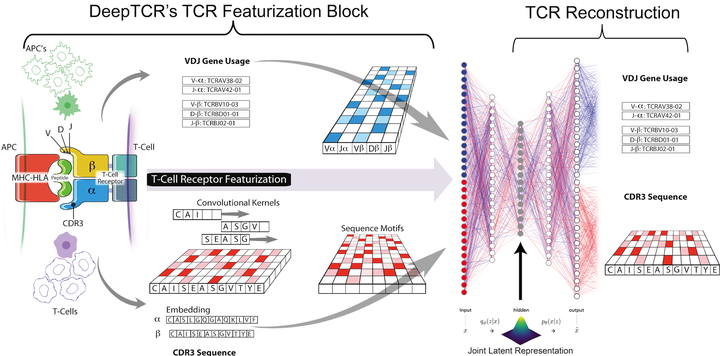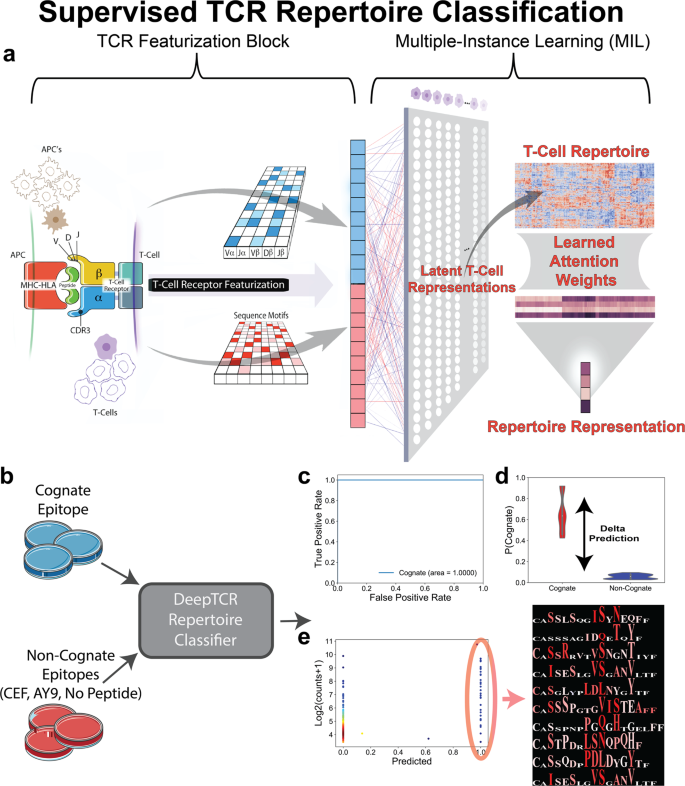
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications

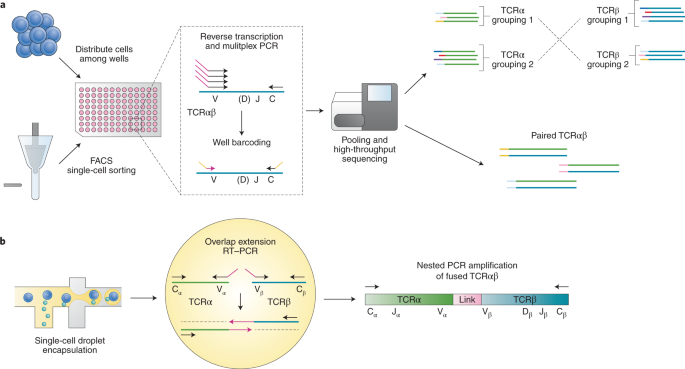
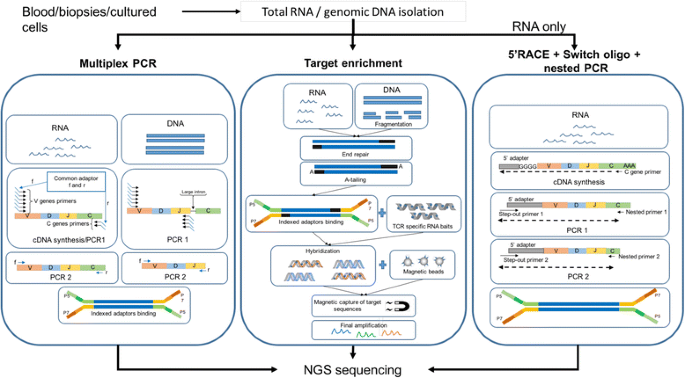
Characterizing immune repertoires by high throughput sequencing: strategies and applications: Trends in Immunology

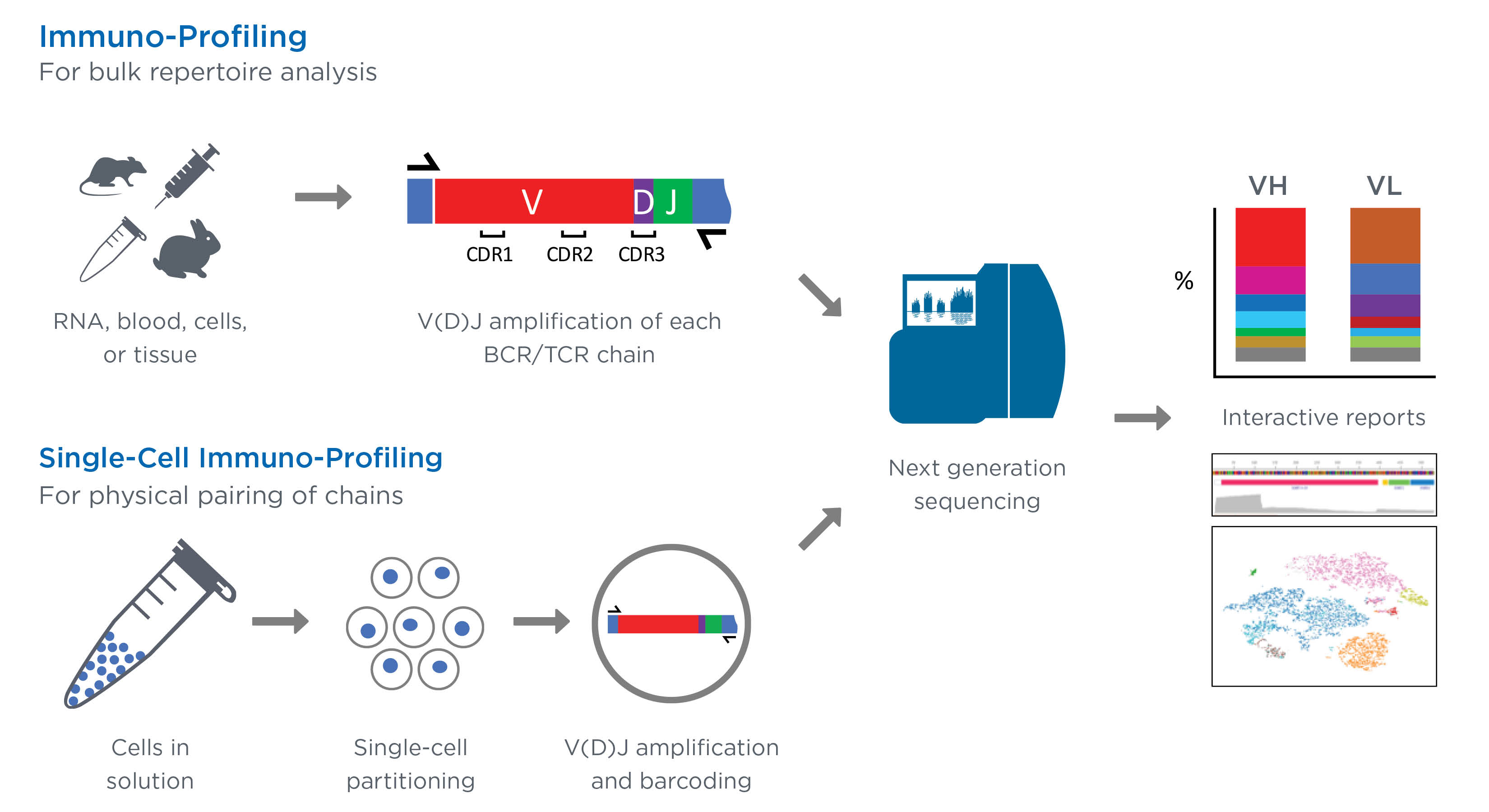
