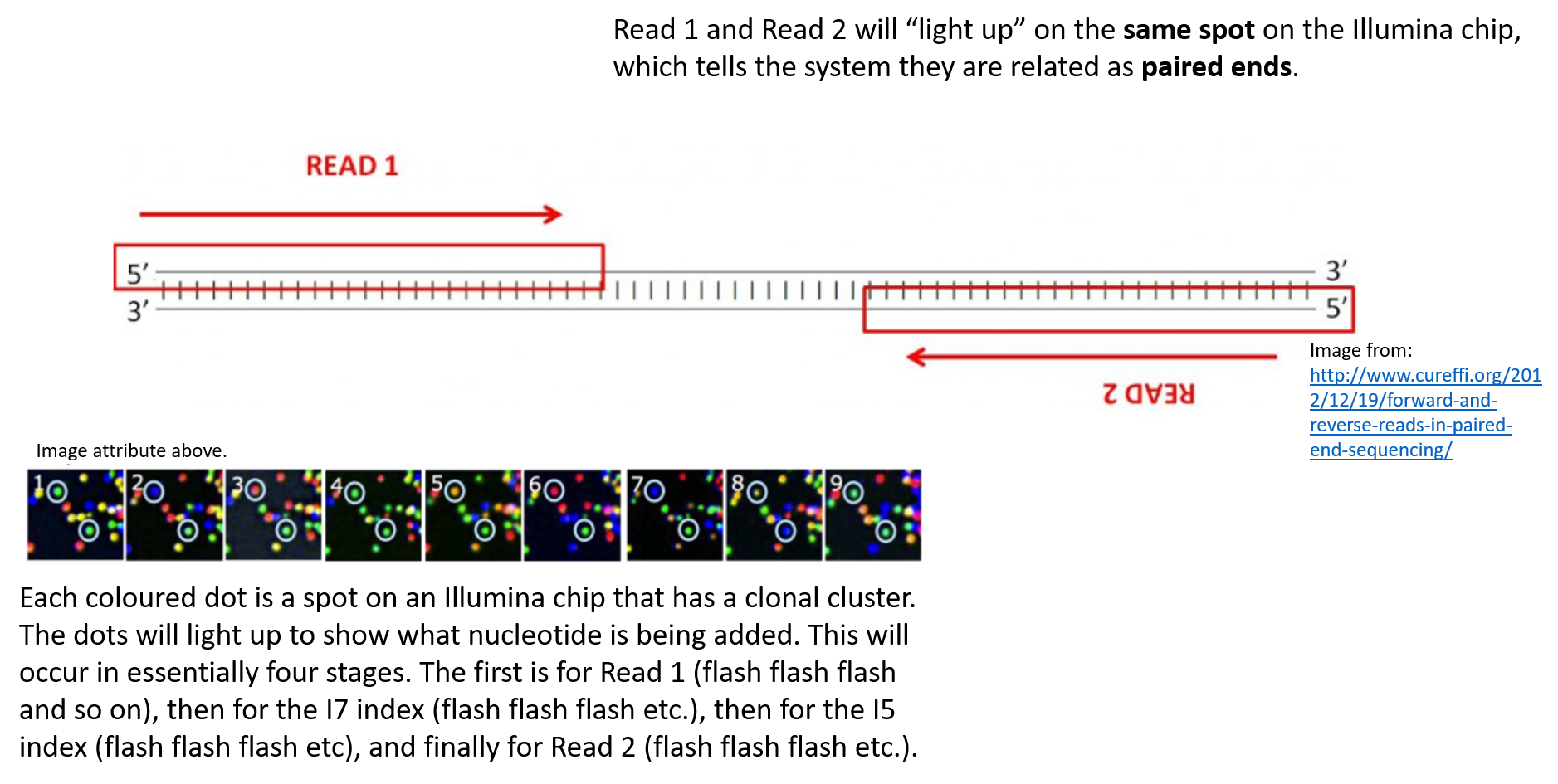
Illumina Sequencing (for Dummies) -An overview on how our samples are sequenced. – kscbioinformatics

Low-Cost, User-Friendly, All-Integrated Smartphone-Based Microplate Reader for Optical-Based Biological and Chemical Analyses | Analytical Chemistry
Decoding of Superimposed Traces Produced by Direct Sequencing of Heterozygous Indels | PLOS Computational Biology

Analysis of Mixed Sequencing Chromatograms and Its Application in Direct 16S rRNA Gene Sequencing of Polymicrobial Samples | Journal of Clinical Microbiology

