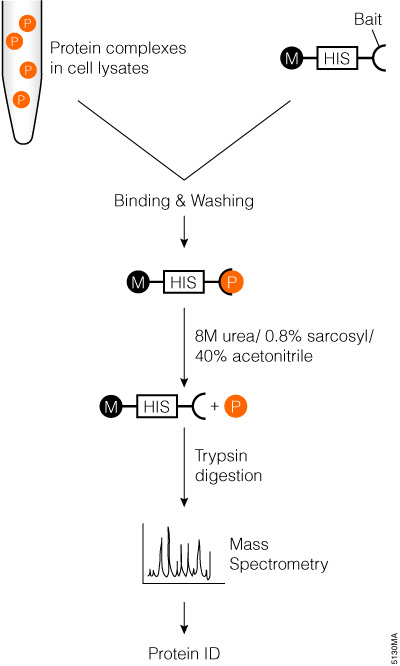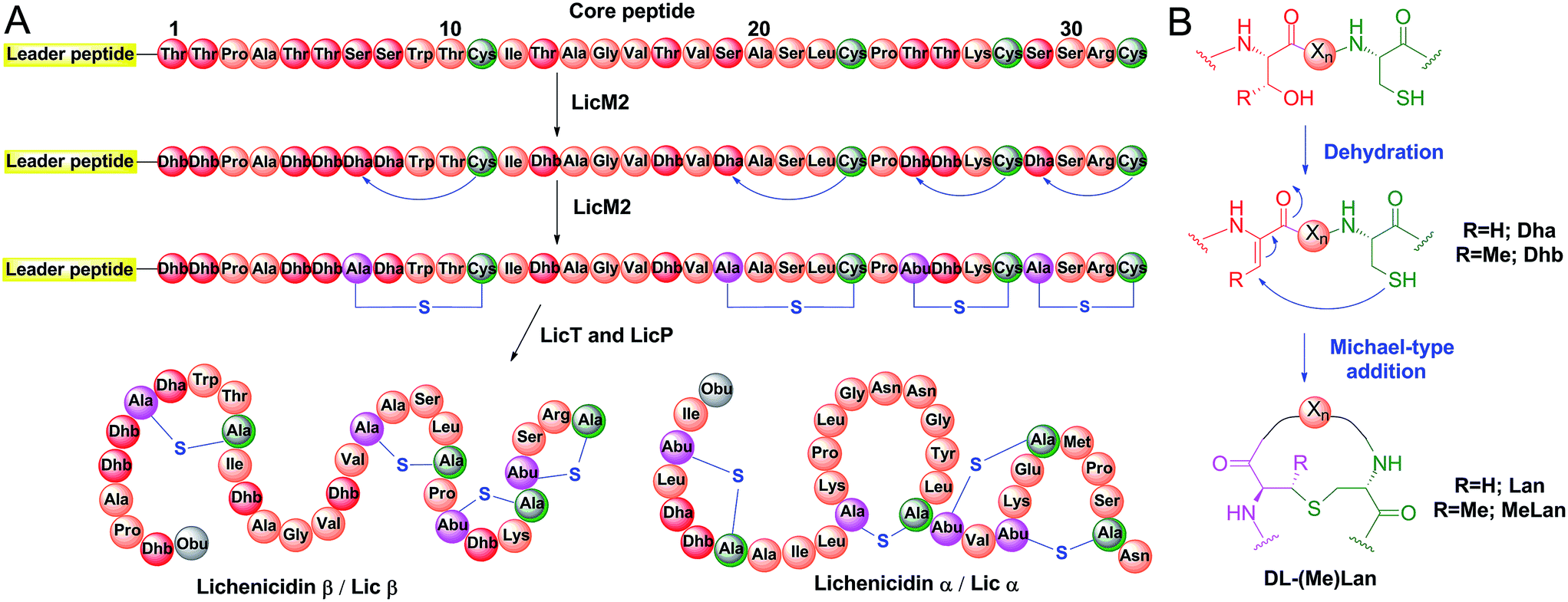
Applications of the class II lanthipeptide protease LicP for sequence-specific, traceless peptide bond cleavage - Chemical Science (RSC Publishing) DOI:10.1039/C5SC02329G
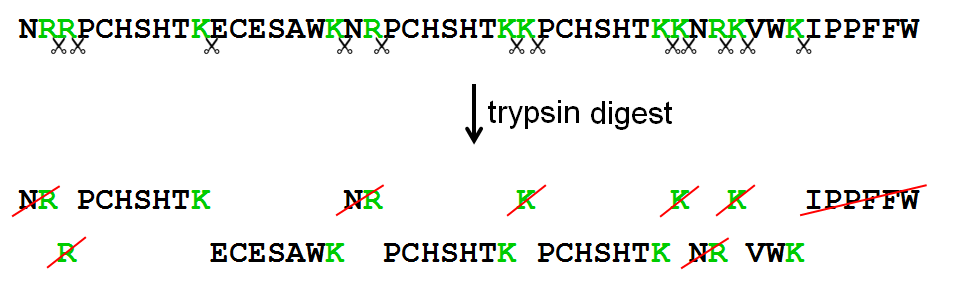
![Sequence of HRPc [1]. The trypsin cleavage sites are indicated by bold... | Download Scientific Diagram Sequence of HRPc [1]. The trypsin cleavage sites are indicated by bold... | Download Scientific Diagram](https://www.researchgate.net/publication/13465969/figure/fig1/AS:602031375003649@1520546873782/Sequence-of-HRPc-1-The-trypsin-cleavage-sites-are-indicated-by-bold-letters-The.png)
Sequence of HRPc [1]. The trypsin cleavage sites are indicated by bold... | Download Scientific Diagram

