![PDF] Artificial signal peptide prediction by a hidden markov model to improve protein secretion via Lactococcus lactis bacteria | Semantic Scholar PDF] Artificial signal peptide prediction by a hidden markov model to improve protein secretion via Lactococcus lactis bacteria | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d1f3750c21e3739318f590ceb206dc8fbf41f06f/4-Table2-1.png)
PDF] Artificial signal peptide prediction by a hidden markov model to improve protein secretion via Lactococcus lactis bacteria | Semantic Scholar

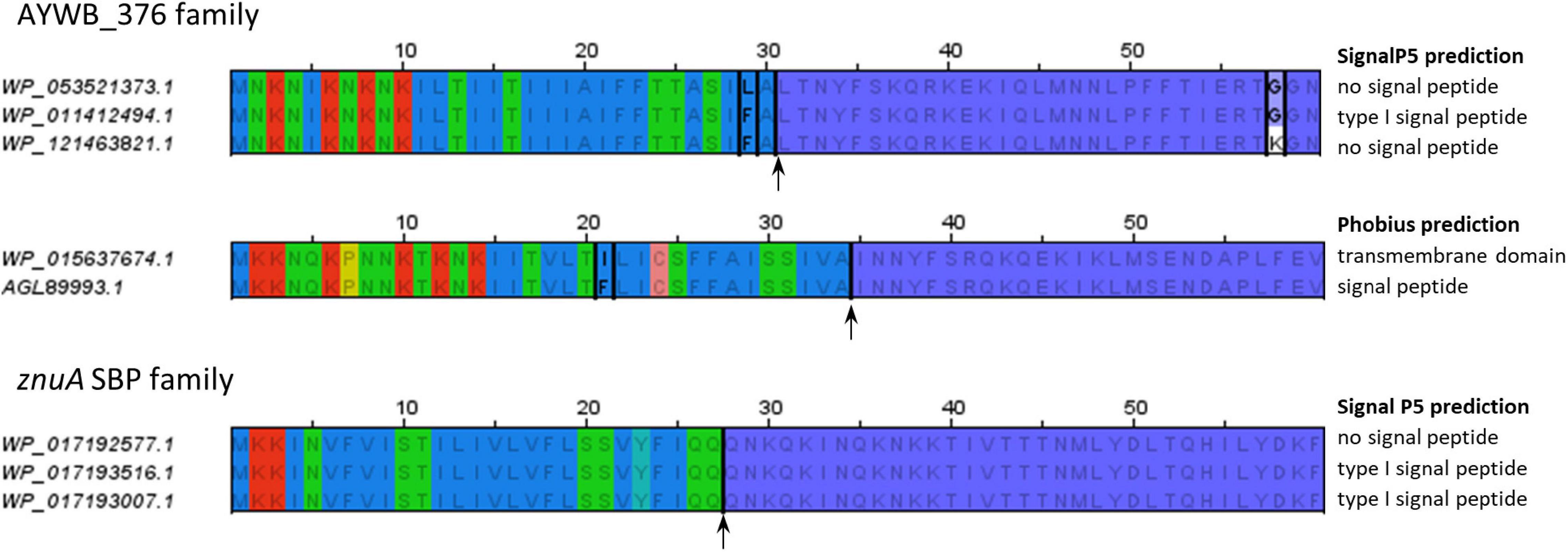
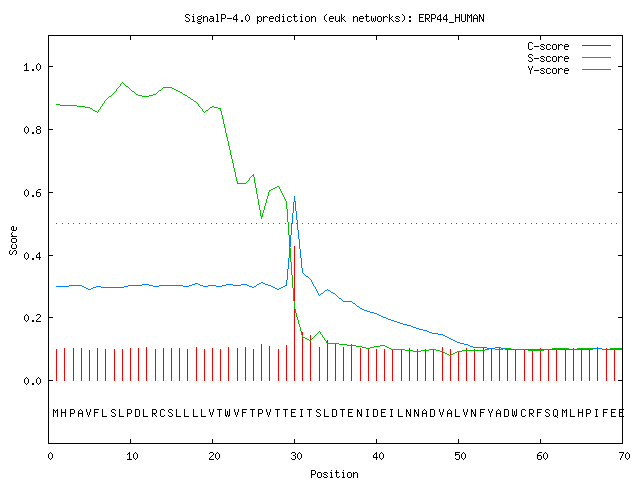
SignalP-5.0 prediction. Signal peptide likelihood was around 0.9505.... | Download Scientific Diagram

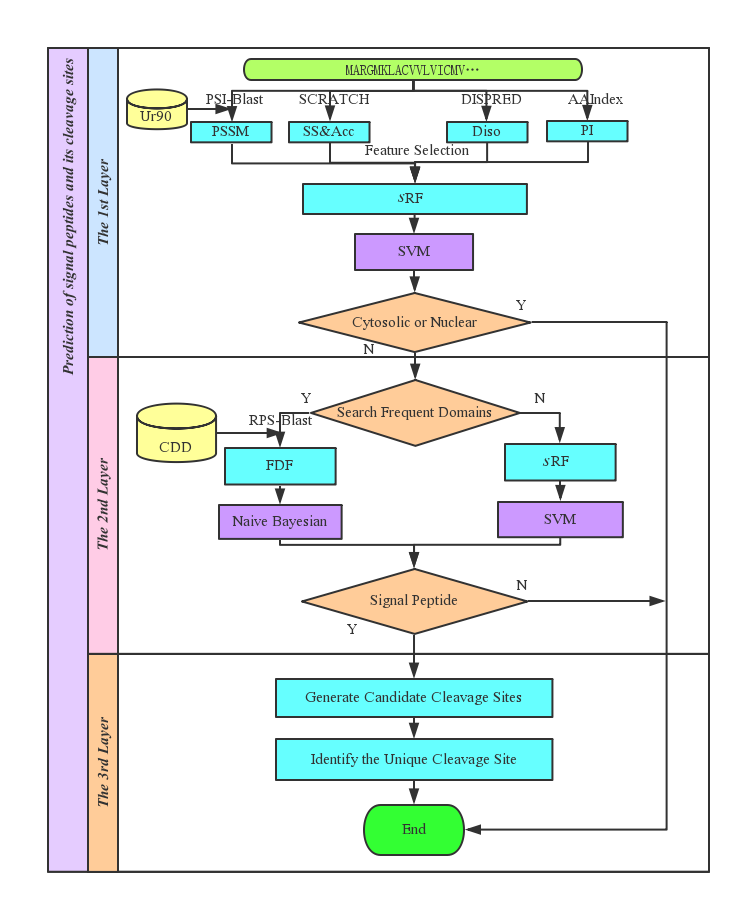
M.W. Mak and S.Y. Kung, ICASSP'09 1 Conditional Random Fields for the Prediction of Signal Peptide Cleavage Sites M.W. Mak The Hong Kong Polytechnic University. - ppt download
