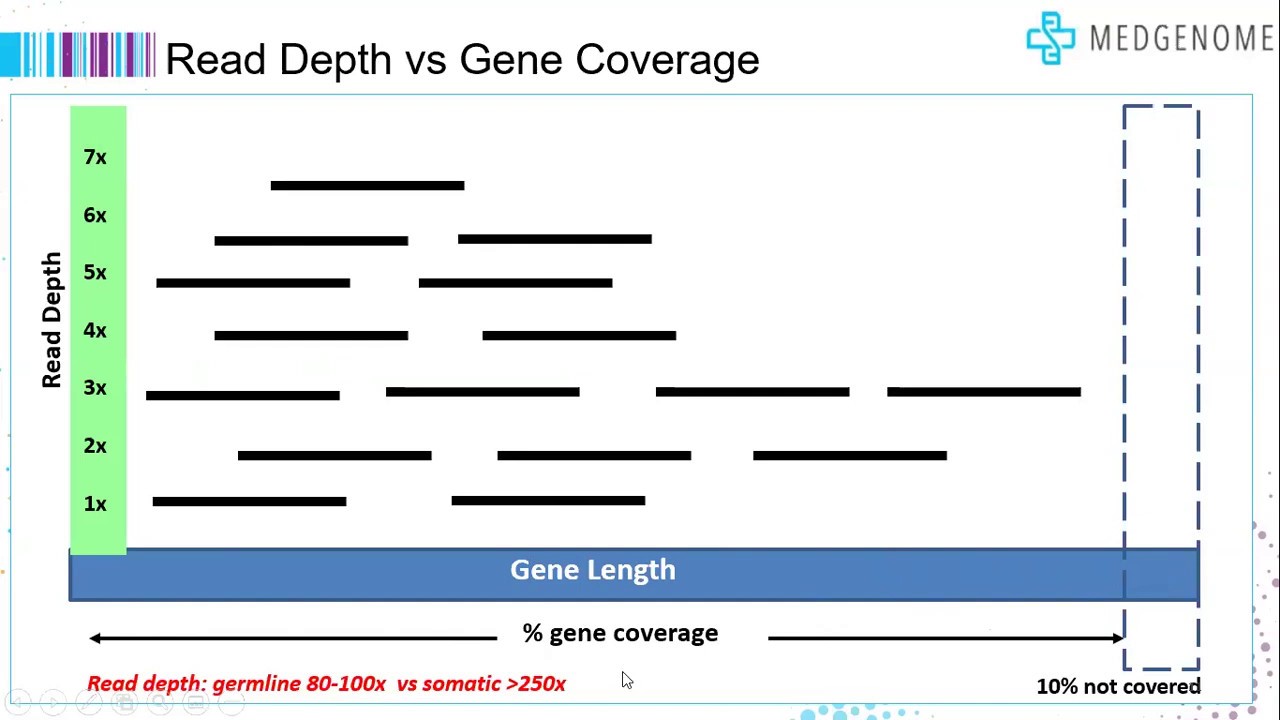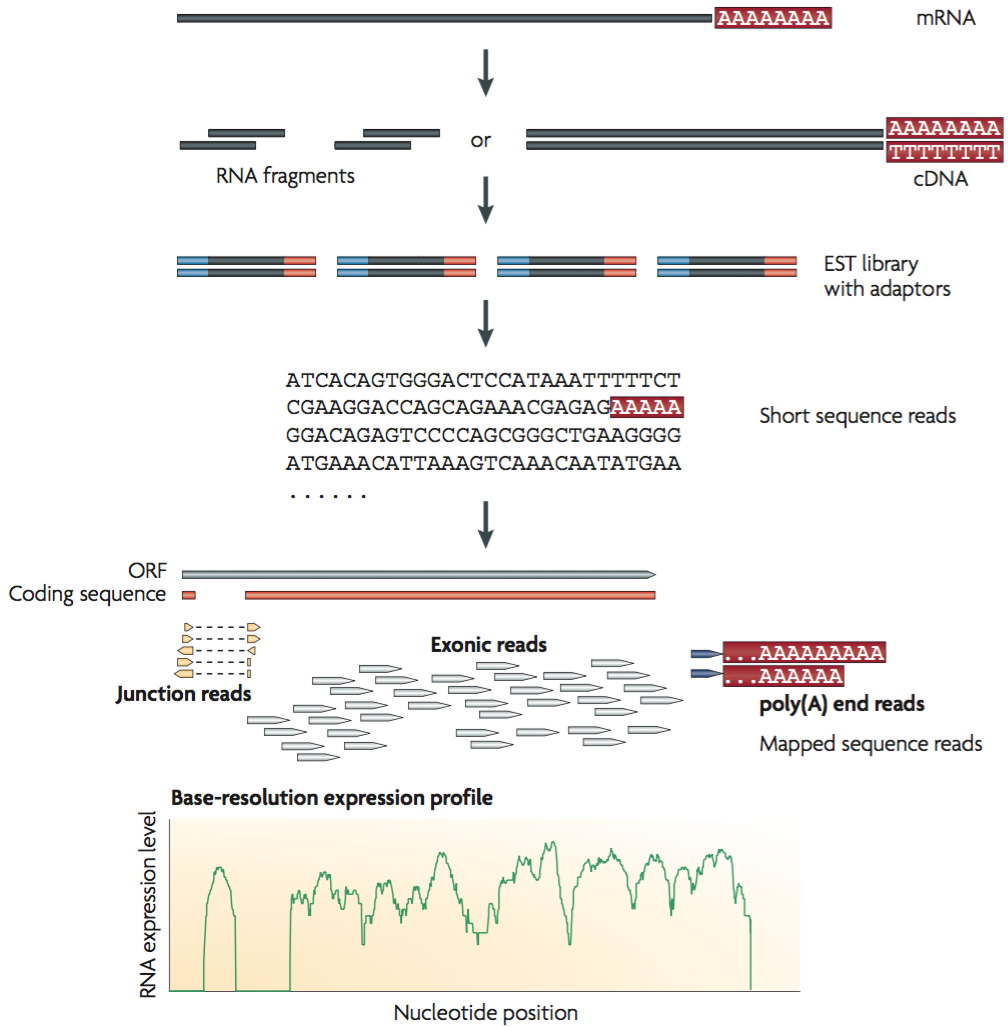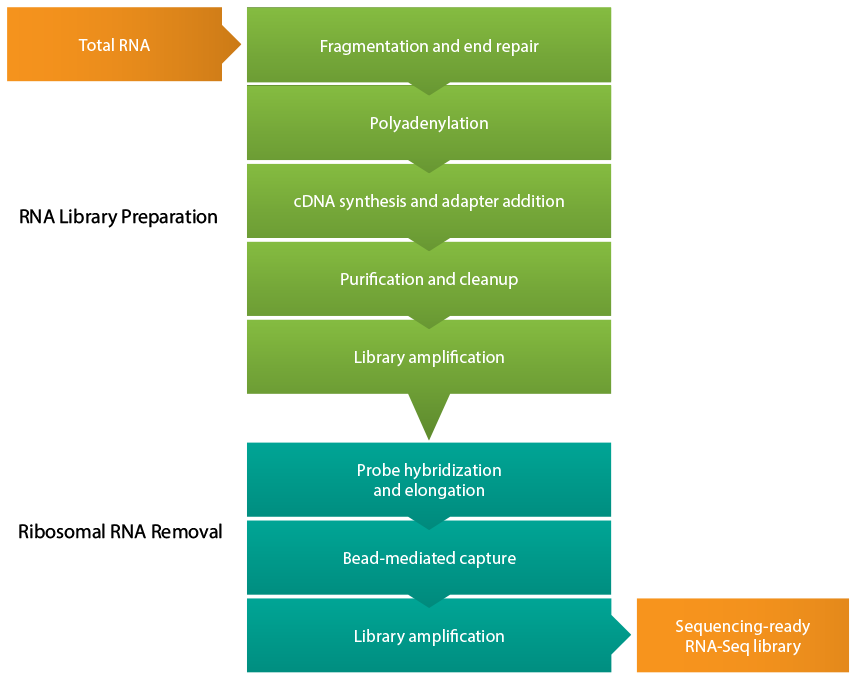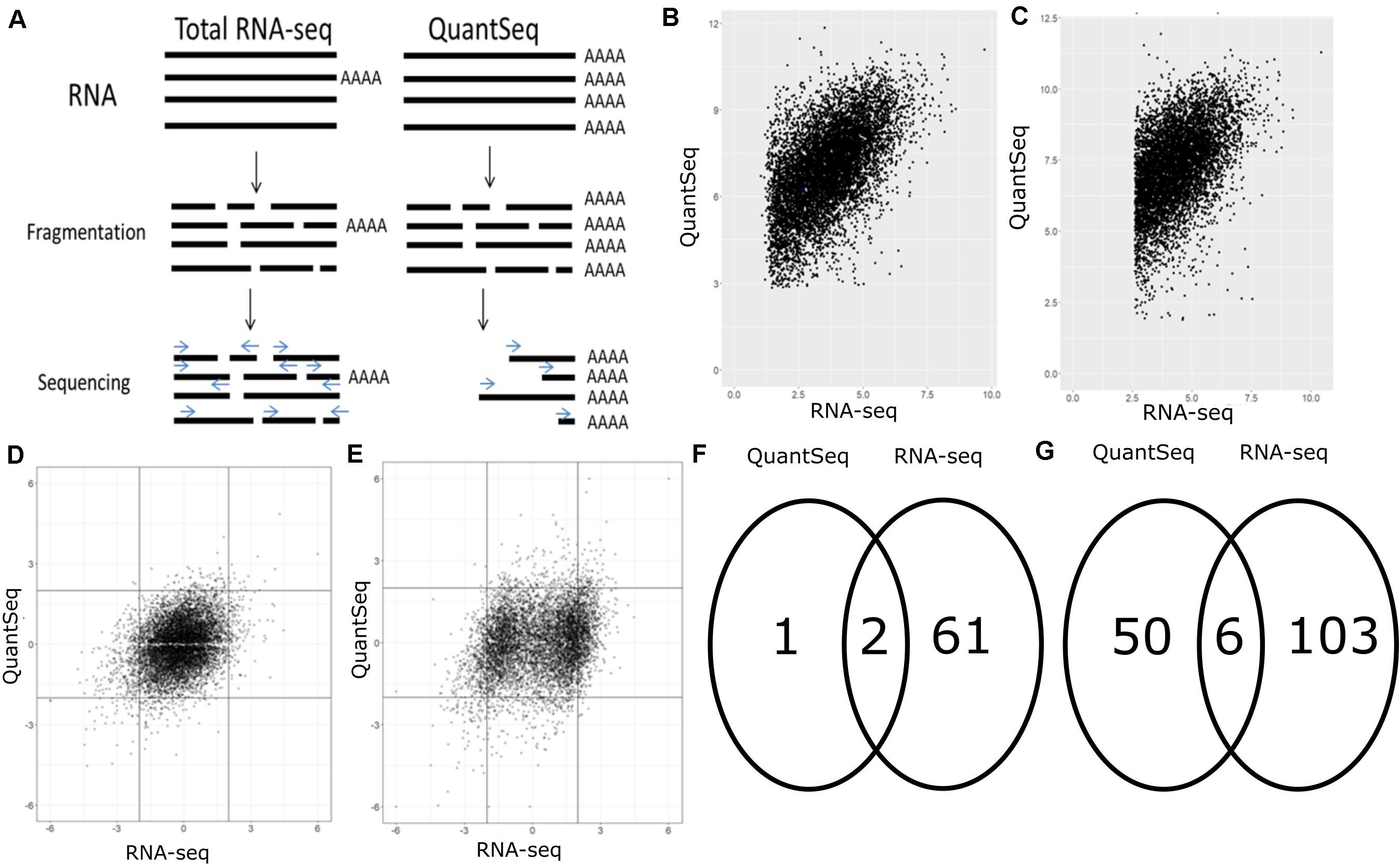
