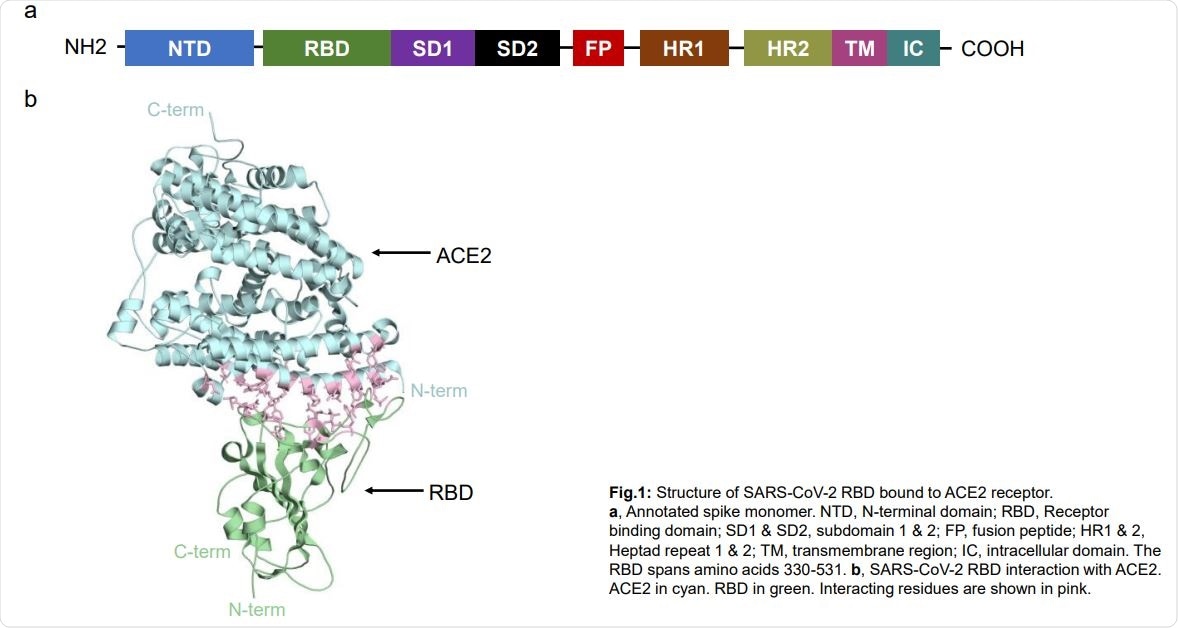
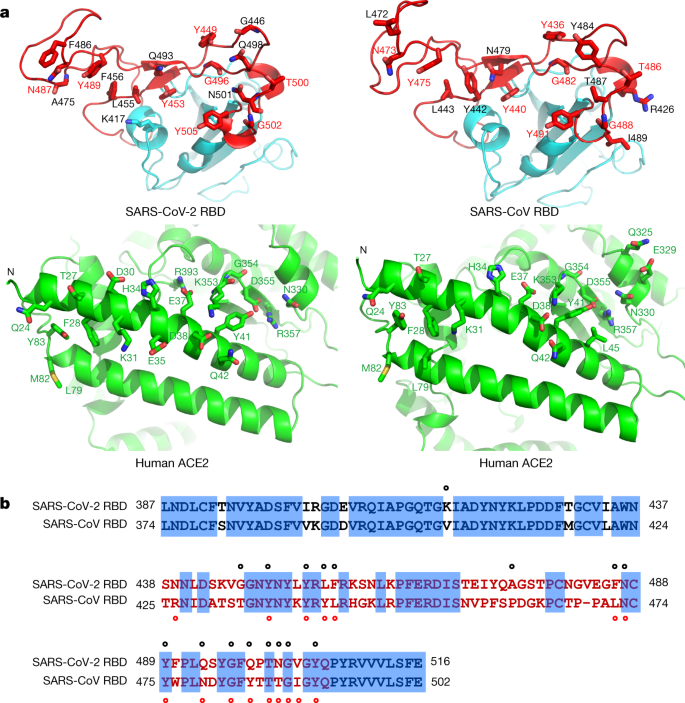
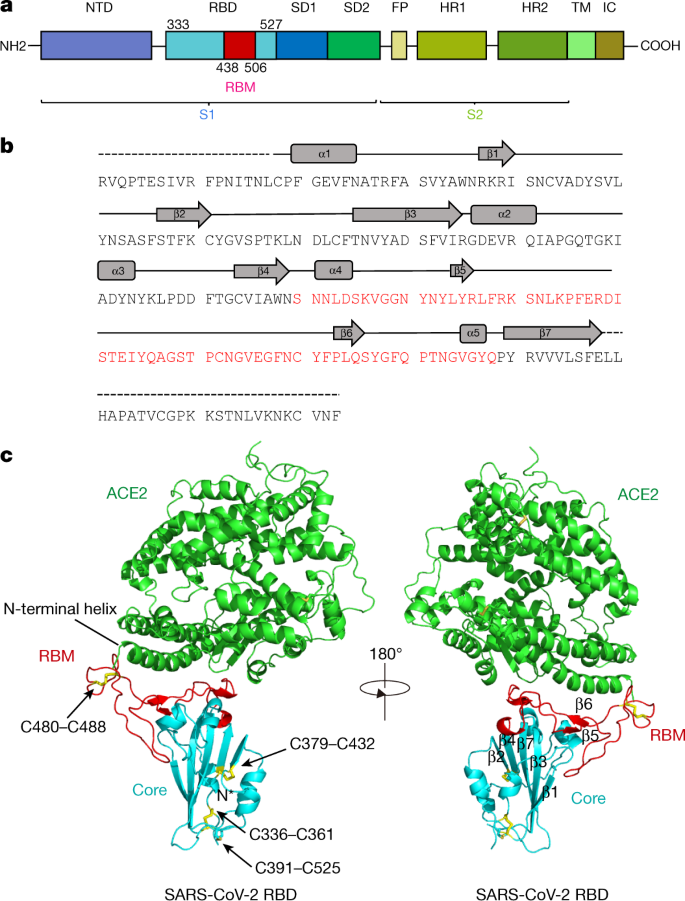
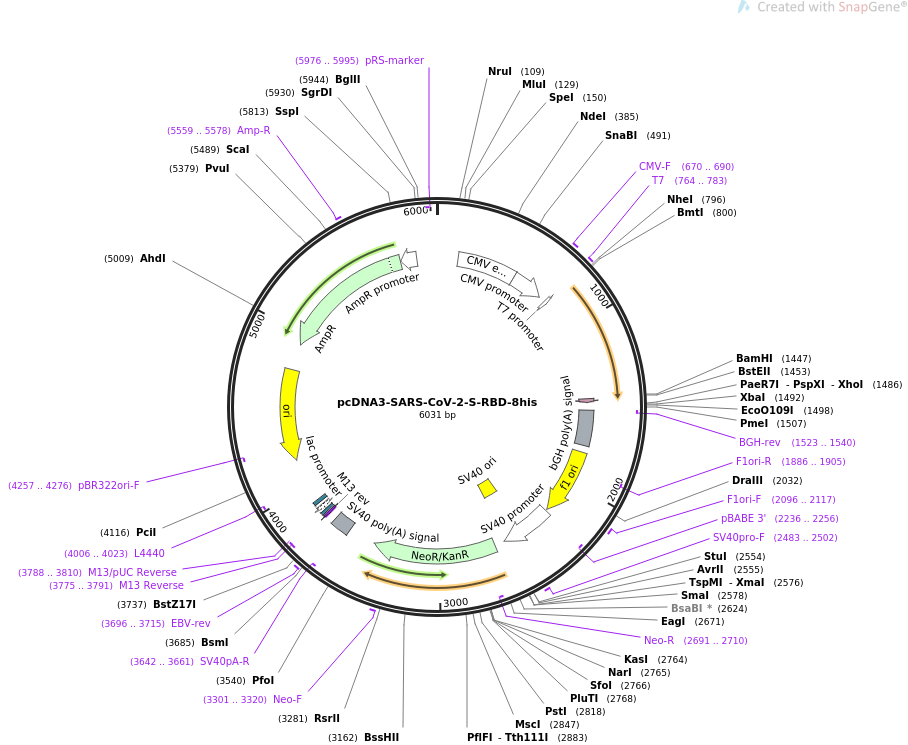
Structural and Functional Characterization of SARS-CoV-2 RBD Domains Produced in Mammalian Cells | Analytical Chemistry
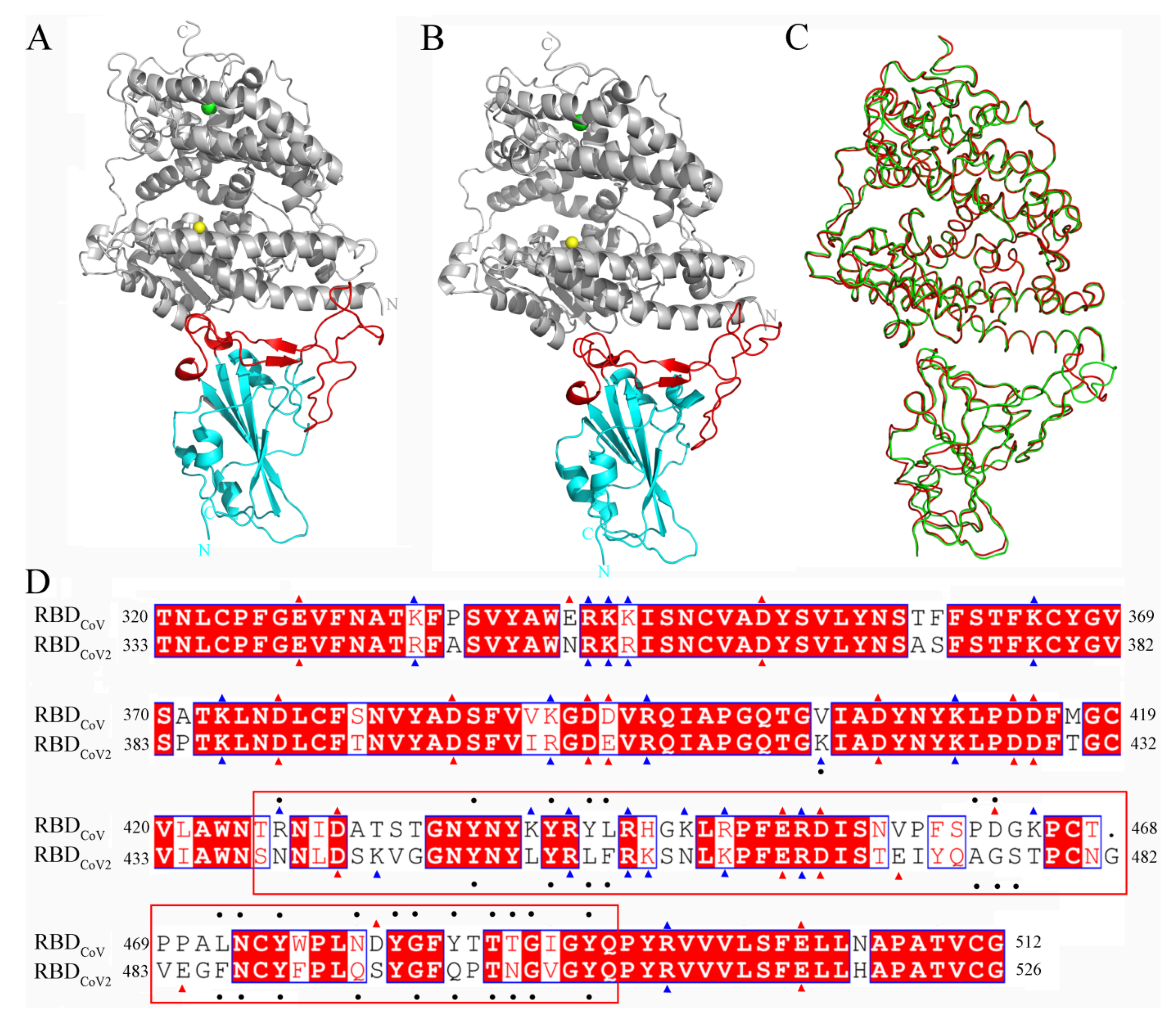
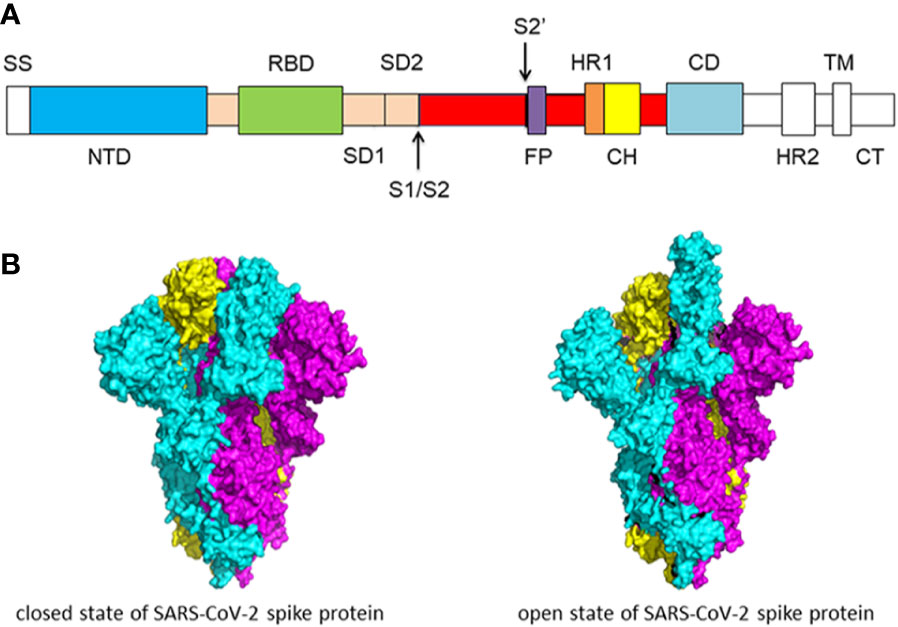