General workflow for TCR repertoire sequencing and analysis. From bulk... | Download Scientific Diagram
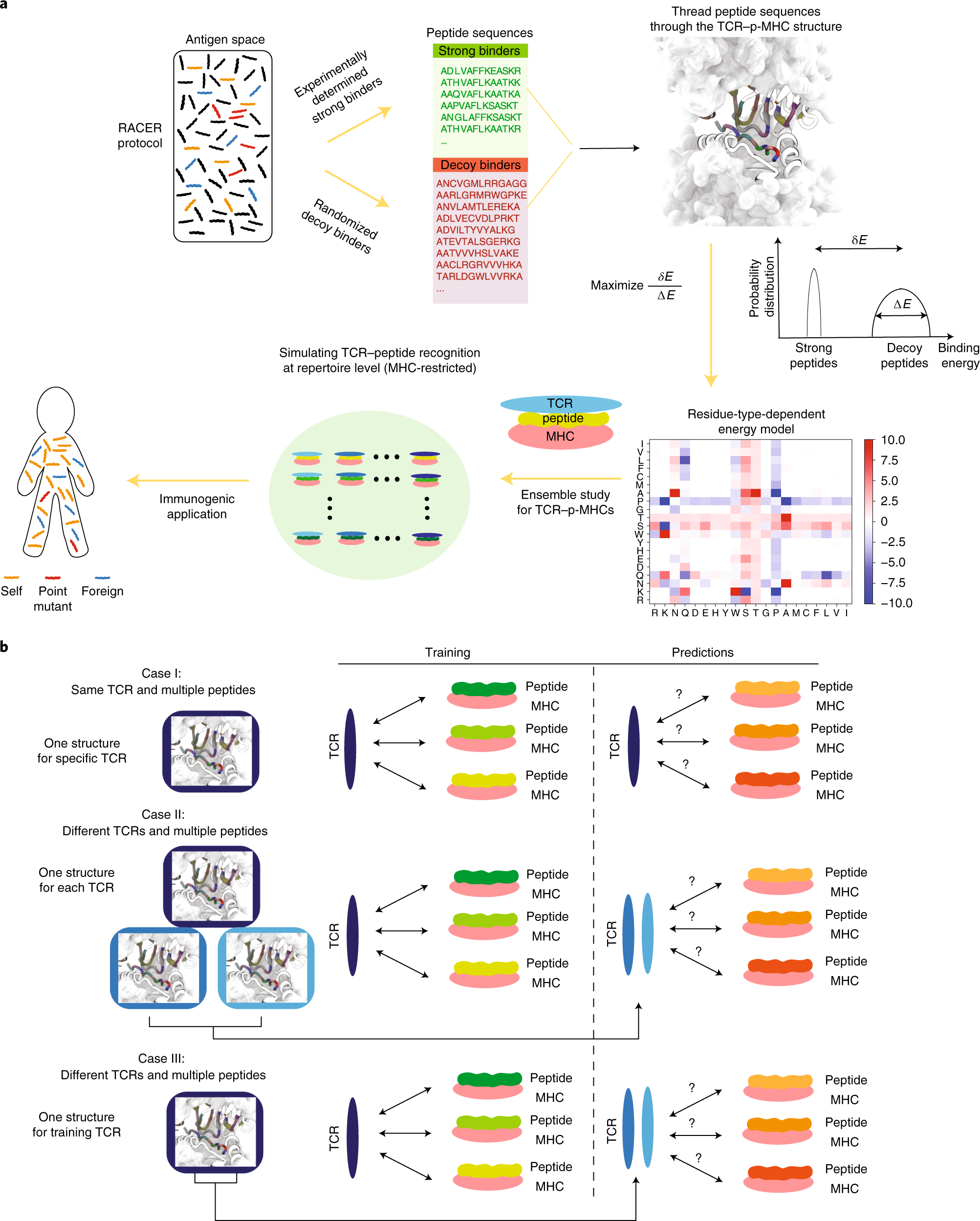
Rapid assessment of T-cell receptor specificity of the immune repertoire | Nature Computational Science
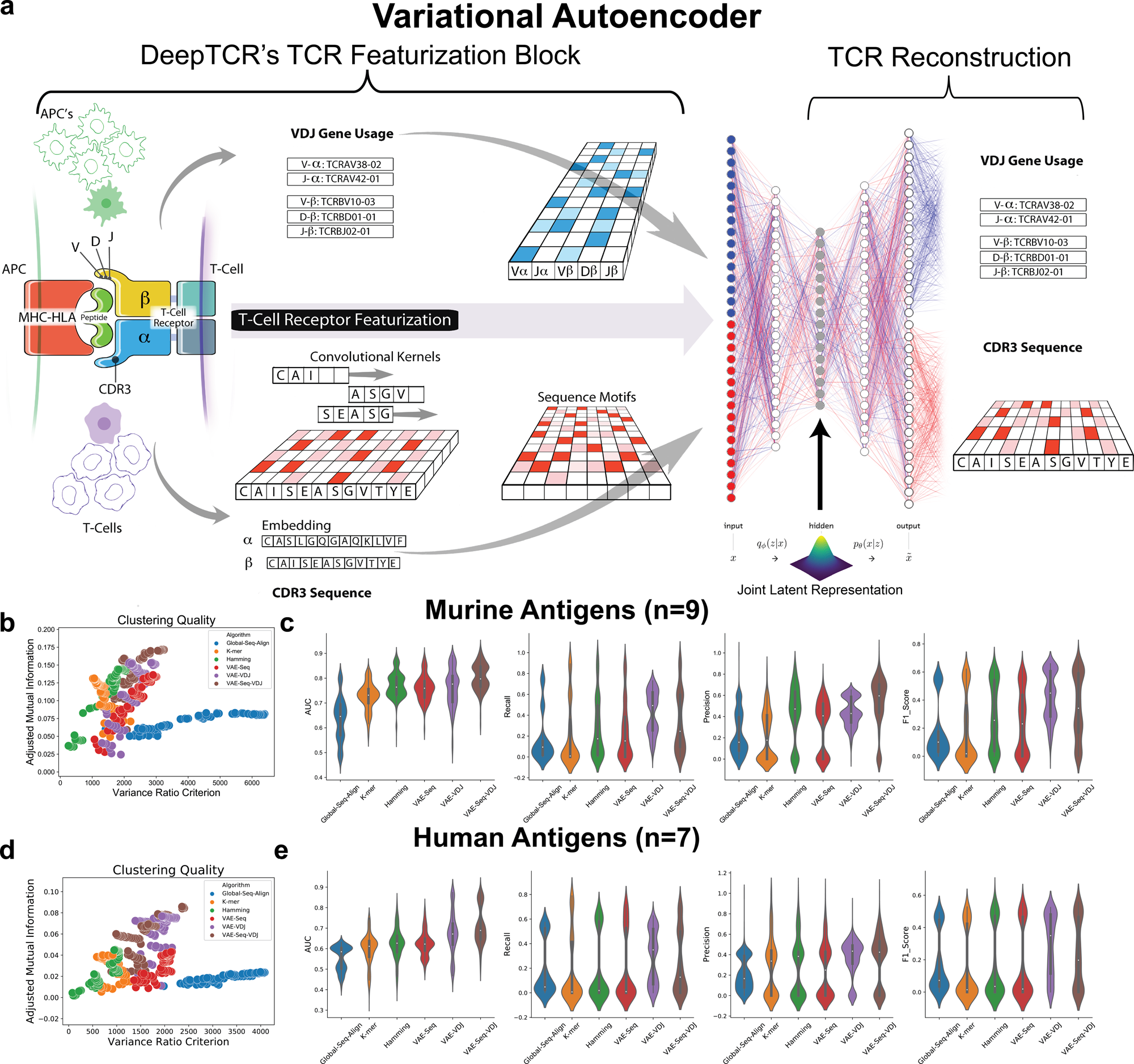
DeepTCR is a deep learning framework for revealing sequence concepts within T-cell repertoires | Nature Communications
