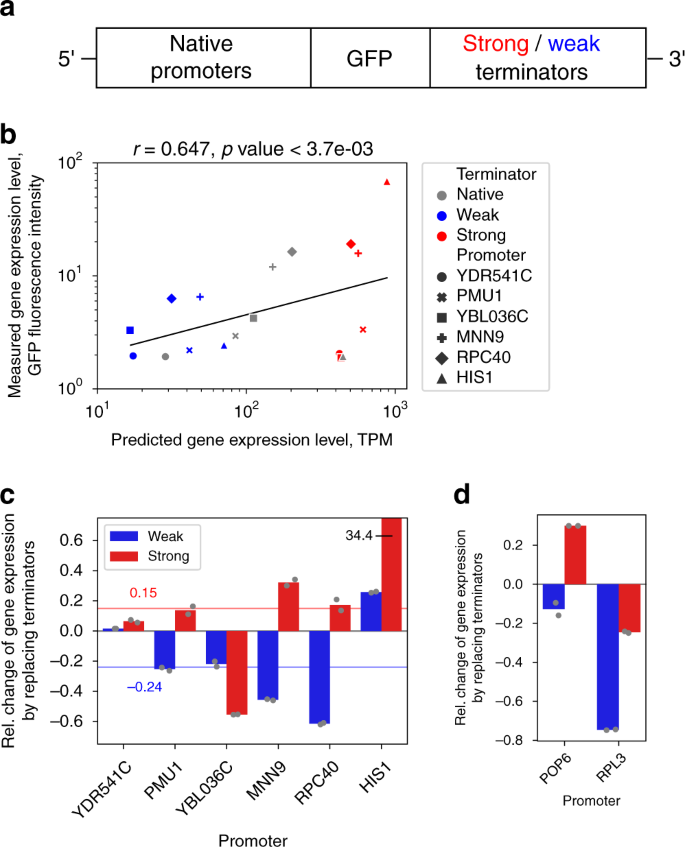
Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure | Nature Communications

Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure | Nature Communications
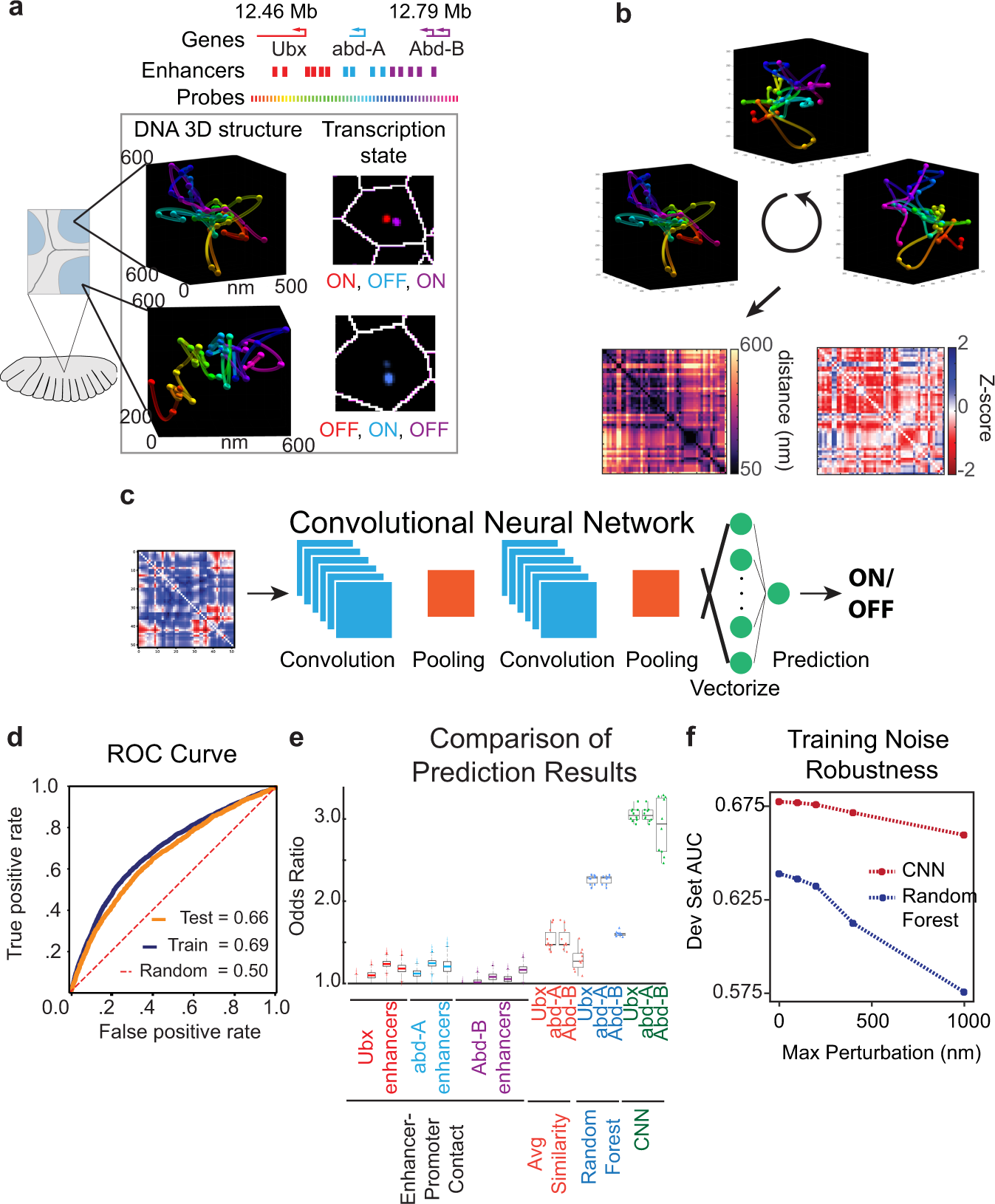
Deep learning connects DNA traces to transcription to reveal predictive features beyond enhancer–promoter contact | Nature Communications

Predicting mRNA Abundance Directly from Genomic Sequence Using Deep Convolutional Neural Networks - ScienceDirect

Predicting mRNA Abundance Directly from Genomic Sequence Using Deep Convolutional Neural Networks - ScienceDirect