Library adaptors with integrated reference controls improve the accuracy and reliability of nanopore sequencing | Nature Communications
![PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/957a53c4cb6cfbc76c655cef9acec295b0123949/4-Figure2-1.png)
PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar
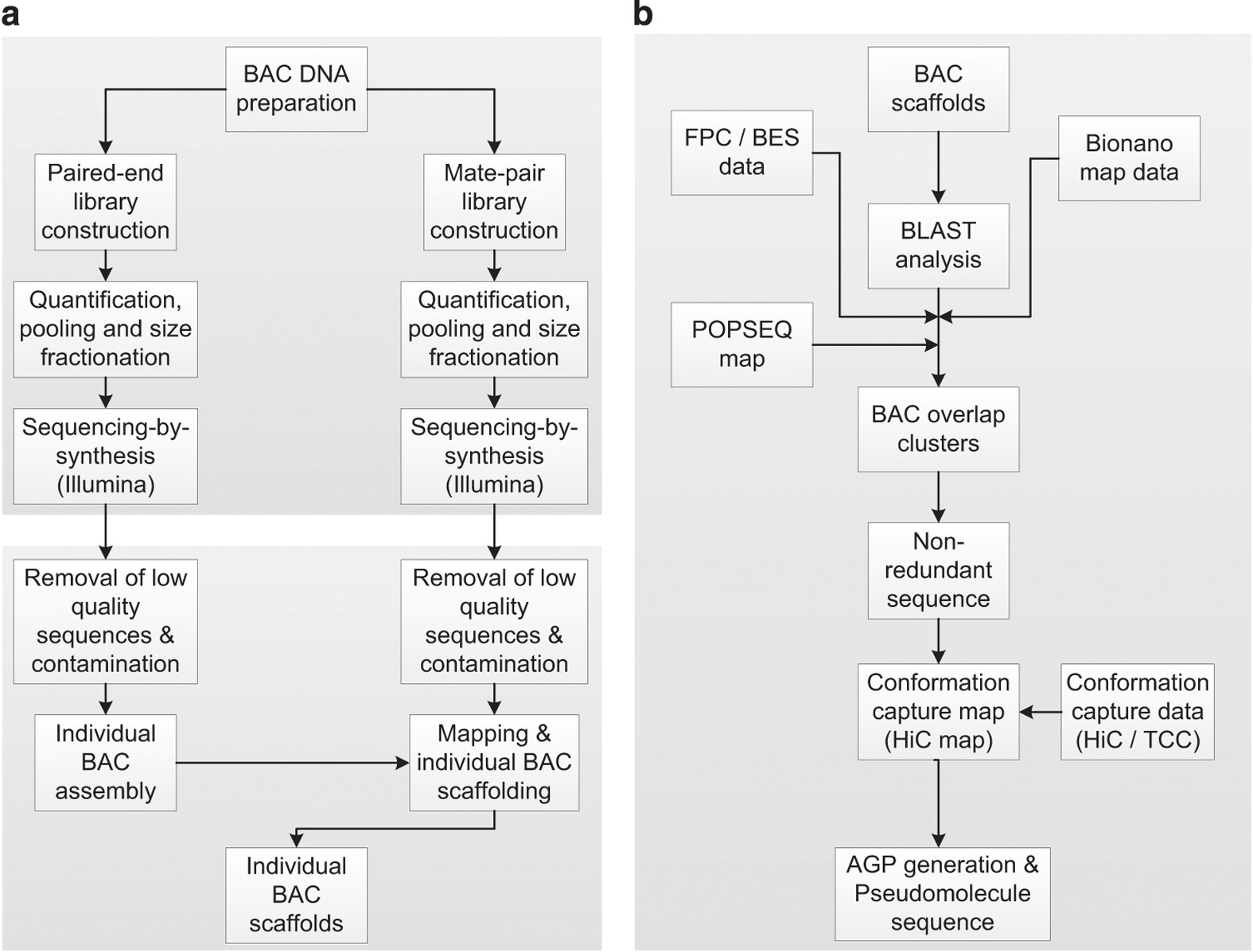
Construction of a map-based reference genome sequence for barley, Hordeum vulgare L. | Scientific Data
![PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/957a53c4cb6cfbc76c655cef9acec295b0123949/6-Figure4-1.png)
PDF] Mixed Sequence Reader: A Program for Analyzing DNA Sequences with Heterozygous Base Calling | Semantic Scholar