Cells | Free Full-Text | Mechanistic Origin of Different Binding Affinities of SARS-CoV and SARS-CoV-2 Spike RBDs to Human ACE2

JCI - Ultrapotent neutralizing antibodies against SARS-CoV-2 with a high degree of mutation resistance
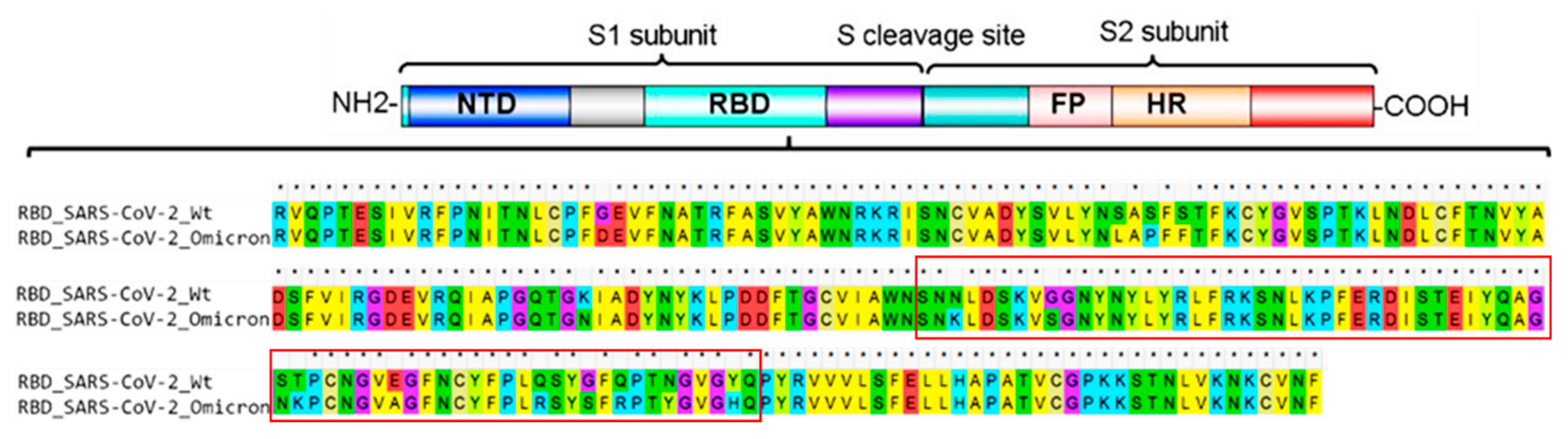
IJMS | Free Full-Text | Improved Binding Affinity of