Frontiers | A Comparison of Low Read Depth QuantSeq 3′ Sequencing to Total RNA-Seq in FUS Mutant Mice
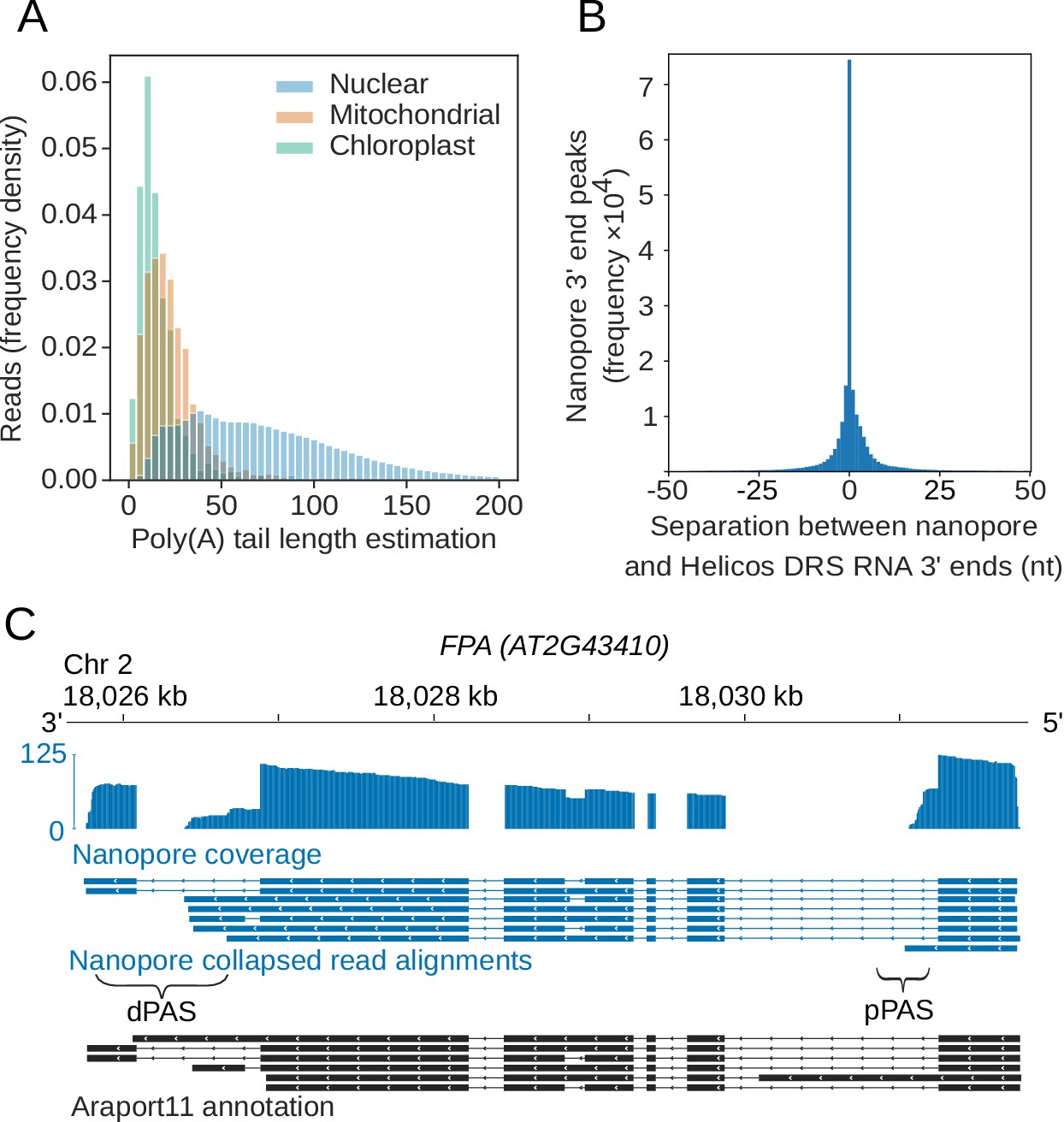
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife

Recommendations for accurate genotyping of SARS-CoV-2 using amplicon-based sequencing of clinical samples - ScienceDirect
Boxplot of sequencing depth data and amplicons size (bp). The range of... | Download Scientific Diagram
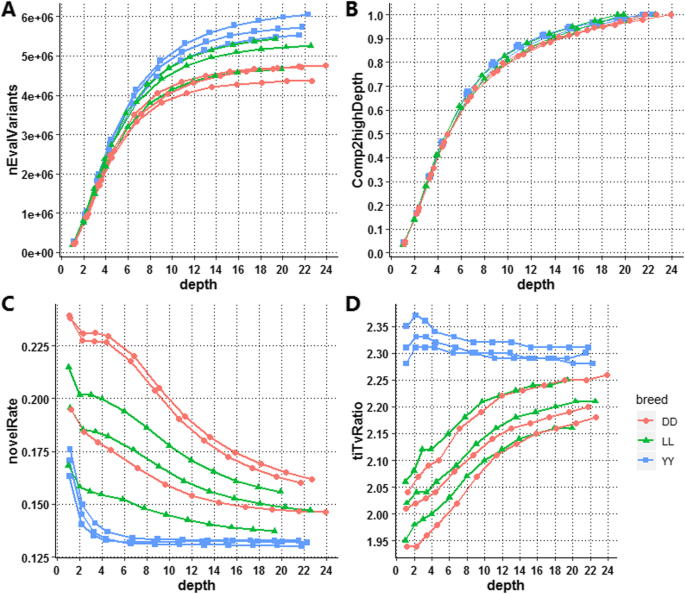
Optimal sequencing depth design for whole genome re-sequencing in pigs | BMC Bioinformatics | Full Text