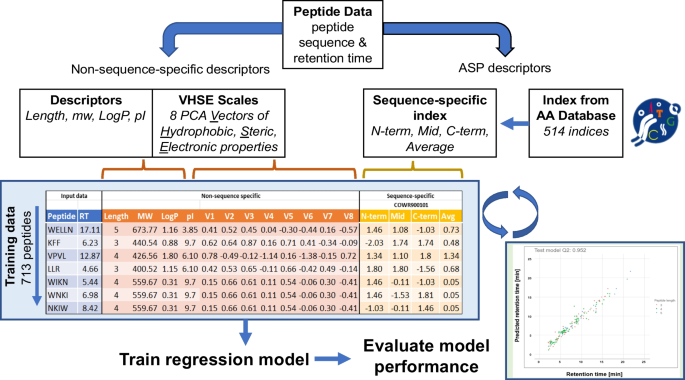
Improved LC–MS identification of short homologous peptides using sequence-specific retention time predictors | SpringerLink

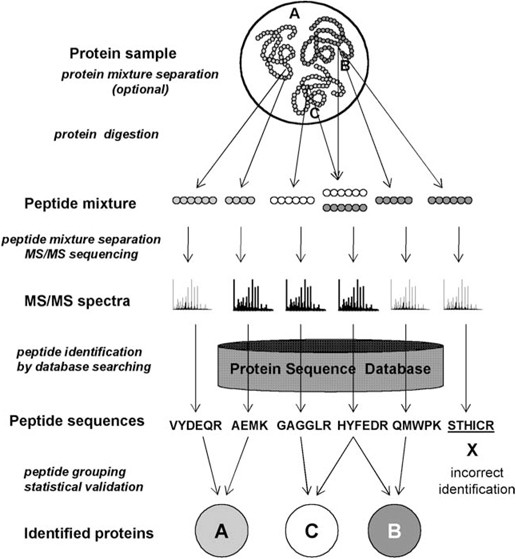
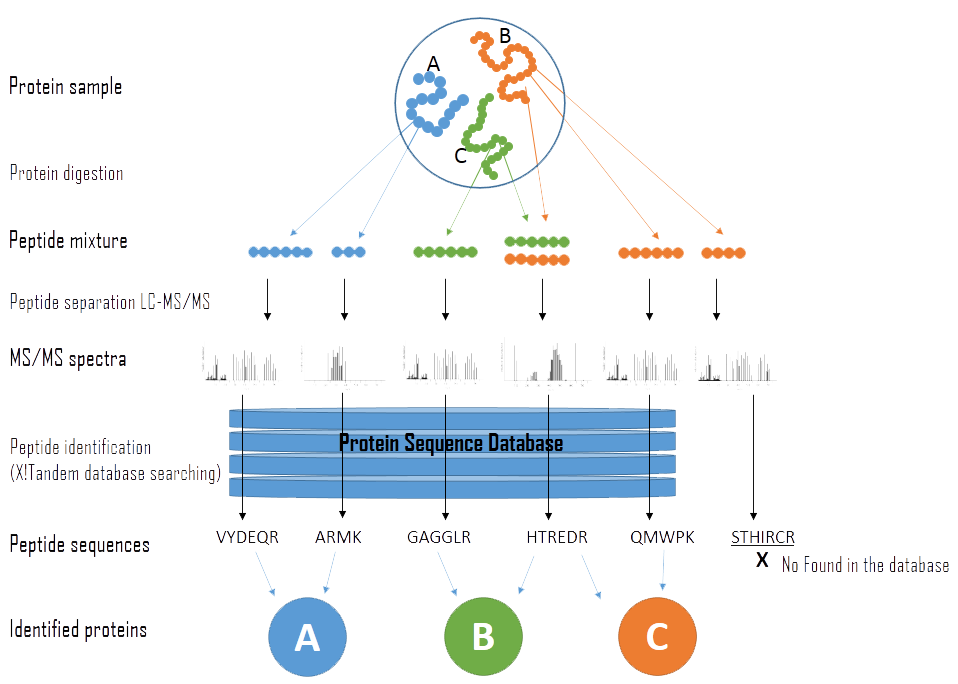
Protein Identification by Database Searching of Mass Spectrometry Data in the Teaching of Proteomics | Journal of Chemical Education
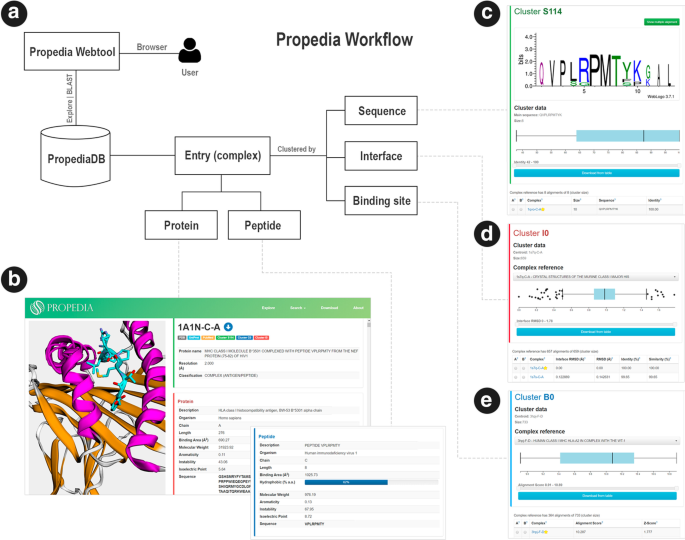
Propedia: a database for protein–peptide identification based on a hybrid clustering algorithm | BMC Bioinformatics | Full Text