Omicron’s Spike Protein for the Human Angiotensin-Converting Enzyme 2 Receptor Is the Key behind Its Increased Virulence

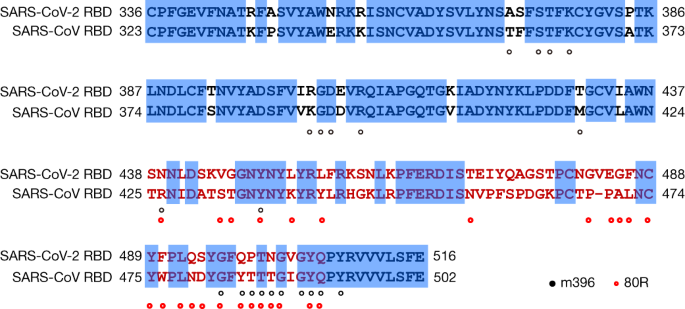
SARS-CoV-2 and SARS-CoV Spike-RBD Structure and Receptor Binding Comparison and Potential Implications on Neutralizing Antibody and Vaccine Development | bioRxiv

Sequence determinants of human-cell entry identified in ACE2-independent bat sarbecoviruses: A combined laboratory and computational network science approach - eBioMedicine

Deep Mutational Scanning of SARS-CoV-2 Receptor Binding Domain Reveals Constraints on Folding and ACE2 Binding - ScienceDirect
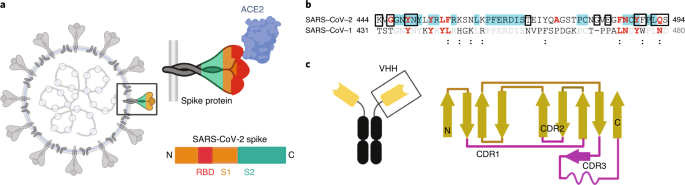
