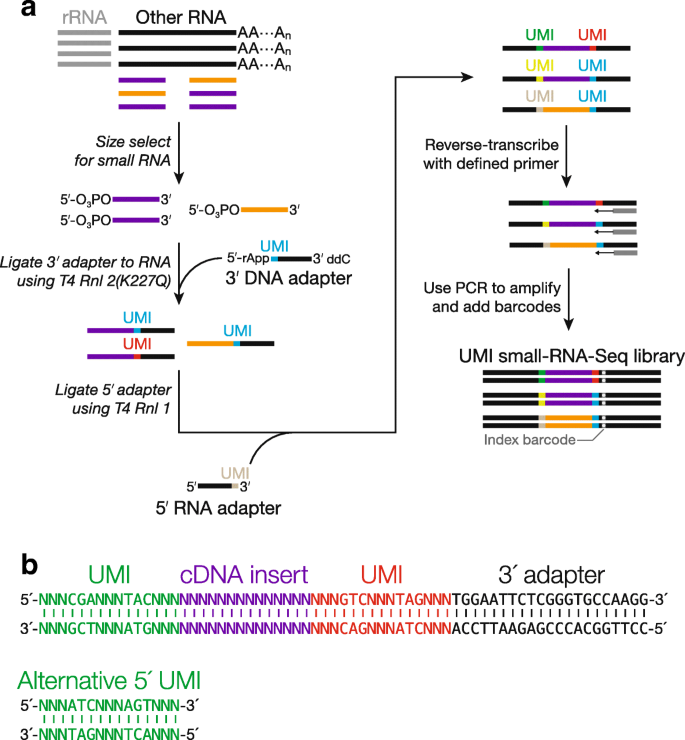
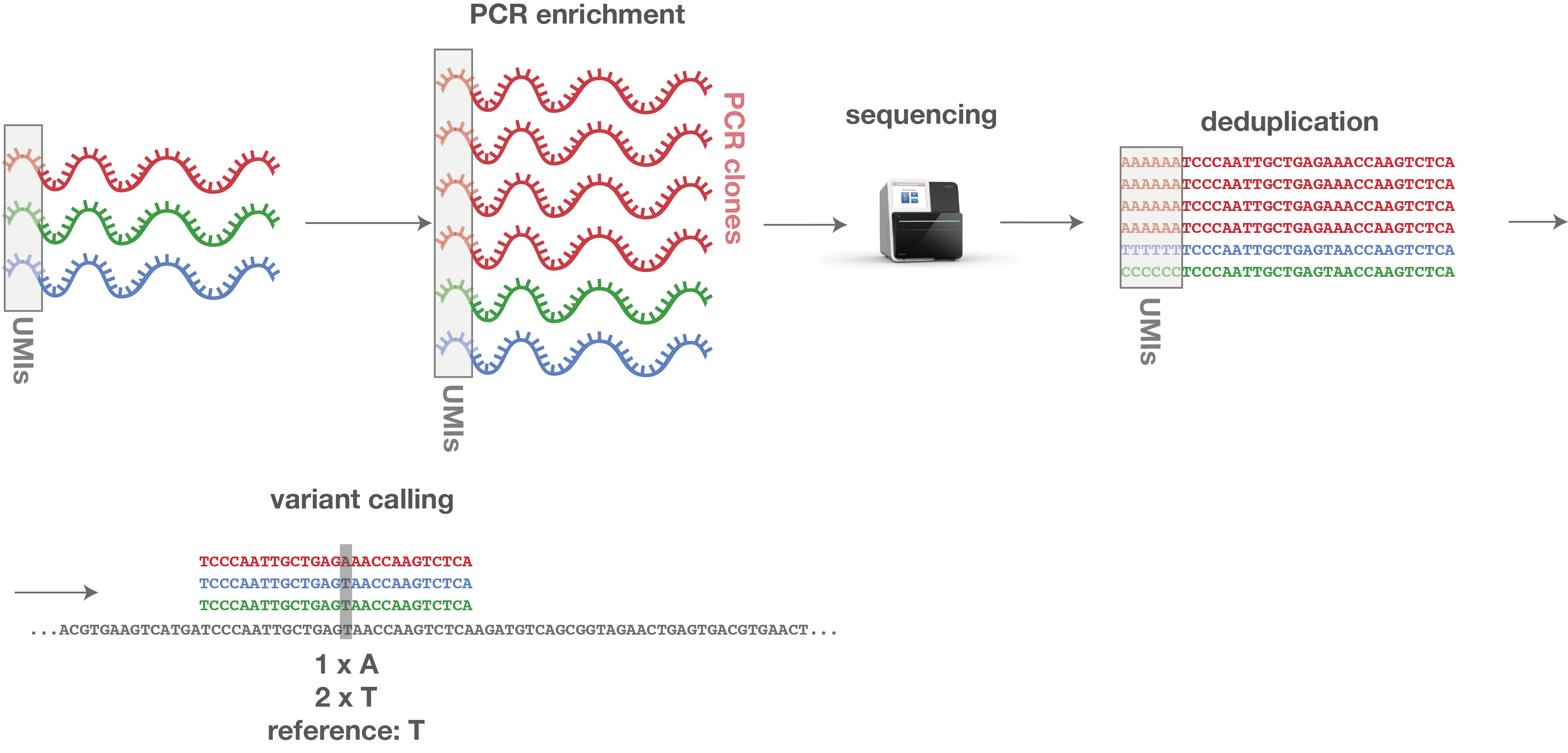
Elimination of PCR duplicates in RNA-seq and small RNA-seq using unique molecular identifiers | BMC Genomics | Full Text
