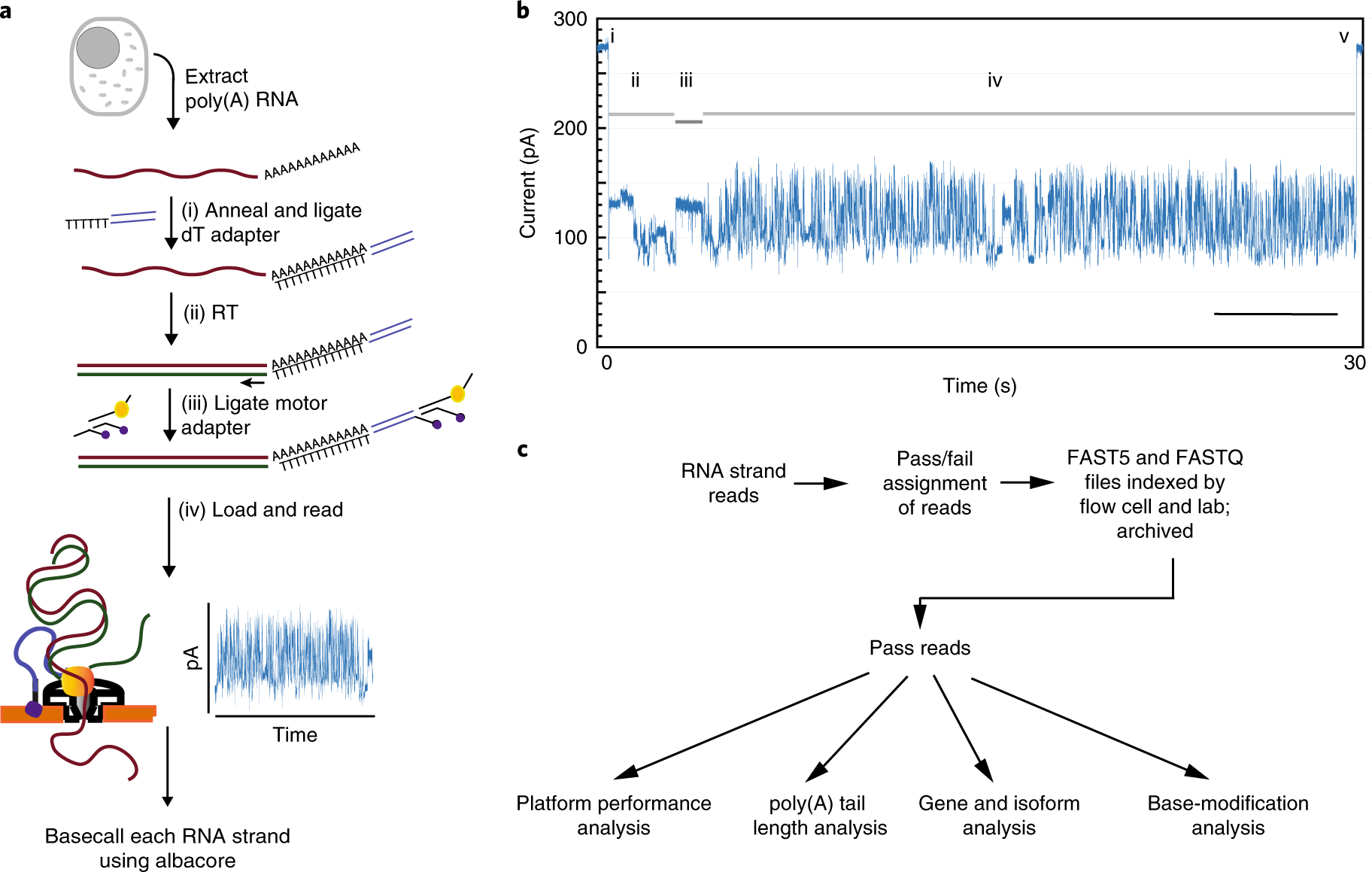
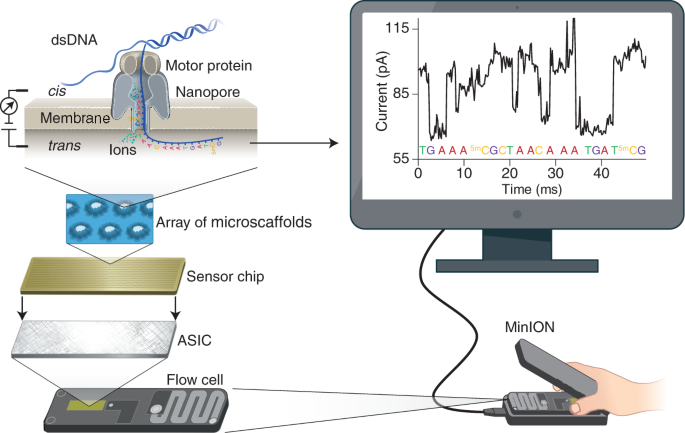
Analysis of Transcriptome and Epitranscriptome in Plants Using PacBio Iso- Seq and Nanopore-Based Direct RNA Sequencing | Semantic Scholar

Direct RNA-seq (a) Library preparation method for direct RNA-seq. (b)... | Download Scientific Diagram
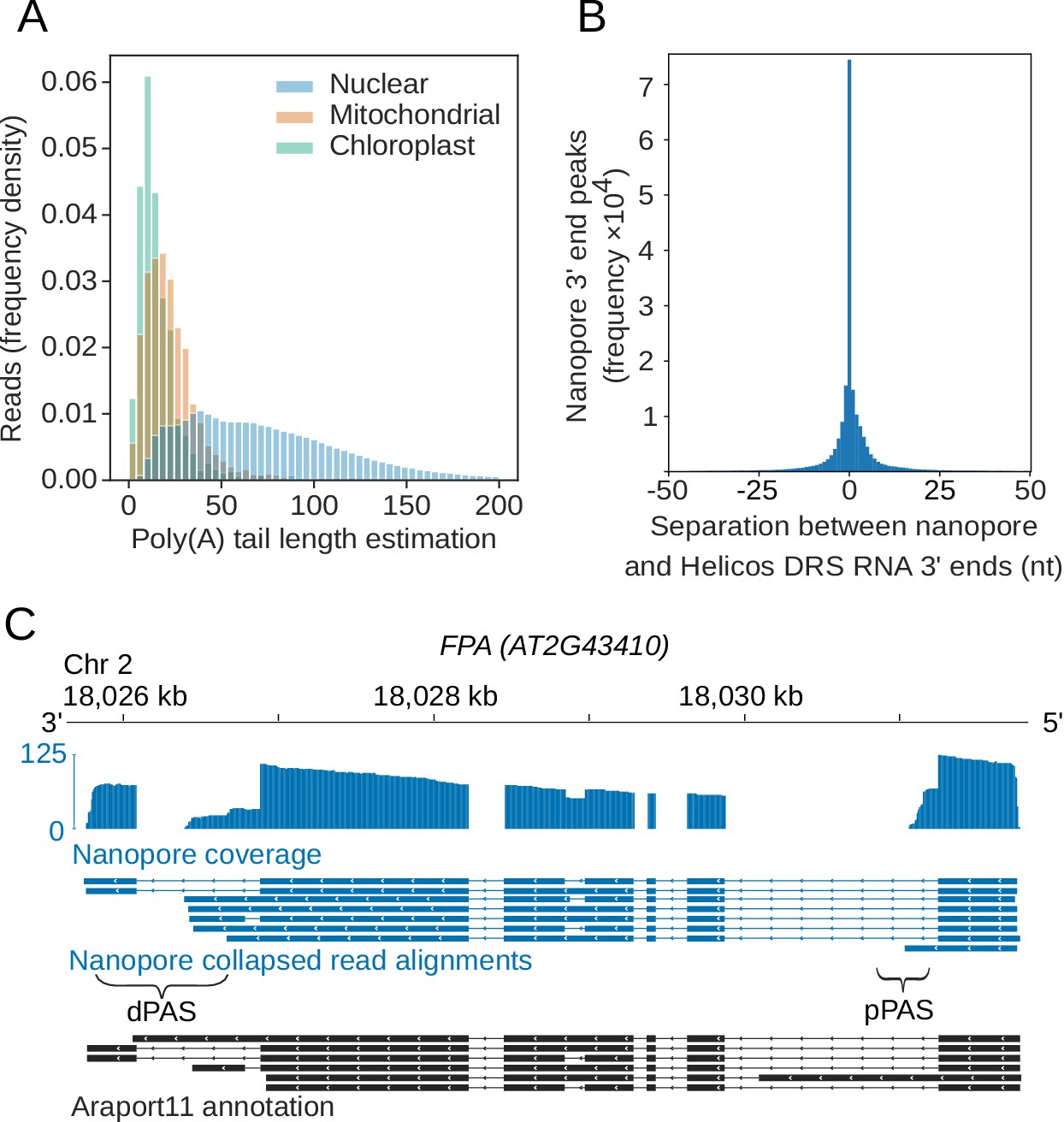
Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m6A modification | eLife
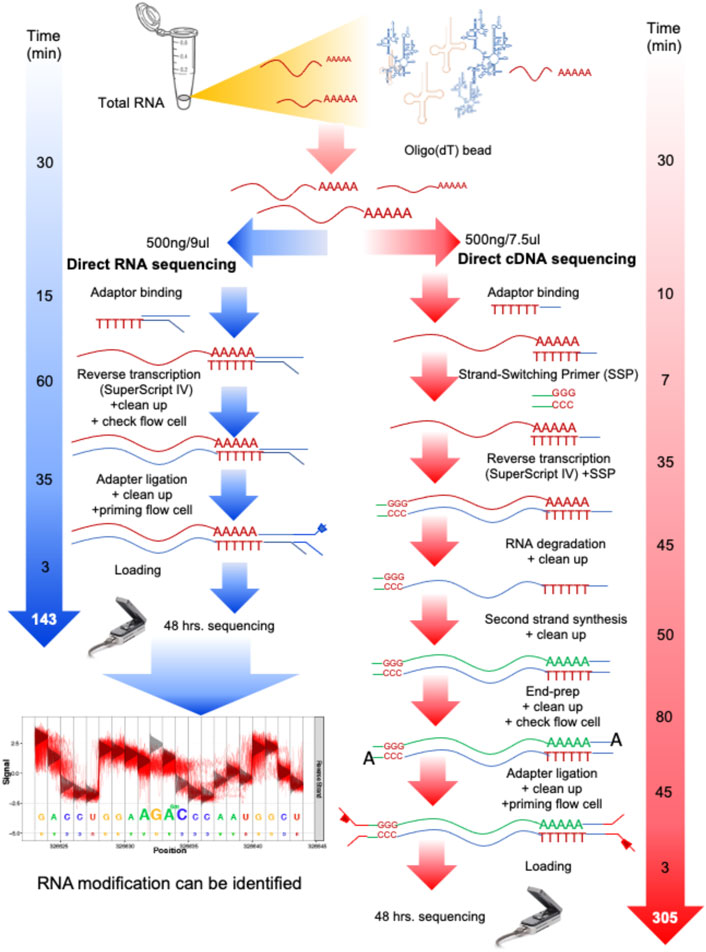
Frontiers | Native RNA or cDNA Sequencing for Transcriptomic Analysis: A Case Study on Saccharomyces cerevisiae
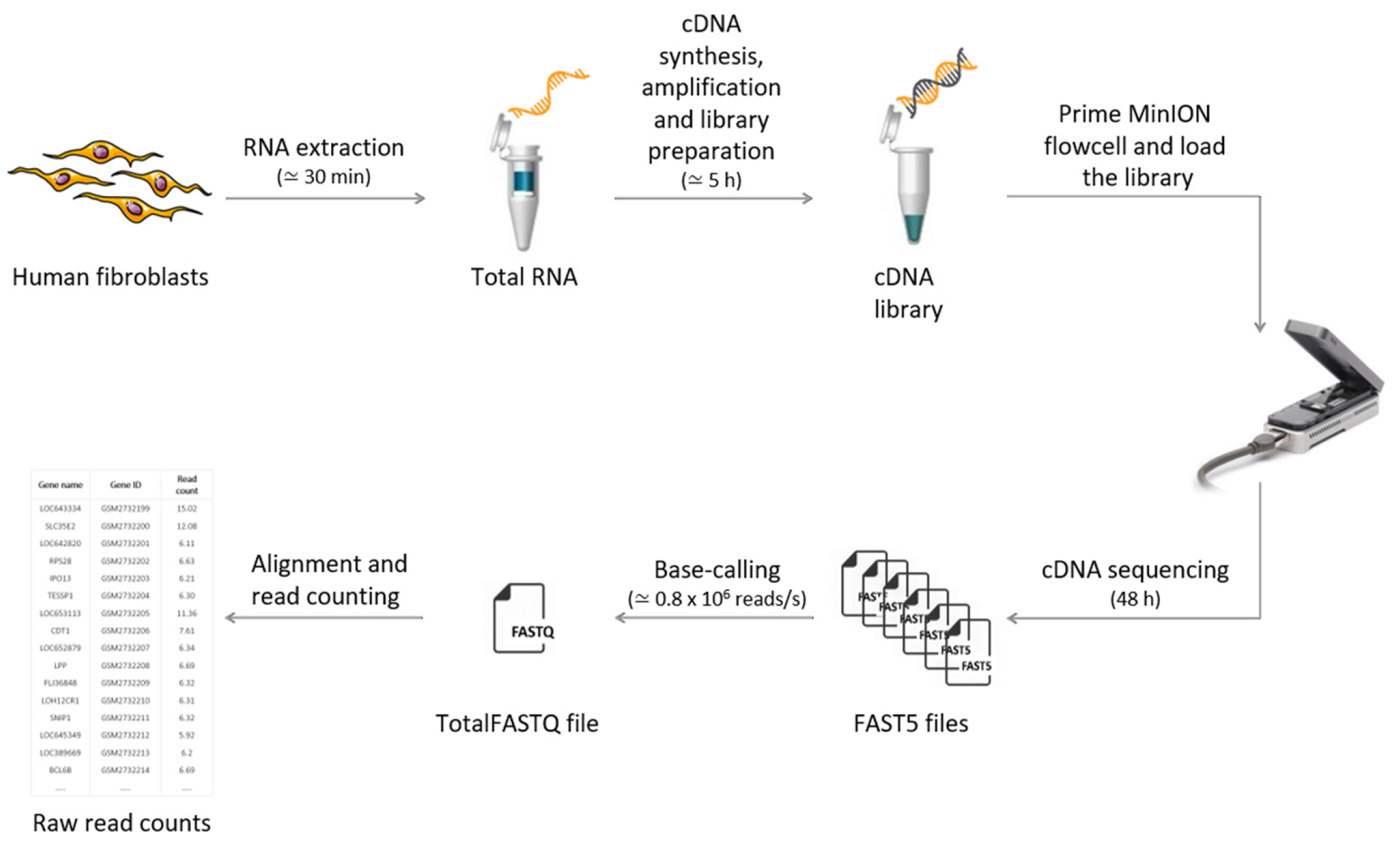
IJMS | Free Full-Text | Evaluation of Oxford Nanopore MinION RNA-Seq Performance for Human Primary Cells
