siRNA Seed Region Is Divided into Two Functionally Different Domains in RNA Interference in Response to 2′-OMe Modifications | ACS Omega

Application of the targeted sequencing approach reveals the single nucleotide polymorphism (SNP) repertoire in microRNA genes in the pig genome | Scientific Reports

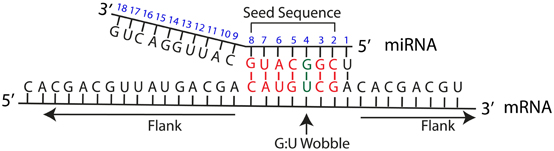
MicroRNA 3′-compensatory pairing occurs through two binding modes, with affinity shaped by nucleotide identity and position | eLife

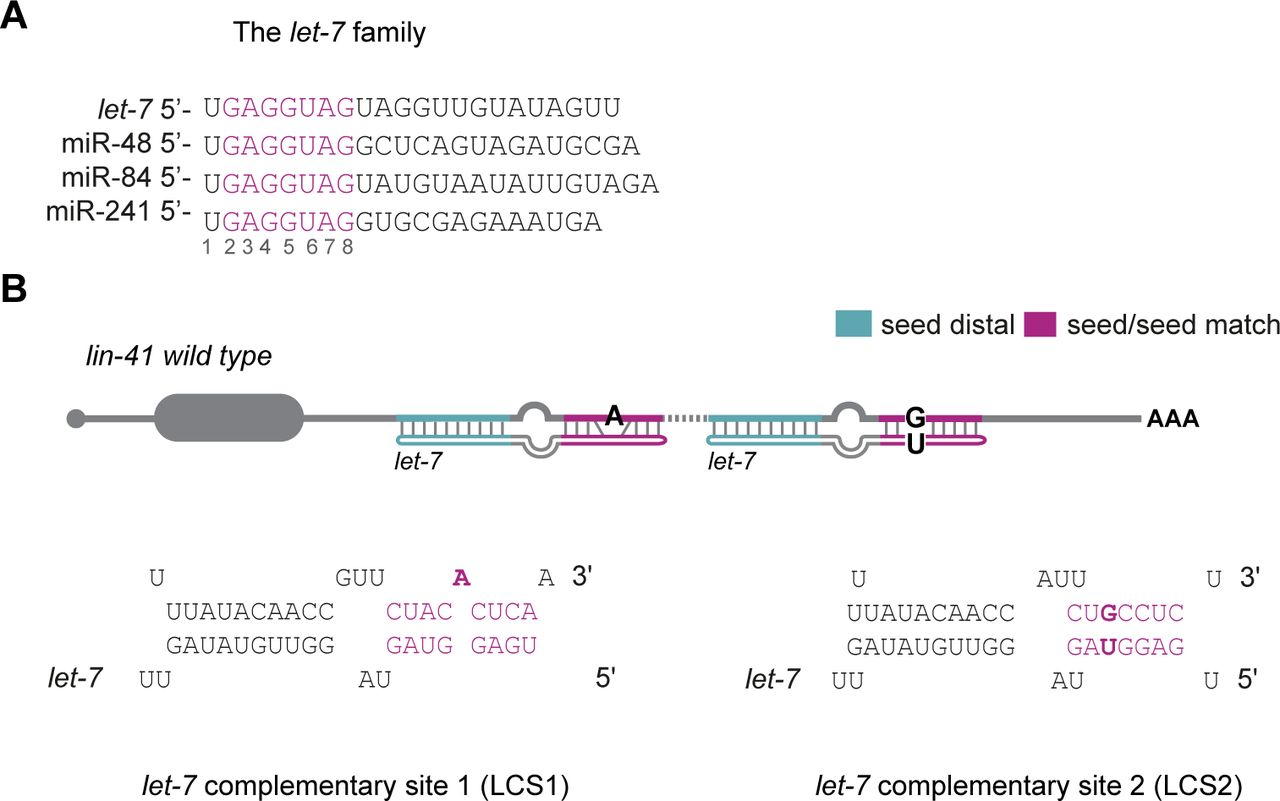
Critical contribution of 3′ non-seed base pairing to the in vivo function of the evolutionarily conserved let-7a microRNA - ScienceDirect
