In Silico Processing of the Complete CRISPR‐Cas Spacer Space for Identification of PAM Sequences - Mendoza - 2018 - Biotechnology Journal - Wiley Online Library
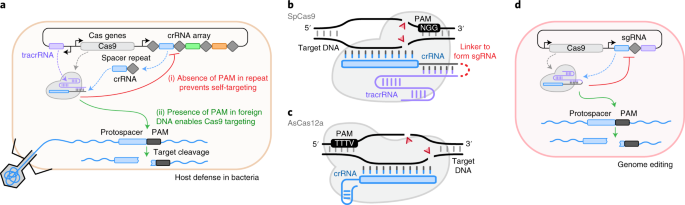
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols
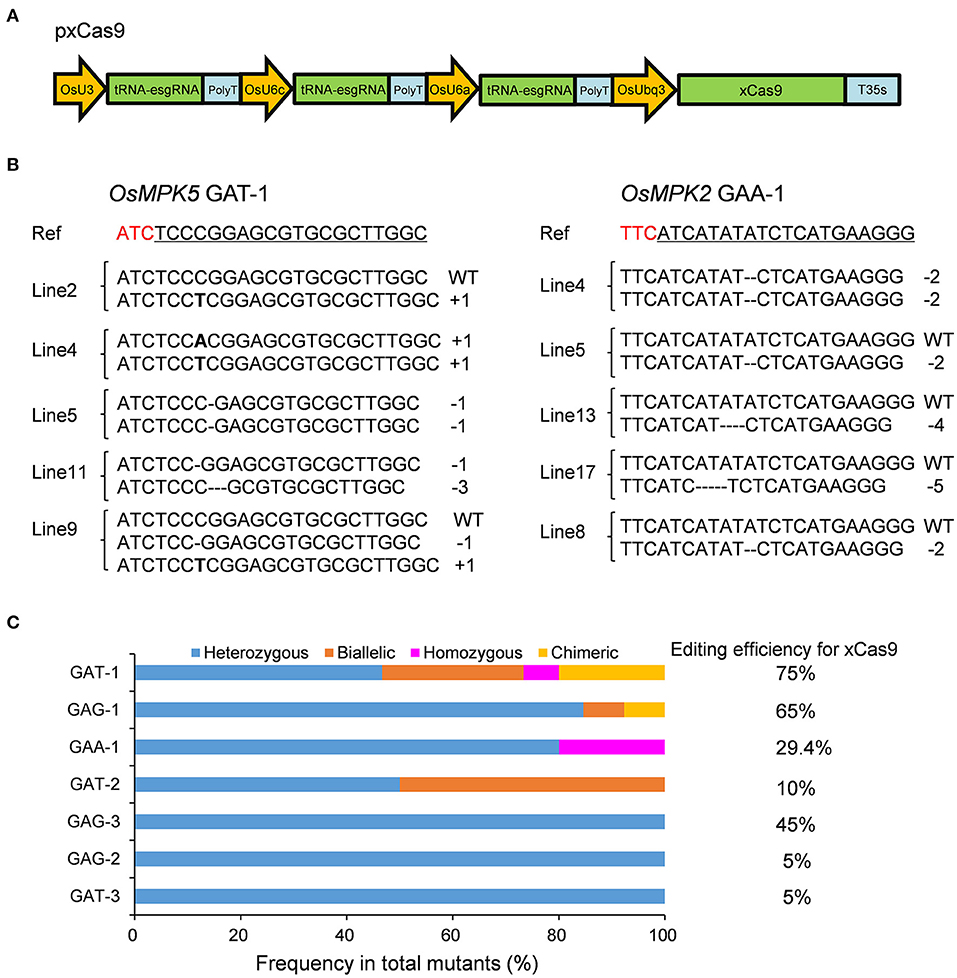
Frontiers | Genome Engineering in Plant Using an Efficient CRISPR-xCas9 Toolset With an Expanded PAM Compatibility