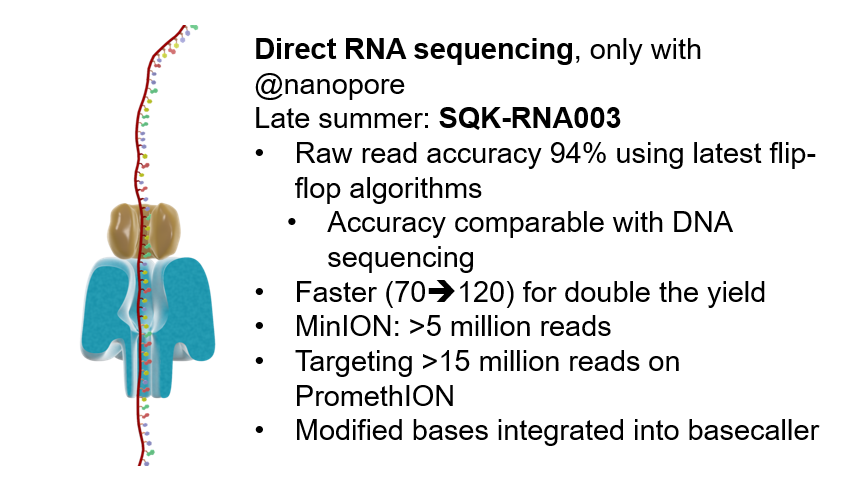

Oxford Nanopore on Twitter: "AH: Direct RNA with @nanopore will get closer to mainstream method this year #nanoporeconf https://t.co/3LrUy8fGog" / Twitter

Nanopore-based native RNA sequencing provides insights into prokaryotic transcription, operon structures, rRNA maturation and modifications | bioRxiv