Predicting mRNA Abundance Directly from Genomic Sequence Using Deep Convolutional Neural Networks - ScienceDirect
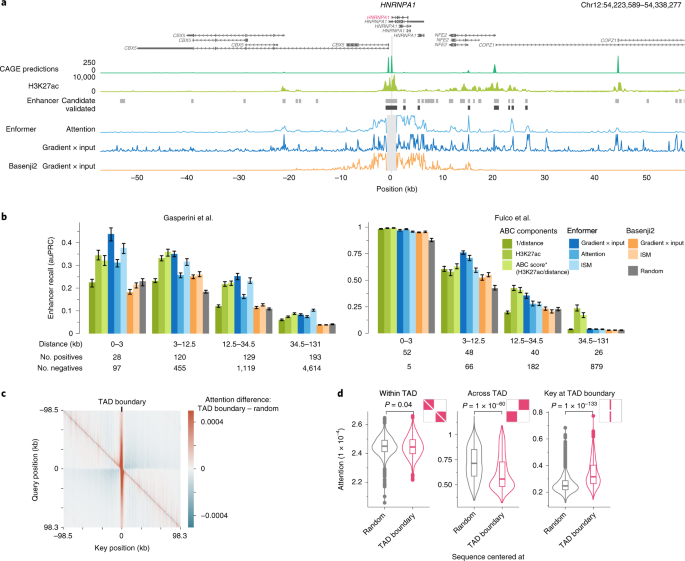
Effective gene expression prediction from sequence by integrating long-range interactions | Nature Methods
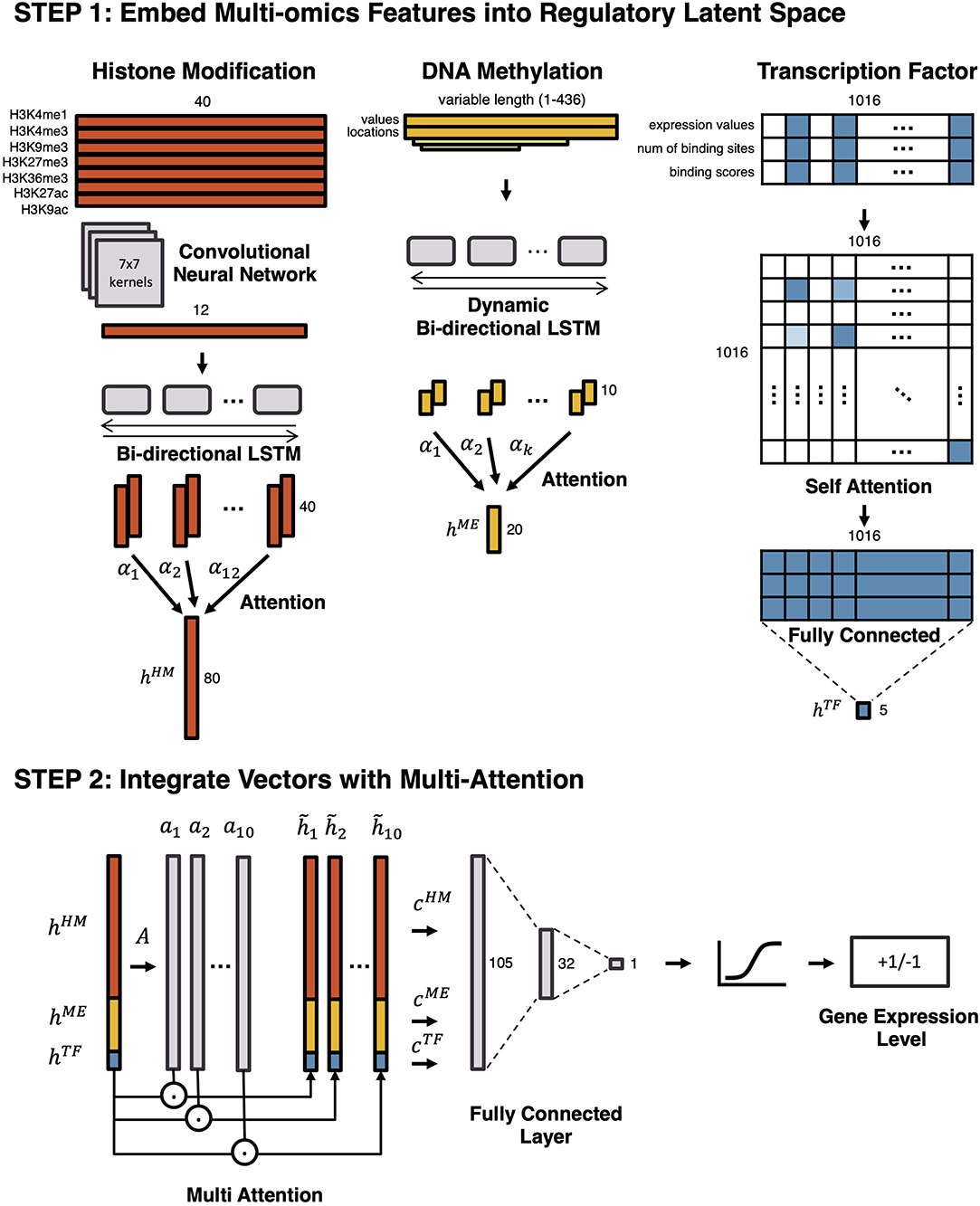
Frontiers | Learning Cell-Type-Specific Gene Regulation Mechanisms by Multi-Attention Based Deep Learning With Regulatory Latent Space

Effective gene expression prediction from sequence by integrating long-range interactions | Nature Methods