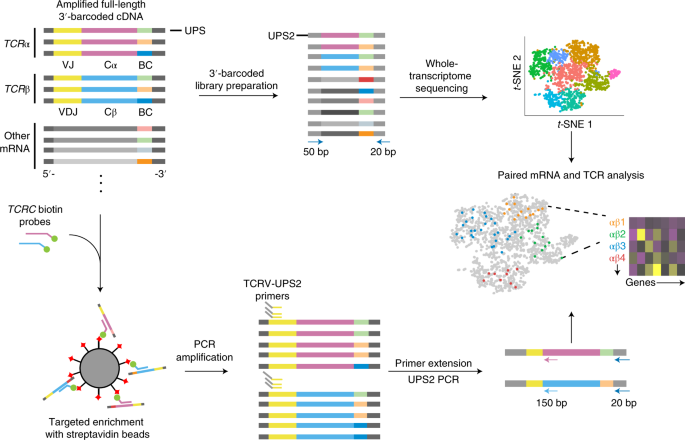
TCR sequencing paired with massively parallel 3′ RNA-seq reveals clonotypic T cell signatures | Nature Immunology

Deep learning reveals predictive sequence concepts within immune repertoires to immunotherapy | Science Advances

Deep autoregressive generative models capture the intrinsics embedded in T-cell receptor repertoires | bioRxiv