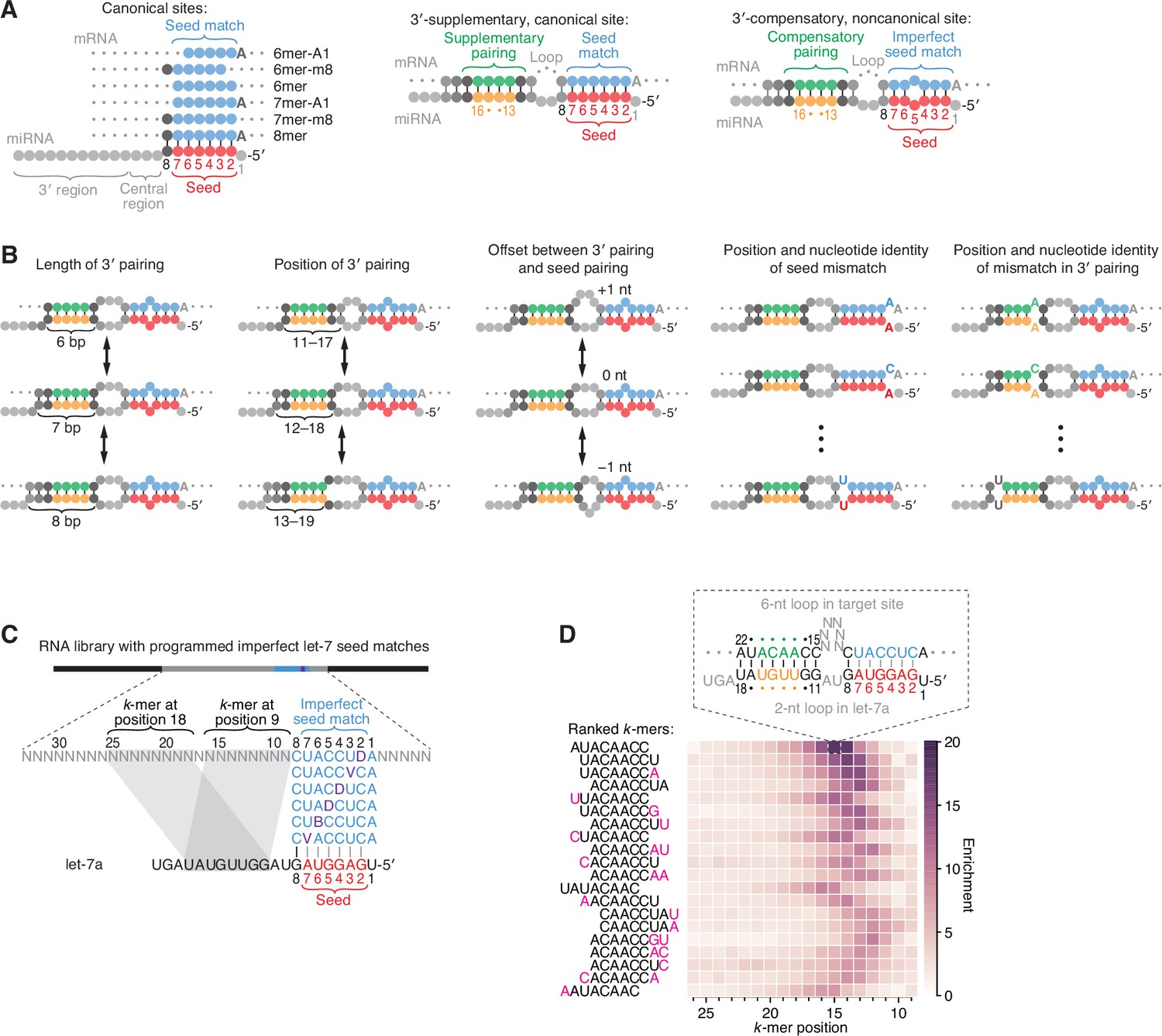
MicroRNA 3′-compensatory pairing occurs through two binding modes, with affinity shaped by nucleotide identity and position | eLife
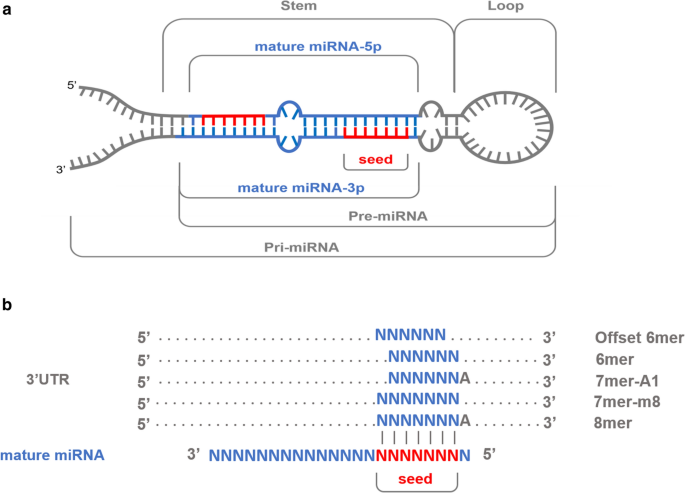
Variability in porcine microRNA genes and its association with mRNA expression and lipid phenotypes | Genetics Selection Evolution | Full Text

Beyond the seed: structural basis for supplementary microRNA targeting by human Argonaute2 | The EMBO Journal
Interplay of target site architecture and miRNA abundance determine miRNA activity and specificity | bioRxiv