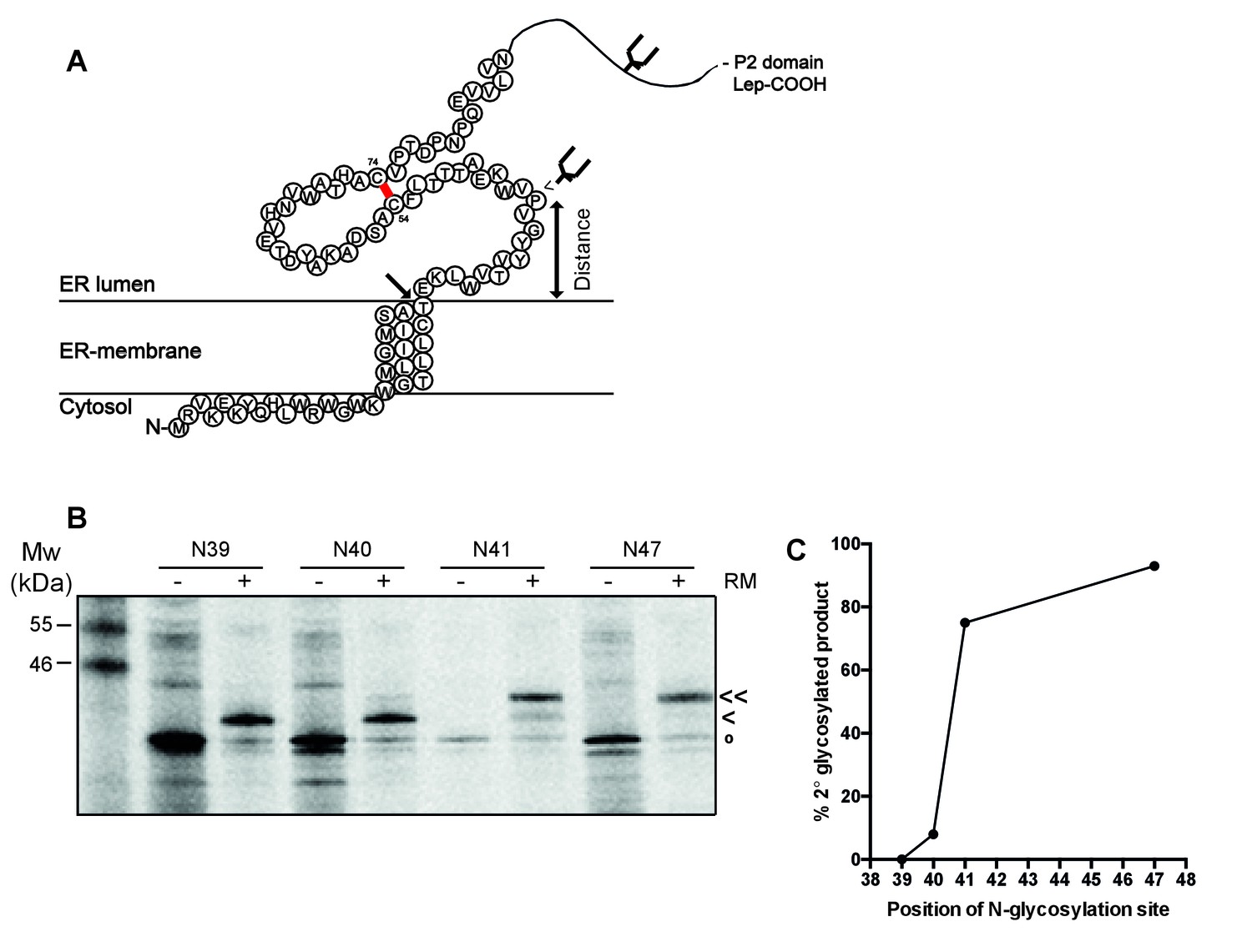
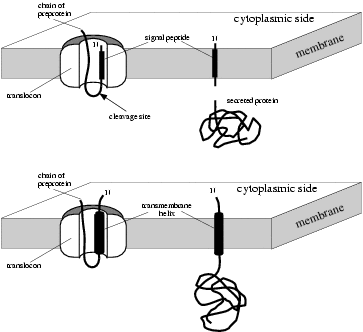
Structure and topology around the cleavage site regulate post-translational cleavage of the HIV-1 gp160 signal peptide | eLife

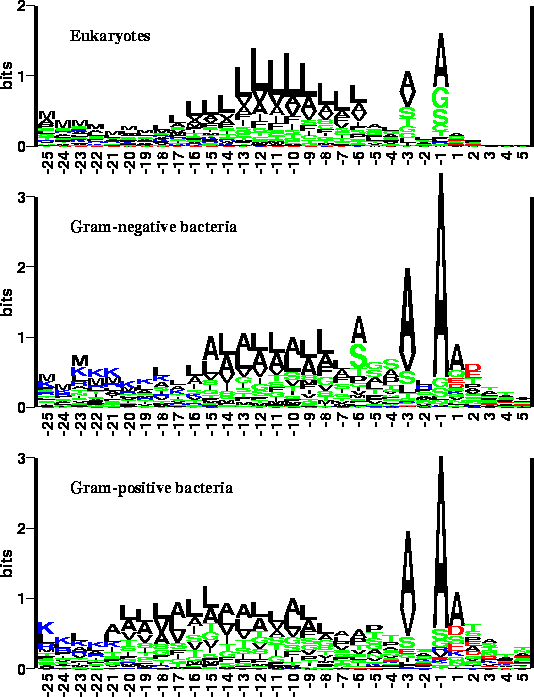
Structure and topology around the cleavage site regulate post-translational cleavage of the HIV-1 gp160 signal peptide | eLife

Nuclear Localization Signal | In Silico Nuclear Localization Signal Prediction | NLS Mapper| - YouTube