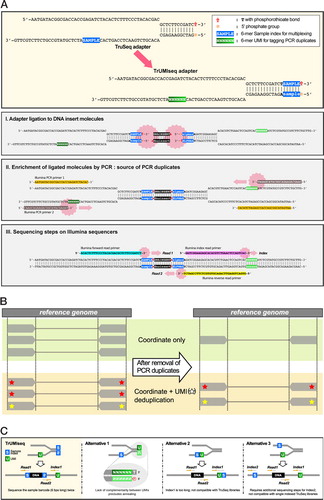
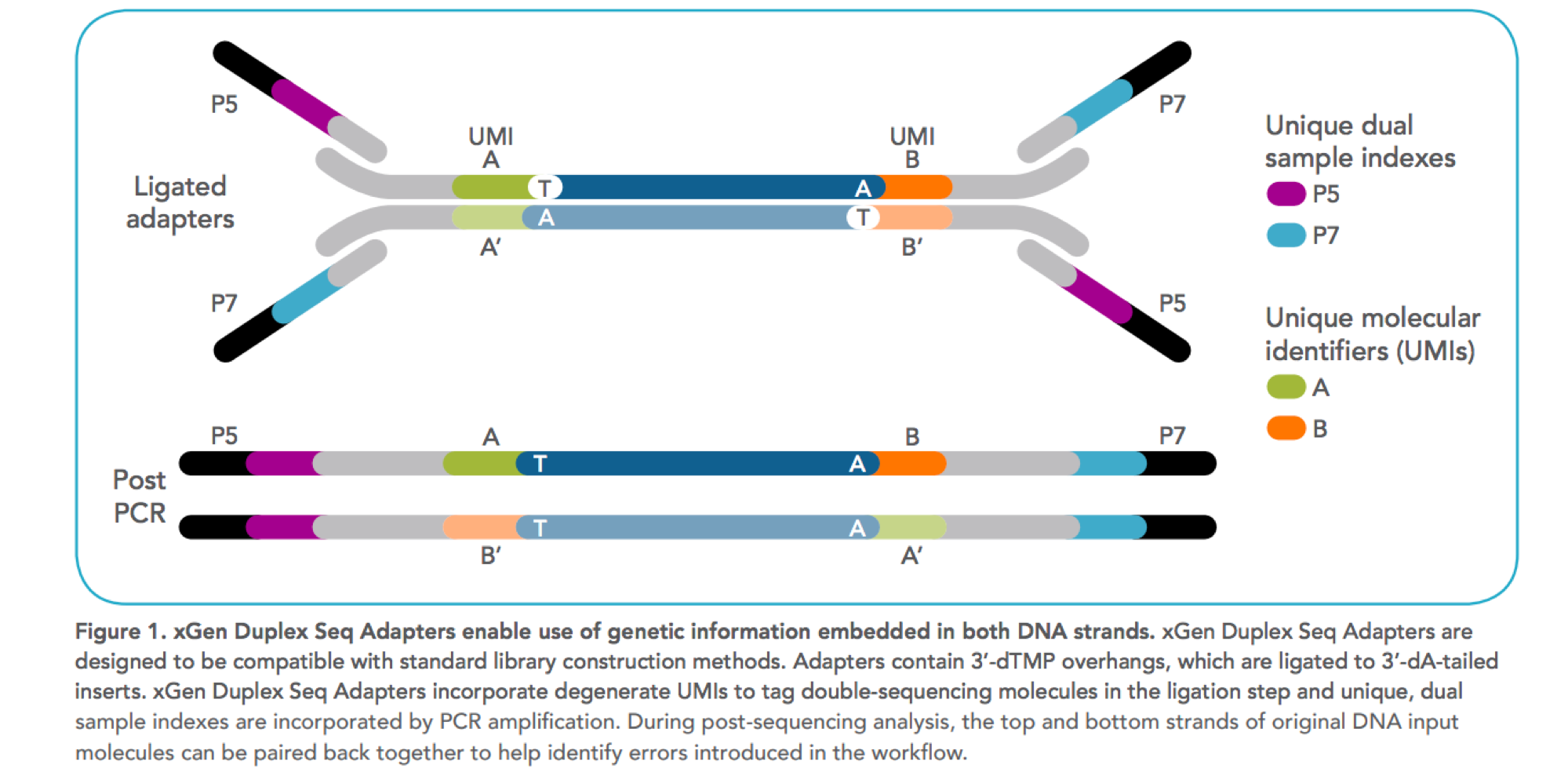
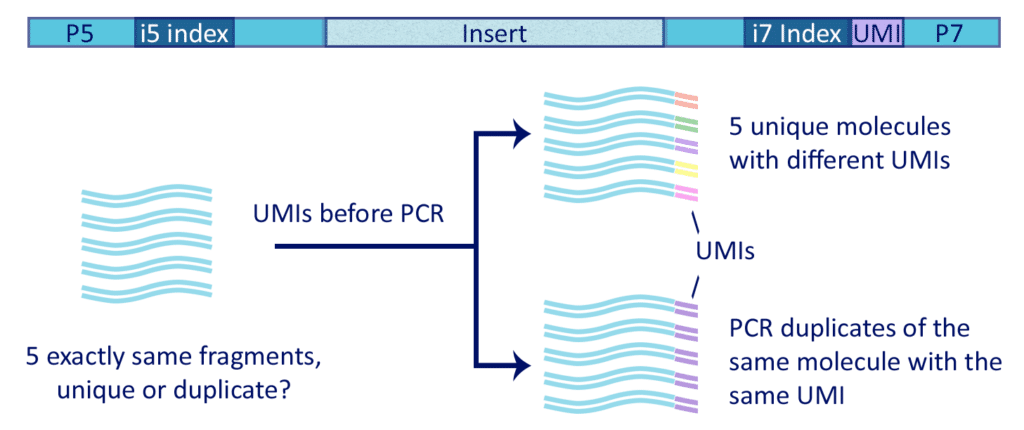
Incorporation of unique molecular identifiers in TruSeq adapters