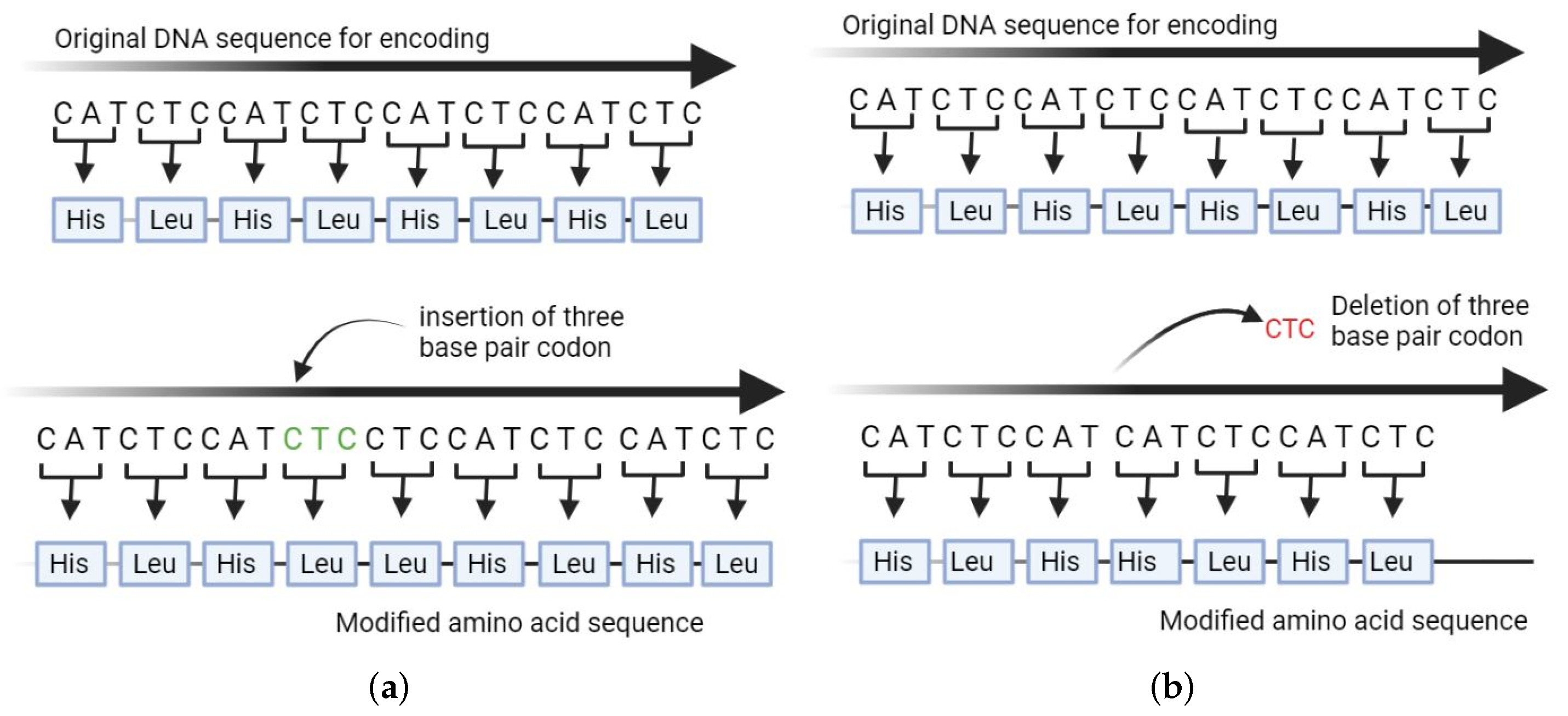
ProtSeq: Toward high-throughput, single-molecule protein sequencing via amino acid conversion into DNA barcodes - ScienceDirect
![Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ] Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/cs-684/1/fig-2-full.png)
Navigating the amino acid sequence space between functional proteins using a deep learning framework [PeerJ]