Predicting antigen specificity of single T cells based on TCR CDR3 regions | Molecular Systems Biology

Single-cell RNA-TCR-seq and TCRB deep sequencing of PsPD from patient... | Download Scientific Diagram

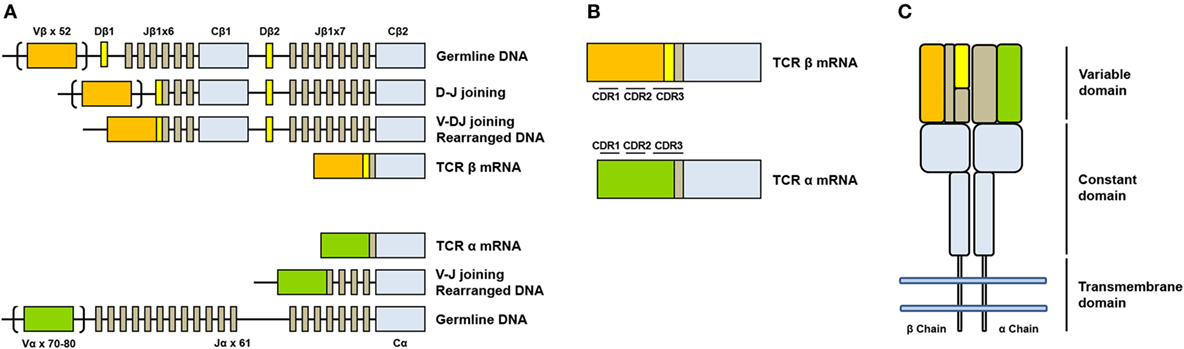
IJMS | Free Full-Text | T-Cell Receptor Repertoire Sequencing and Its Applications: Focus on Infectious Diseases and Cancer
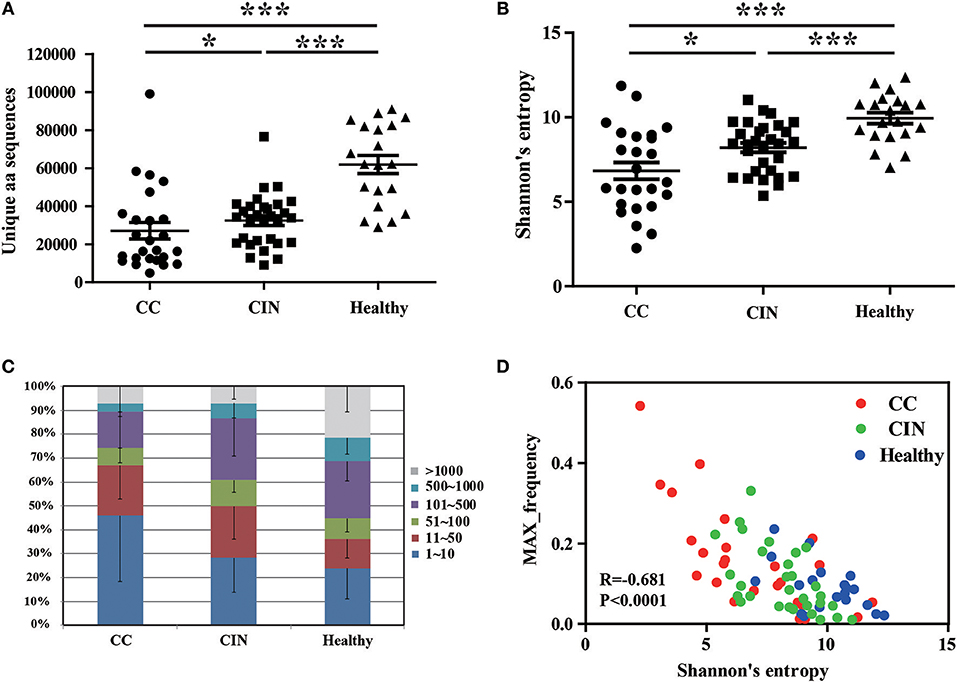