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal

Marked difference in efficiency of the digestive enzymes pepsin, trypsin, chymotrypsin, and pancreatic elastase to cleave tightly folded proteins

Schematic representation of the major trypsin cleavage sites and one... | Download Scientific Diagram

Global Sequencing of Proteolytic Cleavage Sites in Apoptosis by Specific Labeling of Protein N Termini: Cell

Insight into Trypsin Miscleavage: Comparison of Kinetic Constants of Problematic Peptide Sequences | Analytical Chemistry

Proteomic Identification of Protease Cleavage Sites Characterizes Prime and Non-prime Specificity of Cysteine Cathepsins B, L, and S | Journal of Proteome Research

Proteomic Identification of protease Cleavage Sites (PICS) workflow.... | Download Scientific Diagram
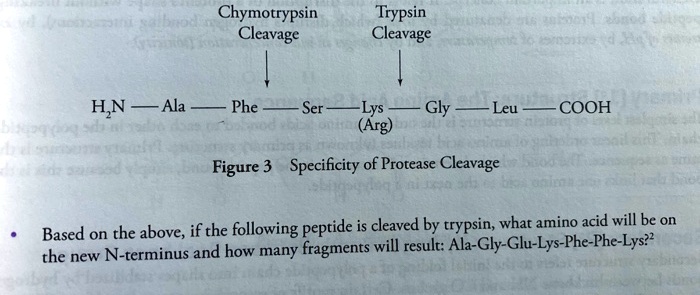