Frontiers | Amino Acid Reduction Can Help to Improve the Identification of Antimicrobial Peptides and Their Functional Activities
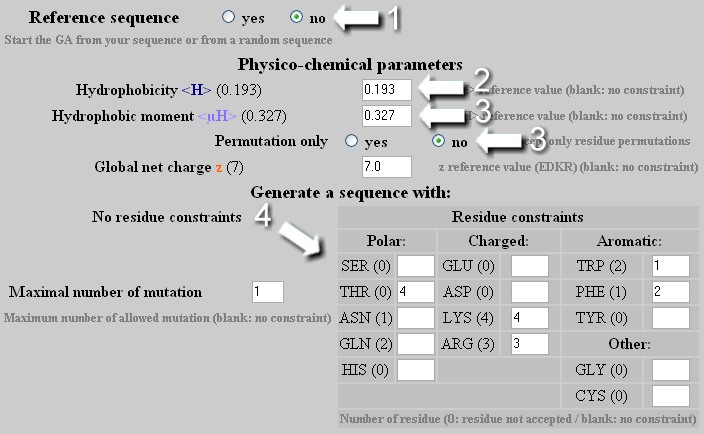
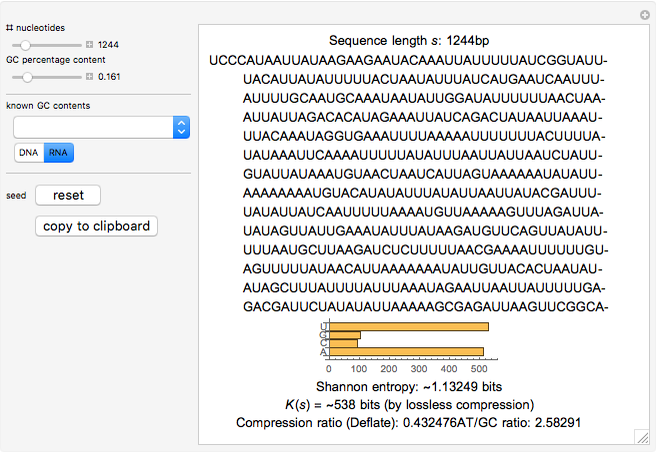
NullSeq: A Tool for Generating Random Coding Sequences with Desired Amino Acid and GC Contents | PLOS Computational Biology
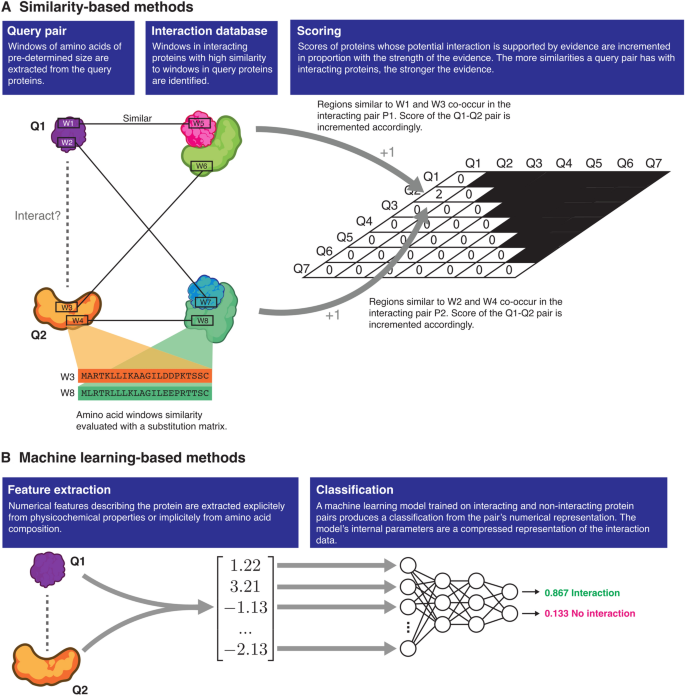
Assessing sequence-based protein–protein interaction predictors for use in therapeutic peptide engineering | Scientific Reports
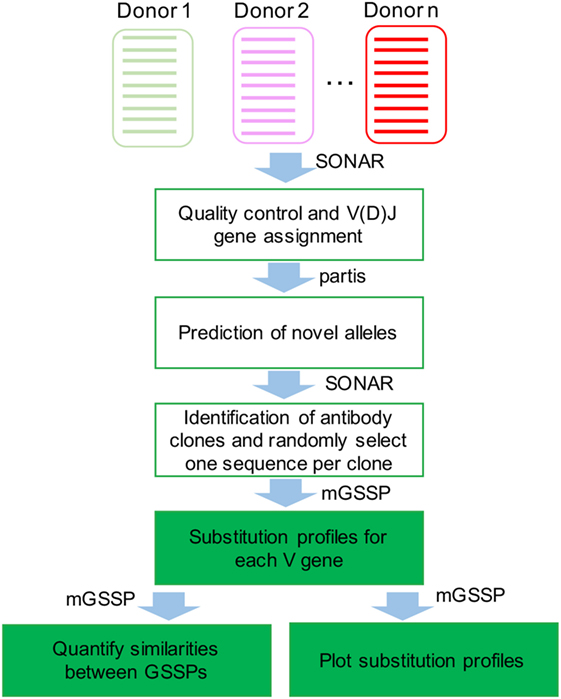
Frontiers | Gene-Specific Substitution Profiles Describe the Types and Frequencies of Amino Acid Changes during Antibody Somatic Hypermutation