Neutralizing nanobodies bind SARS-CoV-2 spike RBD and block interaction with ACE2 | Nature Structural & Molecular Biology
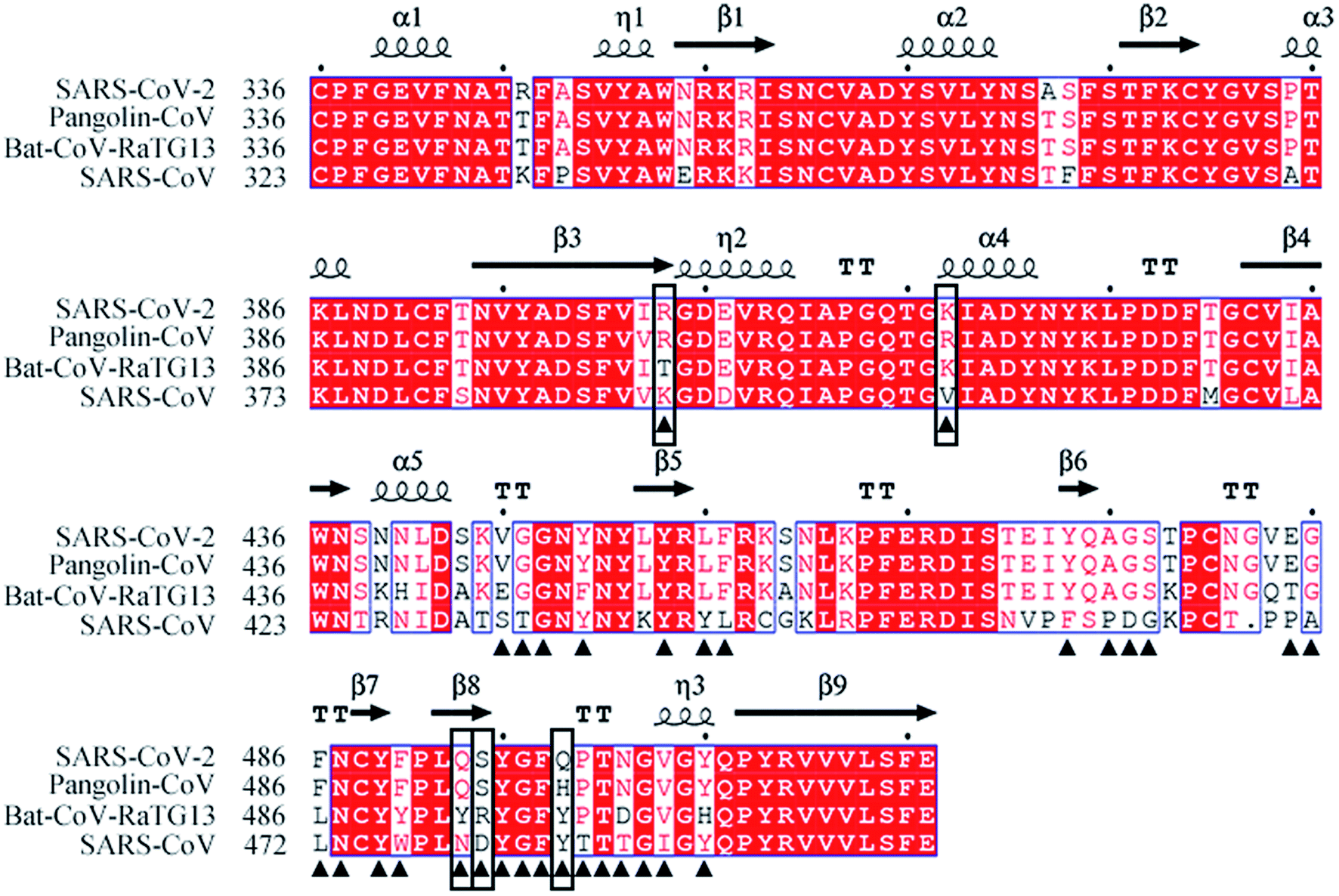
Insight into the origin of SARS-CoV-2 through structural analysis of receptor recognition: a molecular simulation study - RSC Advances (RSC Publishing) DOI:10.1039/D1RA00127B
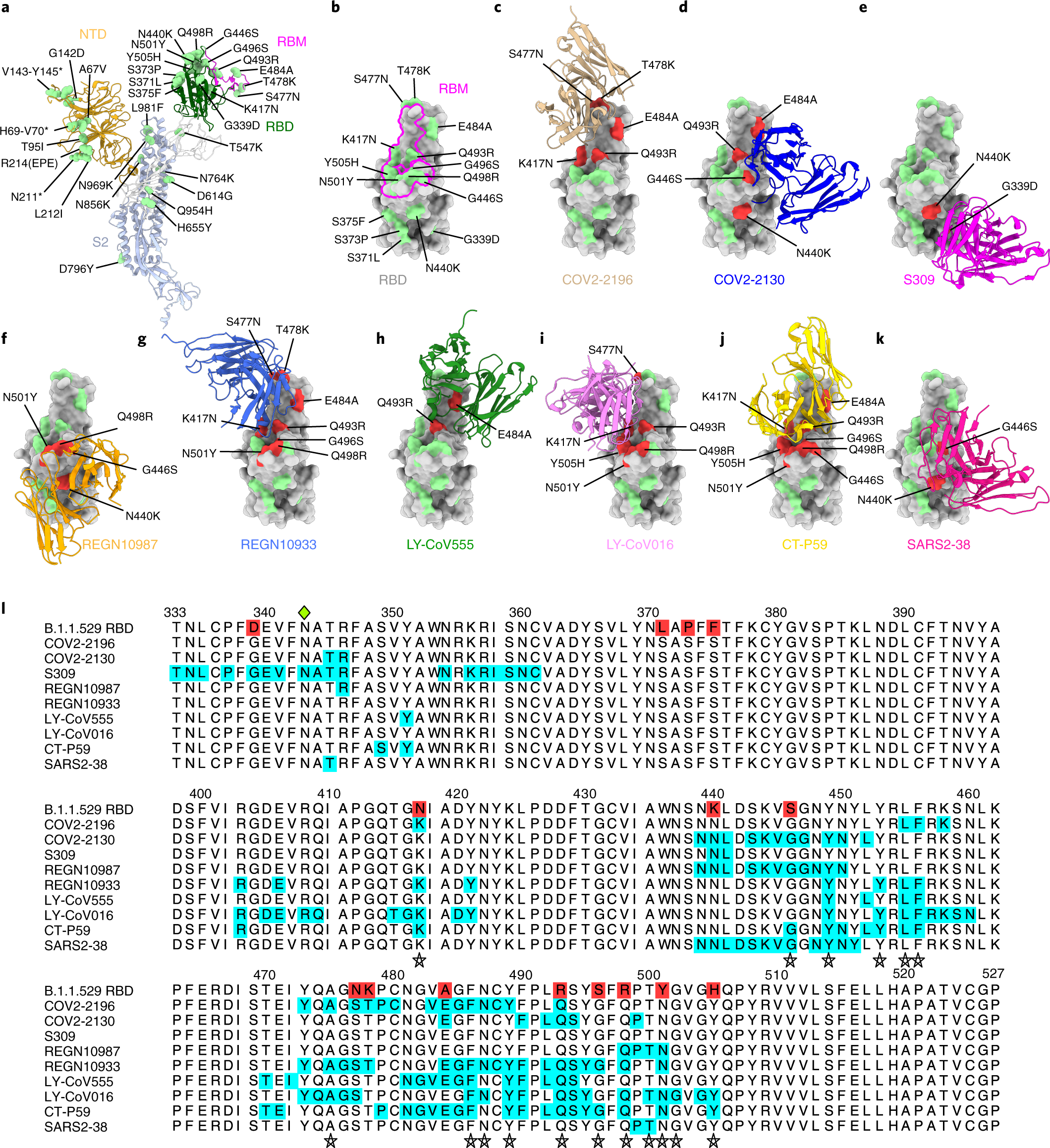