SOLVED: Chymotrypsin Cleavage Trypsin Cleavage HN Ala Phe Ser Lys (Arg) Gly Leu COOH Figure 3 Specificity of Protease Cleavage Based on the above; if the following peptide is cleaved by trypsin,
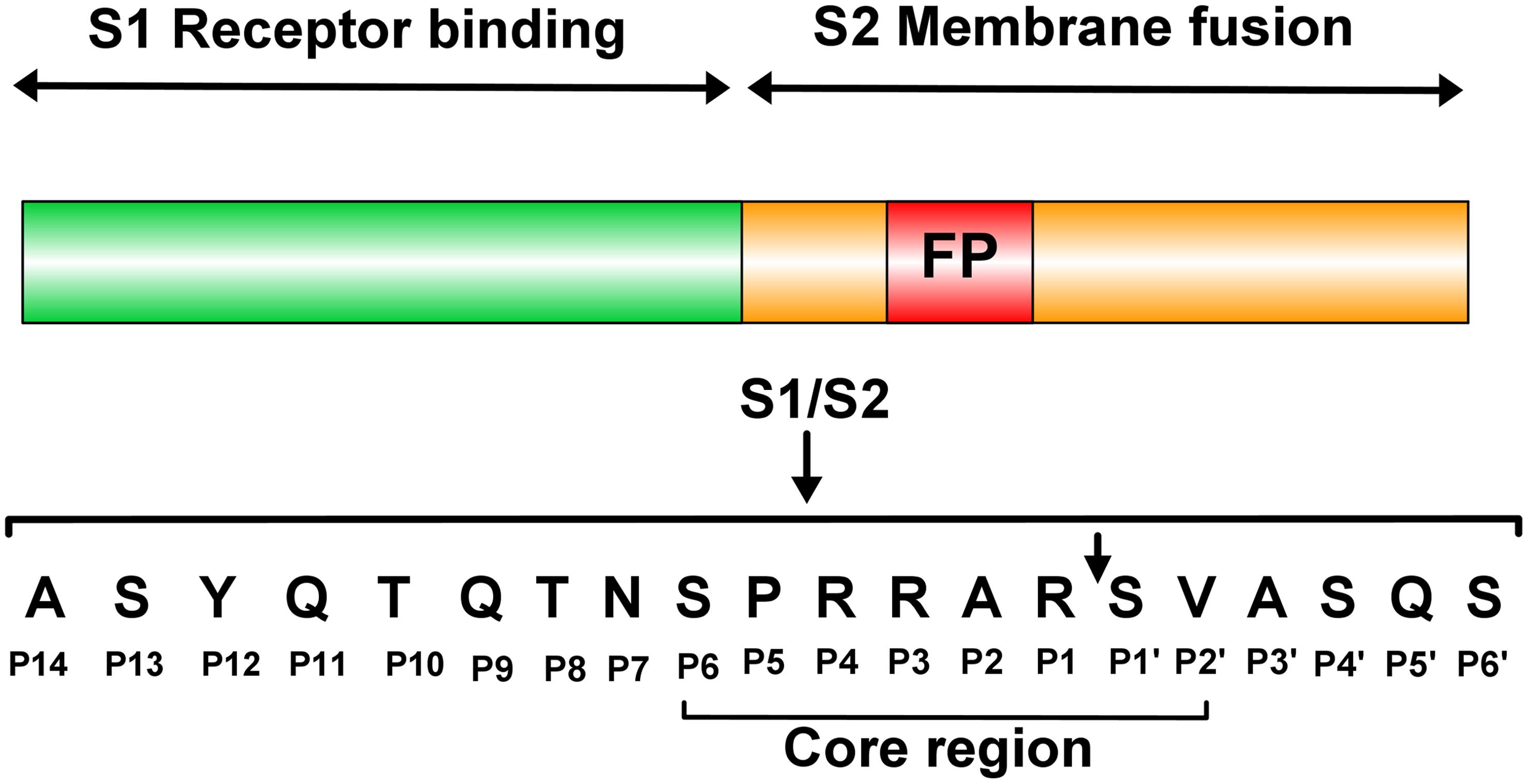
Hypertonic Saline and Aprotinin Inhibit Furin and Nasal Protease to Reduce SARS-CoV-2 Specific Furin Site Cleavage Activity
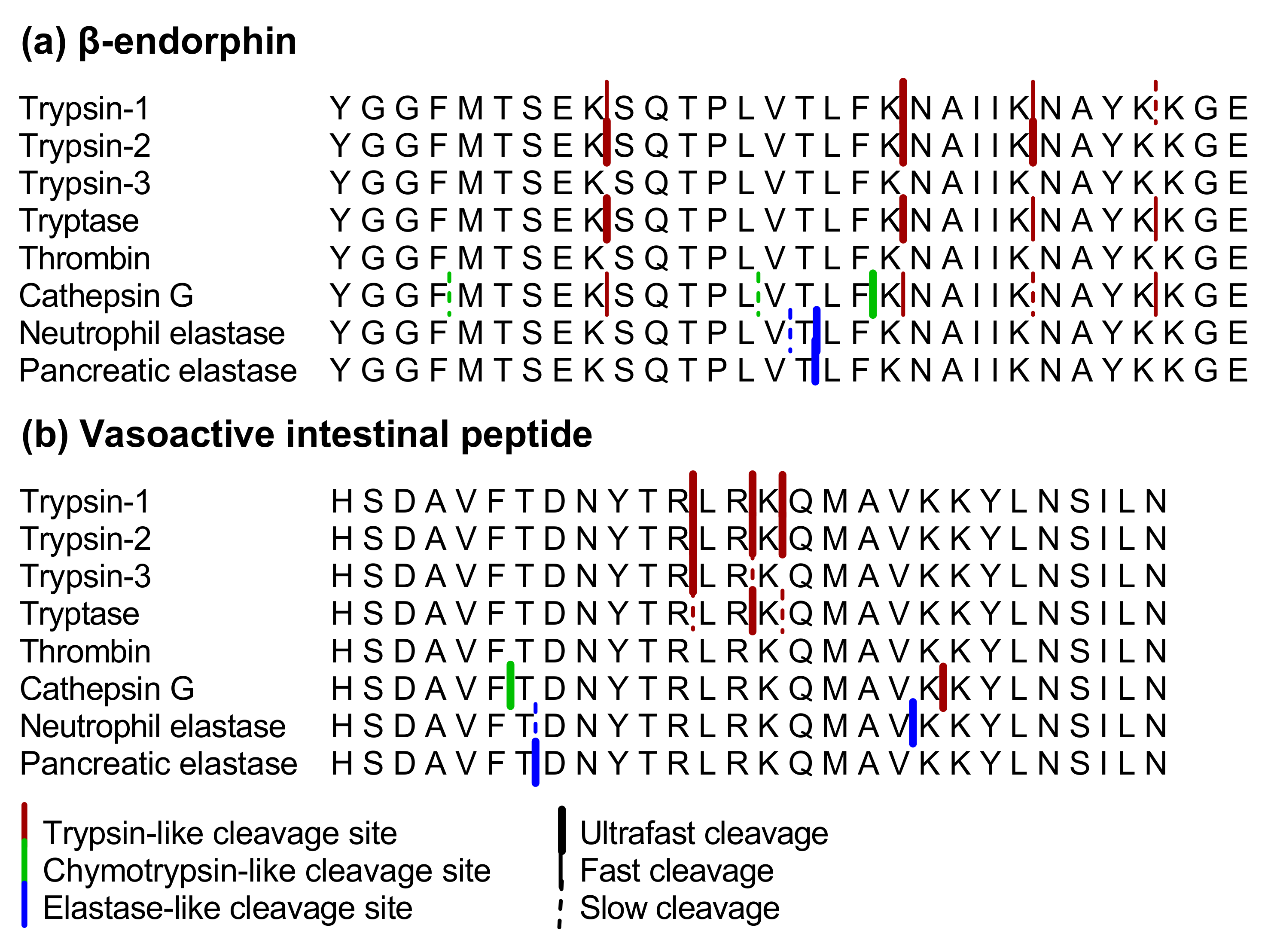
IJMS | Free Full-Text | Proteolytic Cleavage of Bioactive Peptides and Protease-Activated Receptors in Acute and Post-Colitis