Frontiers | TCR Repertoire as a Novel Indicator for Immune Monitoring and Prognosis Assessment of Patients With Cervical Cancer

Recent advances in T-cell receptor repertoire analysis: Bridging the gap with multimodal single-cell RNA sequencing - ScienceDirect
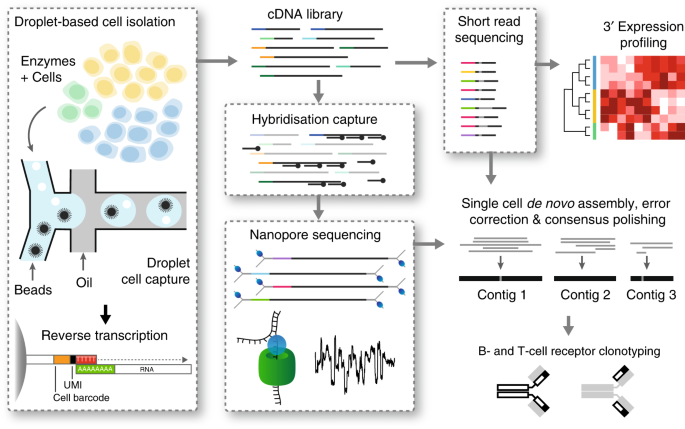
High-throughput targeted long-read single cell sequencing reveals the clonal and transcriptional landscape of lymphocytes | Nature Communications