An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies | Nature Medicine
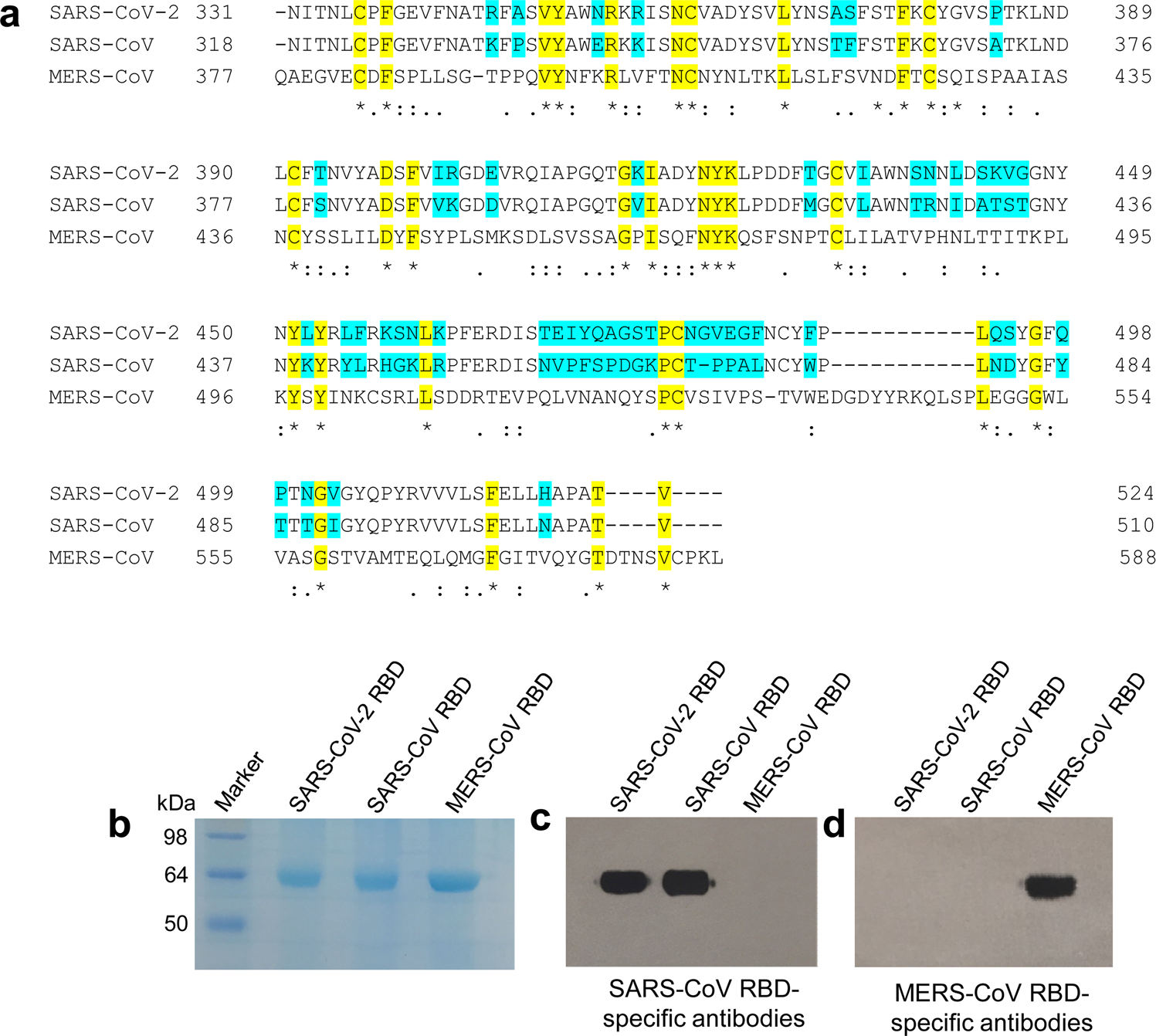
Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine | Cellular & Molecular Immunology

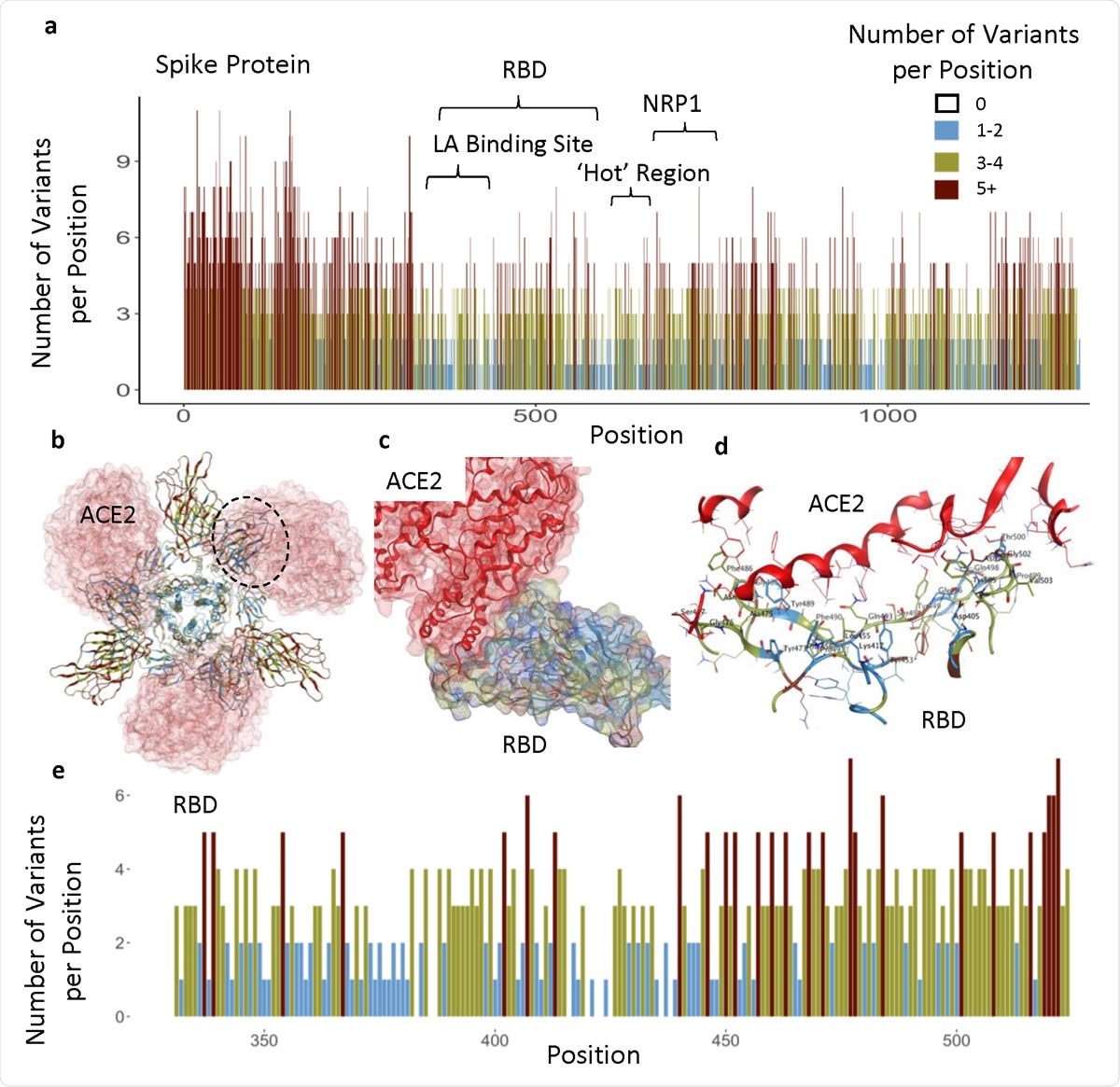
Analysis of the mutation dynamics of SARS-CoV-2 reveals the spread history and