Uncovering Thousands of New Peptides with Sequence-Mask-Search Hybrid De Novo Peptide Sequencing Framework - ScienceDirect

MaCPepDB: A Database to Quickly Access All Tryptic Peptides of the UniProtKB | Journal of Proteome Research
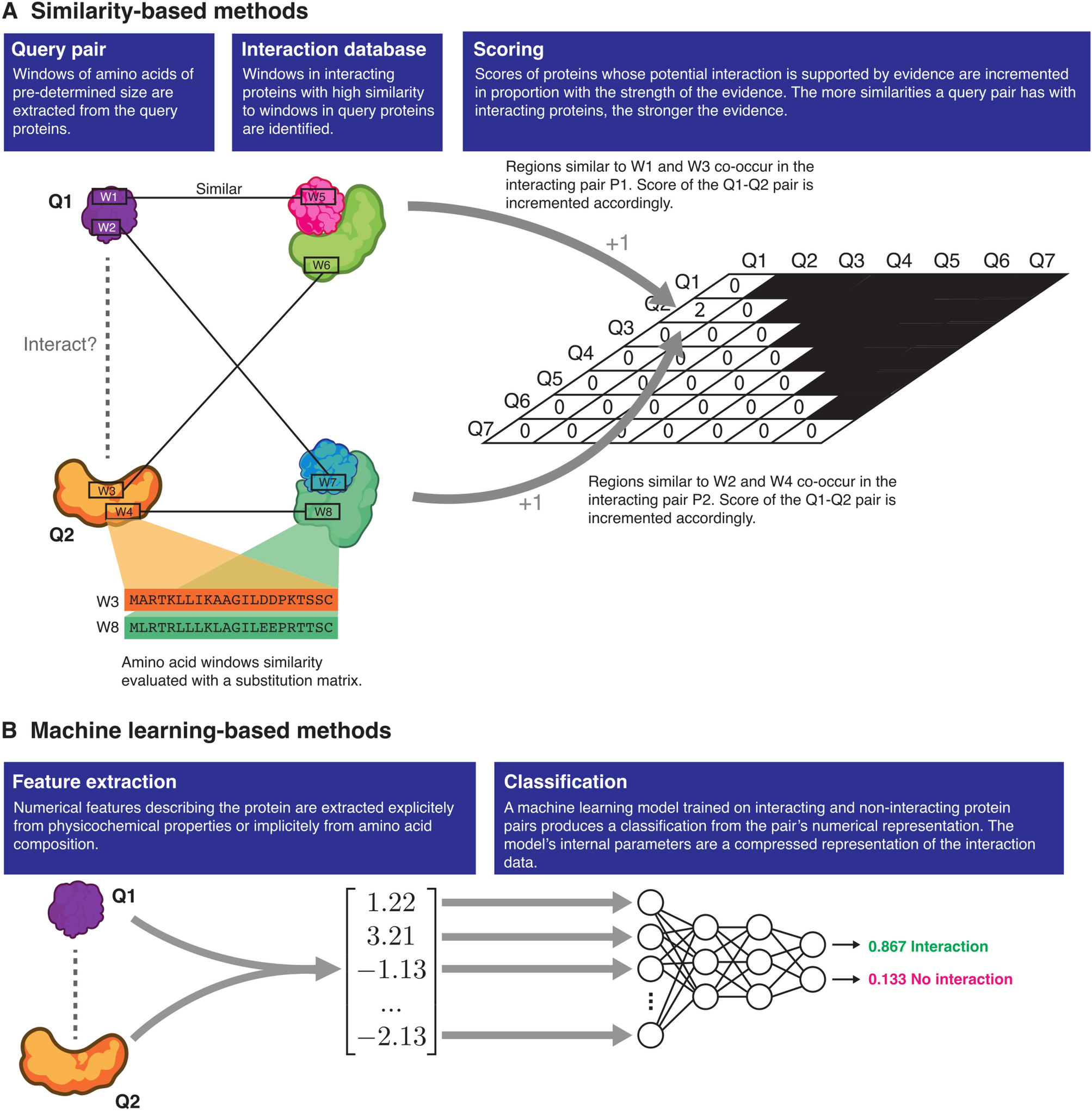
Assessing sequence-based protein–protein interaction predictors for use in therapeutic peptide engineering | Scientific Reports

PepBank - a database of peptides based on sequence text mining and public peptide data sources – topic of research paper in Biological sciences. Download scholarly article PDF and read for free

How To Use the Conserved Domain Database (CDD): identify amino acids involved in binding or catalysis


![Help [PIR - Protein Information Resource] Help [PIR - Protein Information Resource]](https://proteininformationresource.org/pirwww/images/creationpirsf.jpg)






![Peptide Match User Guide [PIR - Protein Information Resource] Peptide Match User Guide [PIR - Protein Information Resource]](https://research.bioinformatics.udel.edu/peptidematch/docs/images/singlePeptideMatchCPAnnotated.png)


