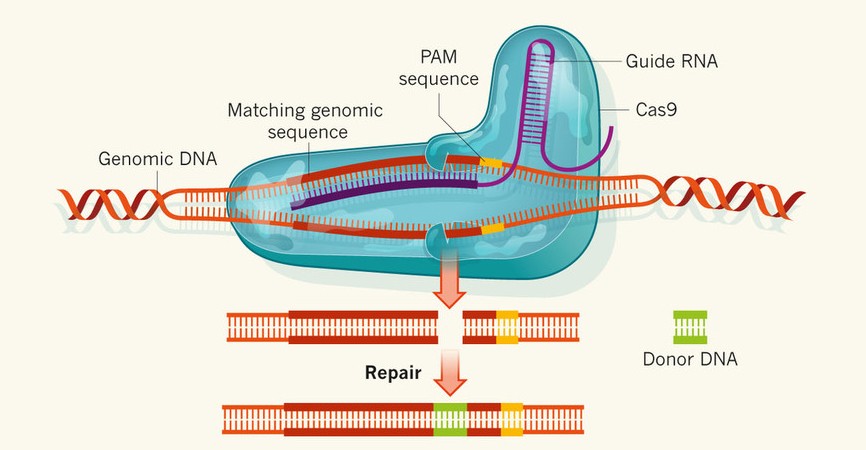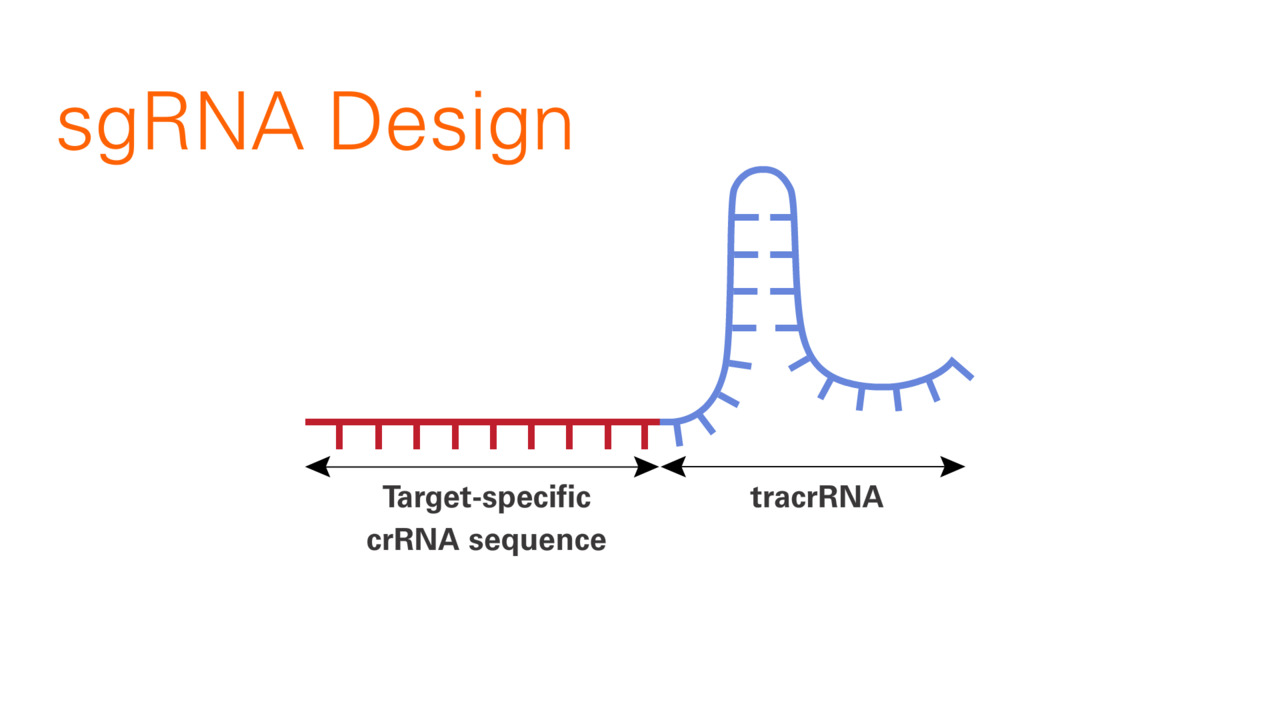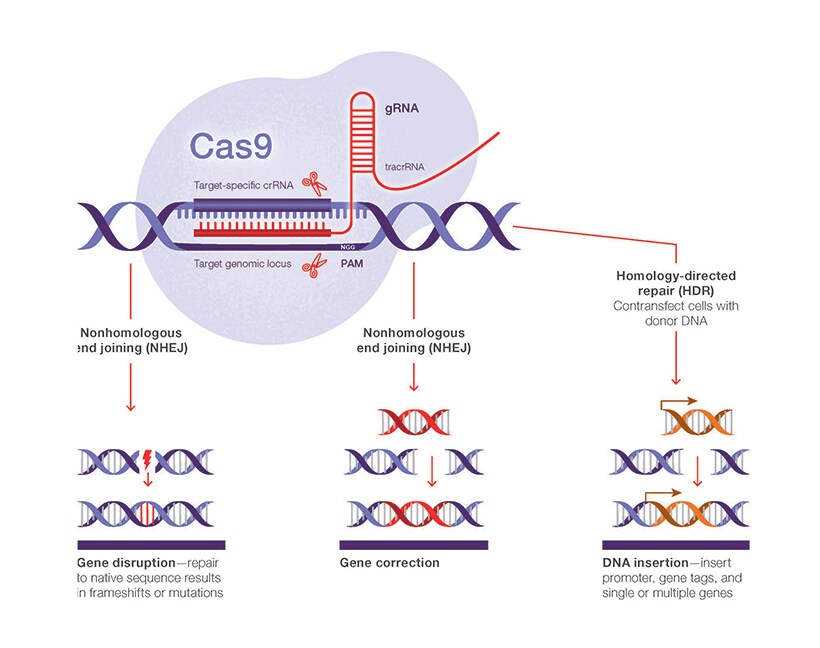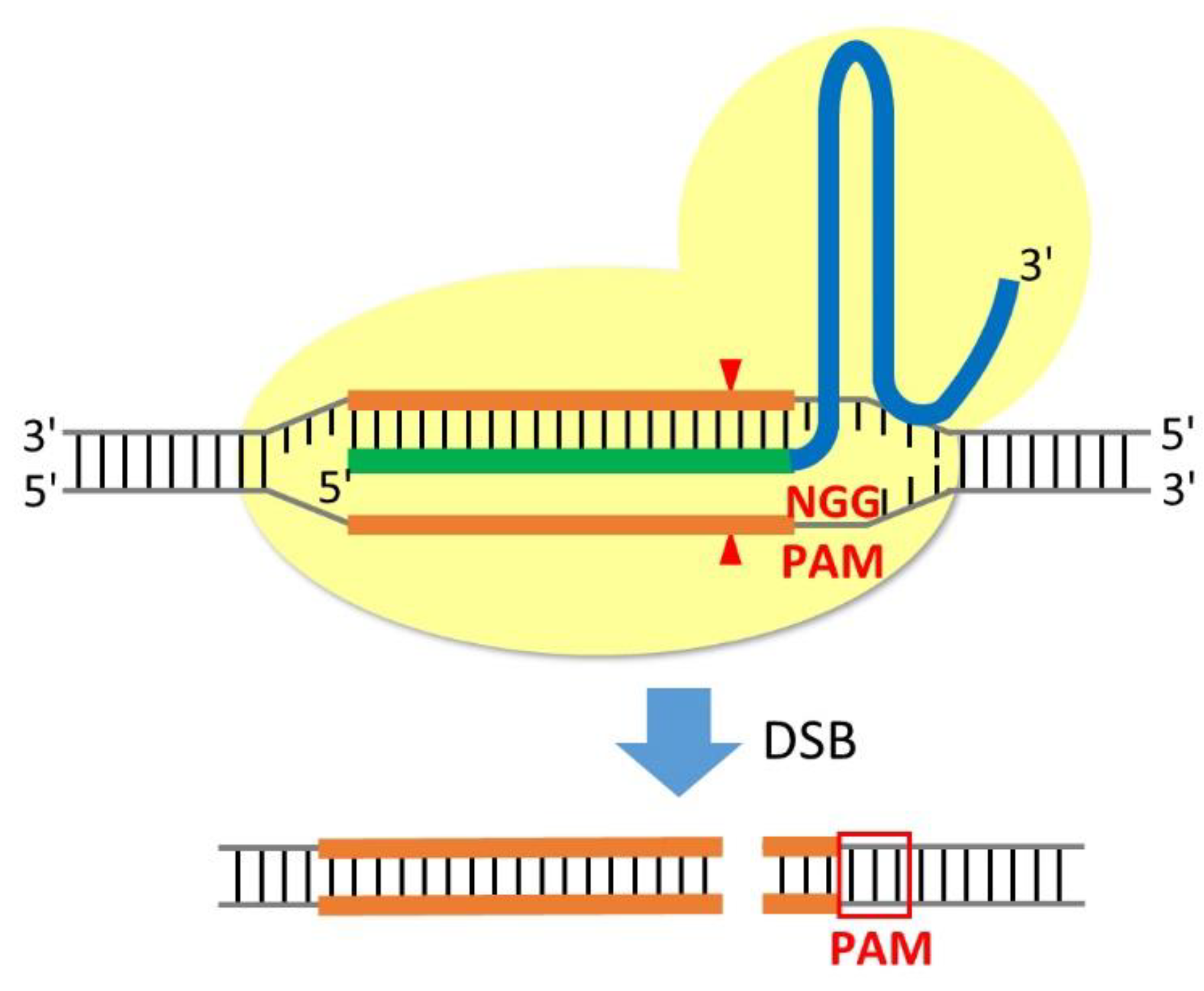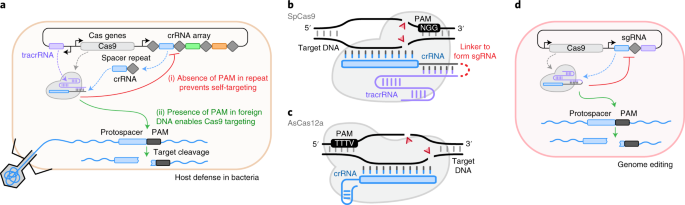
Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA | Nature Protocols
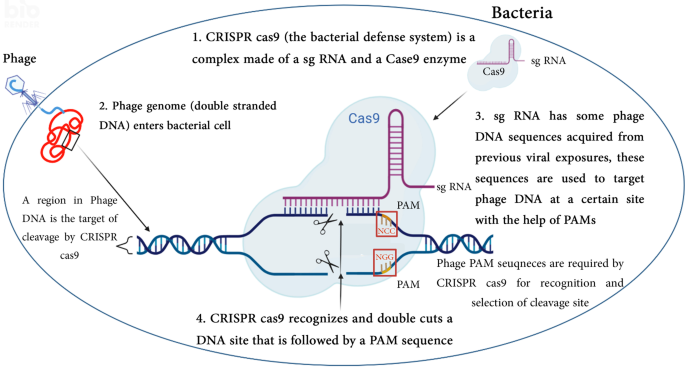
Bio-informatic analysis of CRISPR protospacer adjacent motifs (PAMs) in T4 genome | BMC Genomic Data | Full Text

A positive, growth-based PAM screen identifies noncanonical motifs recognized by the S. pyogenes Cas9 | Science Advances

Systematic analysis of Type I‐E Escherichia coli CRISPR‐Cas PAM sequences ability to promote interference and primed adaptation - Musharova - 2019 - Molecular Microbiology - Wiley Online Library

In Silico Processing of the Complete CRISPR‐Cas Spacer Space for Identification of PAM Sequences - Mendoza - 2018 - Biotechnology Journal - Wiley Online Library

Schematic for identification of PAM preferences by Cas9 cleavage in... | Download Scientific Diagram

Identification of functional PAM sequences for CRISPR-interference.... | Download Scientific Diagram