emergence of RBD mutant with lower ACE2 binding affinity | bioRxiv

All-Atom Simulations of Human ACE2-Spike Protein RBD Complexes for SARS-CoV-2 and Some of its Variants: Nature of Interactions and Free Energy Diagrams for Dissociation of the Protein Complexes | The Journal of
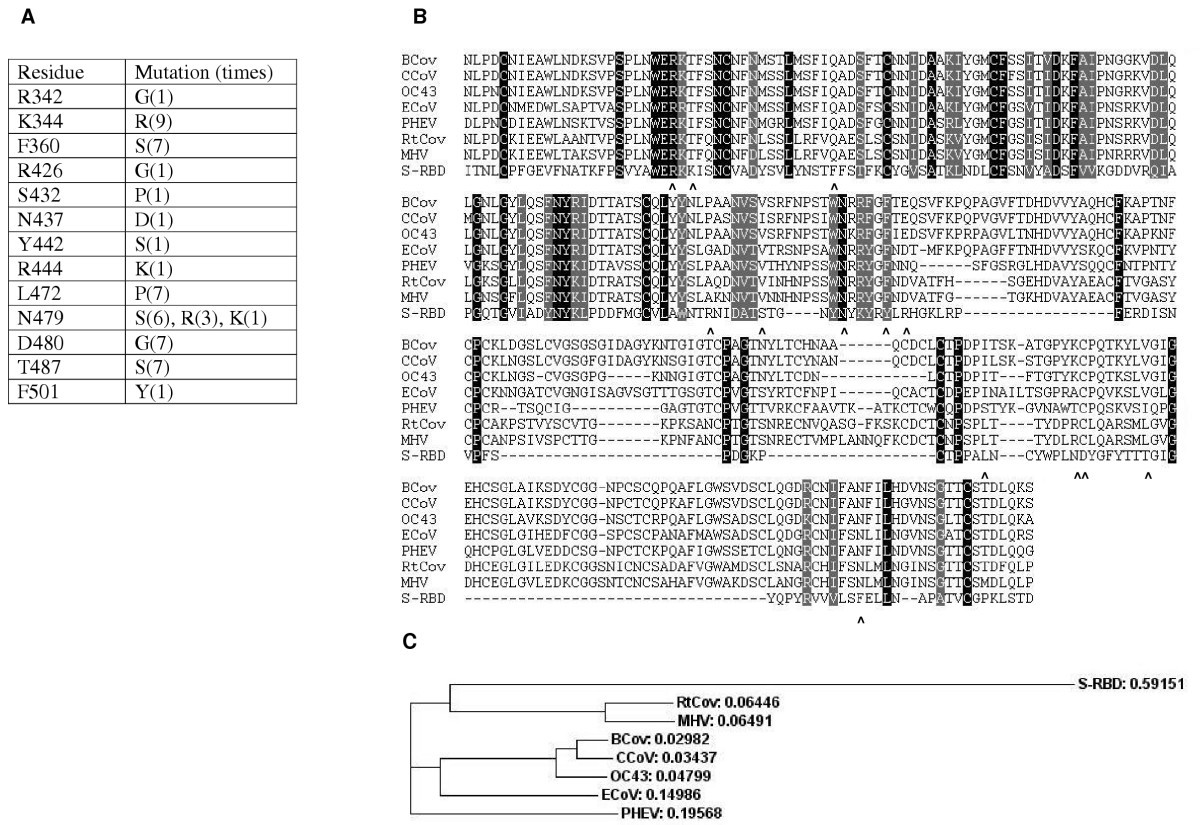