DeepTCR: a deep learning framework for understanding T-cell receptor sequence signatures within complex T-cell repertoires | bioRxiv
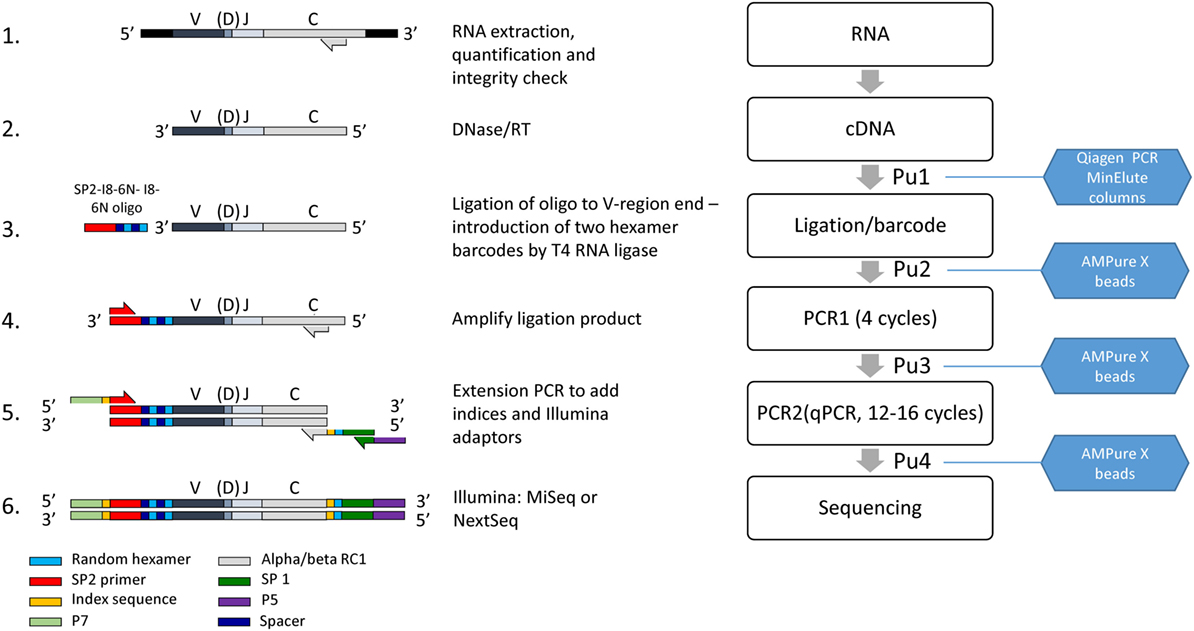
Frontiers | Quantitative Characterization of the T Cell Receptor Repertoire of Naïve and Memory Subsets Using an Integrated Experimental and Computational Pipeline Which Is Robust, Economical, and Versatile