IJMS | Free Full-Text | Intelligent De Novo Design of Novel Antimicrobial Peptides against Antibiotic-Resistant Bacteria Strains

CodSeqGen: A tool for generating synonymous coding sequences with desired GC-contents - ScienceDirect

Methods for enzyme library creation: Which one will you choose? - Alejaldre - 2021 - BioEssays - Wiley Online Library
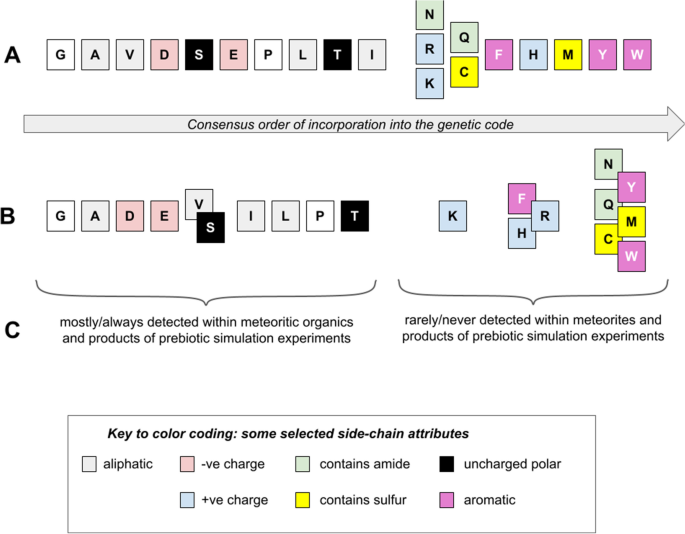
A Closer Look at Non-random Patterns Within Chemistry Space for a Smaller, Earlier Amino Acid Alphabet | SpringerLink



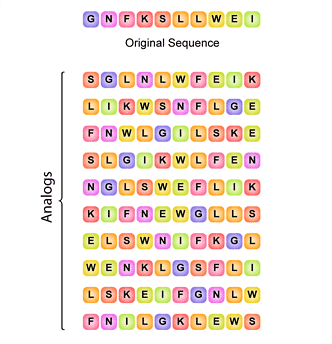

![PDF] Randomseq: python command–line random sequence generator | Semantic Scholar PDF] Randomseq: python command–line random sequence generator | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/517d4e899d52fd7536562b4ee597578c80f409e9/3-Figure1-1.png)