The SARS Coronavirus S Glycoprotein Receptor Binding Domain: Fine Mapping and Functional Characterization | Virology Journal | Full Text
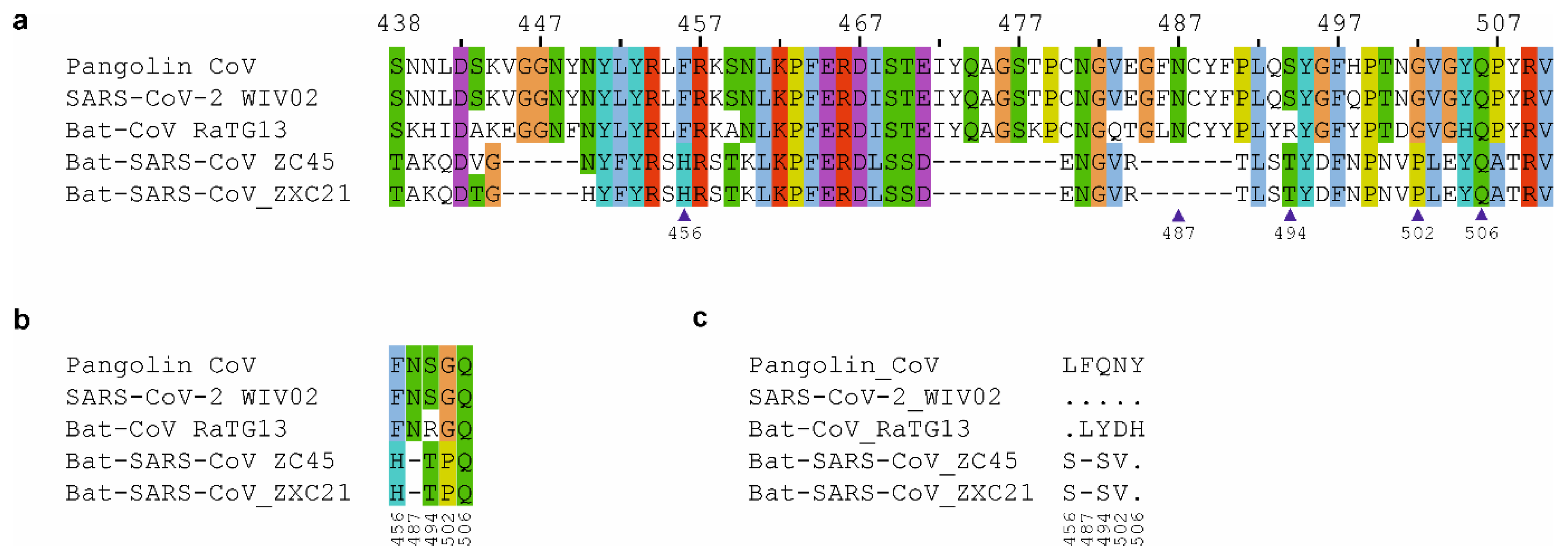
nCoV-2019 Spike Protein Receptor Binding Domain Shares High Amino Acid Identity With a Coronavirus Recovered from a Pangolin Viral Metagenomic Dataset - nCoV-2019 Evolutionary History - Virological