Applications and potentials of nanopore sequencing in the (epi)genome and (epi)transcriptome era - ScienceDirect
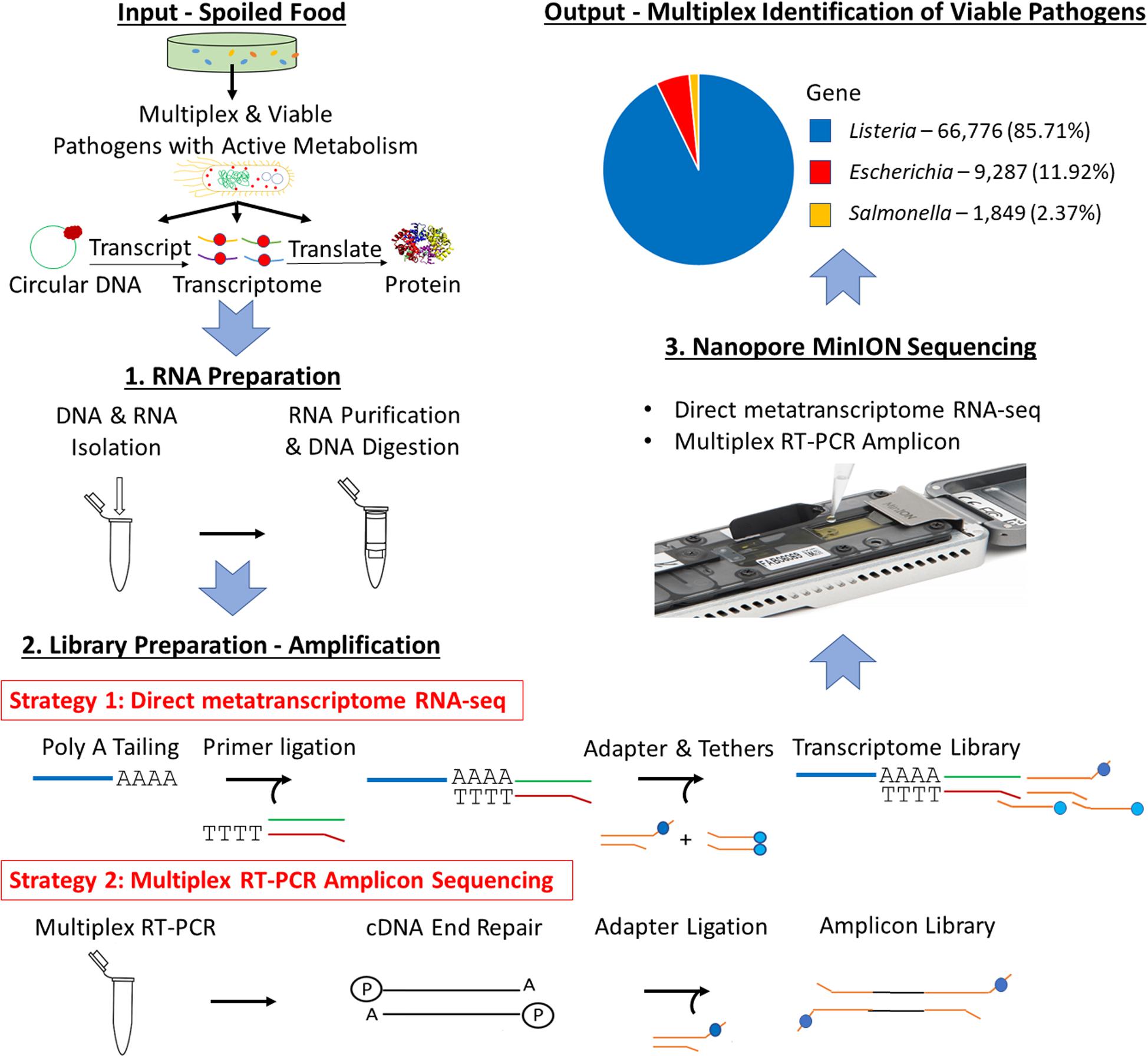
Frontiers | Direct Metatranscriptome RNA-seq and Multiplex RT-PCR Amplicon Sequencing on Nanopore MinION – Promising Strategies for Multiplex Identification of Viable Pathogens in Food

Improved nanopore direct RNA sequencing of cardiac myocyte samples by selective mt-RNA depletion - Journal of Molecular and Cellular Cardiology

Recent advances in the detection of base modifications using the Nanopore sequencer | Journal of Human Genetics

Frontiers | MasterOfPores: A Workflow for the Analysis of Oxford Nanopore Direct RNA Sequencing Datasets