improves the accuracy of quantitative sequencing | BioTechniques

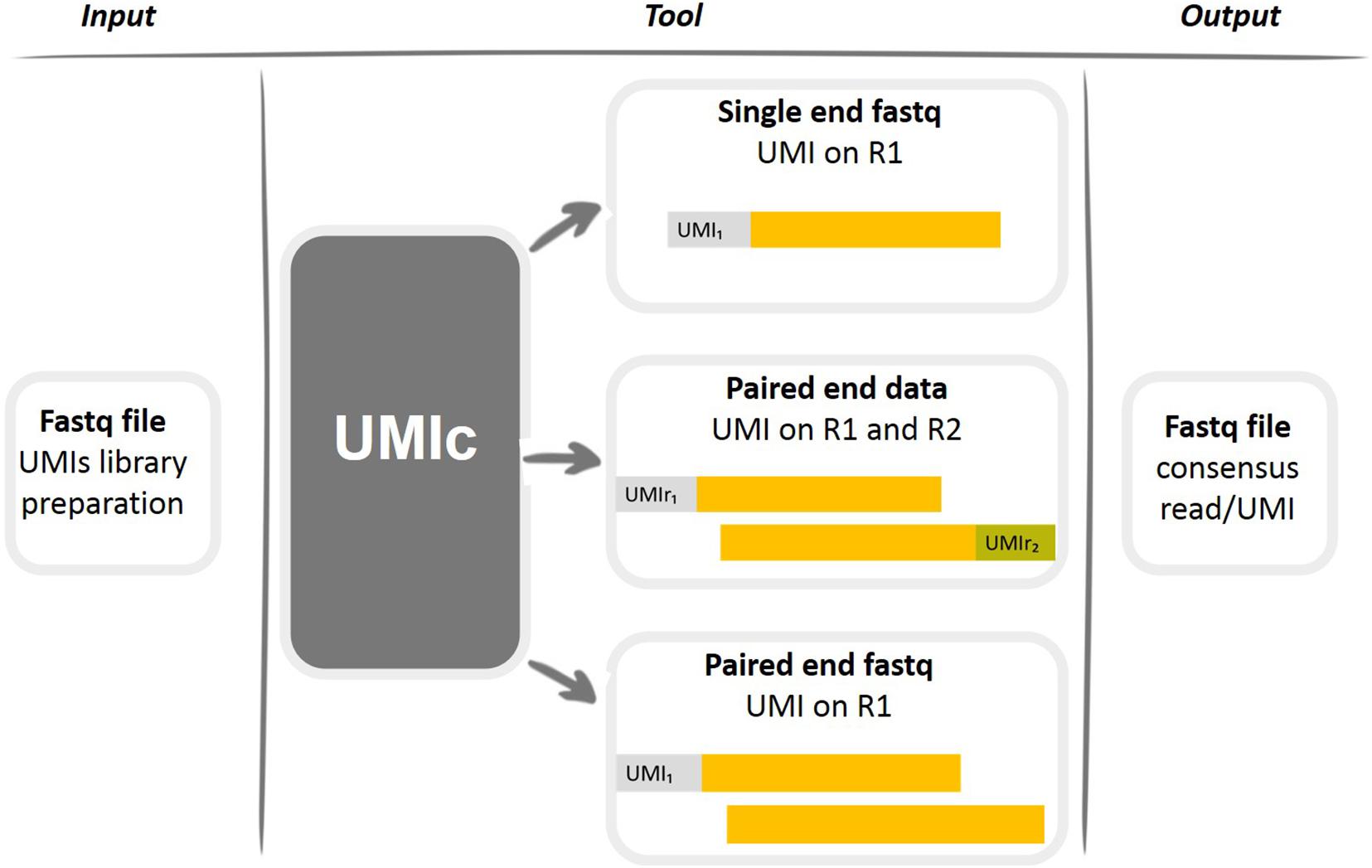
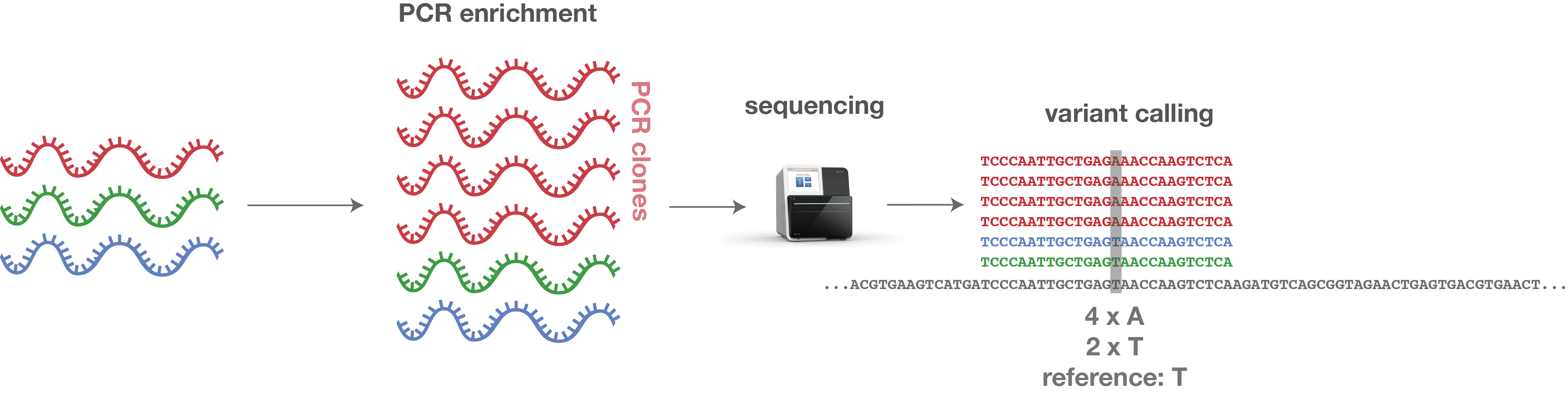
UMI-Gen: A UMI-based read simulator for variant calling evaluation in paired-end sequencing NGS libraries - ScienceDirect
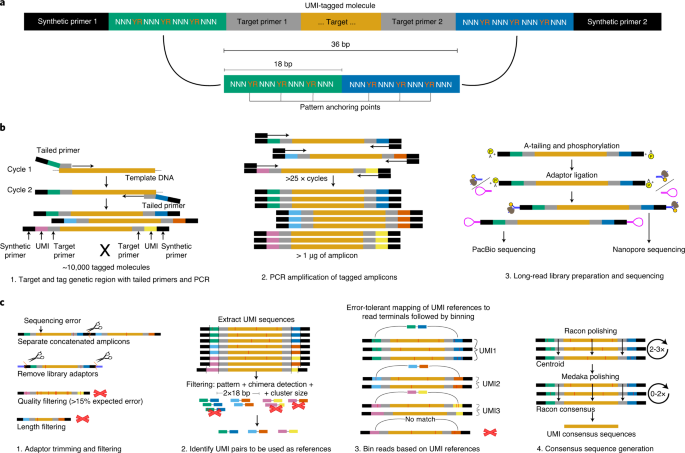
High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing | Nature Methods