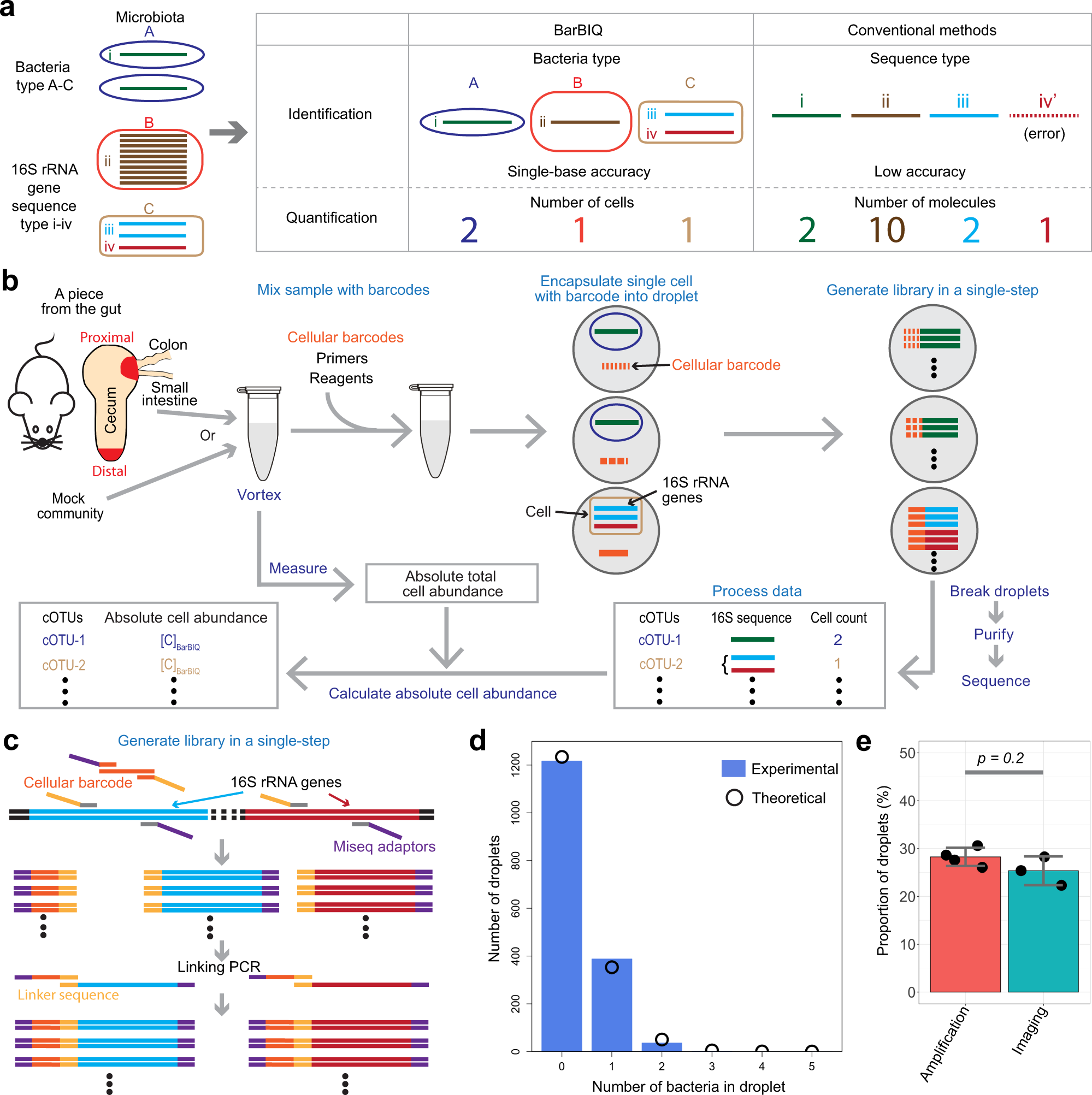
High-throughput identification and quantification of single bacterial cells in the microbiota | Nature Communications
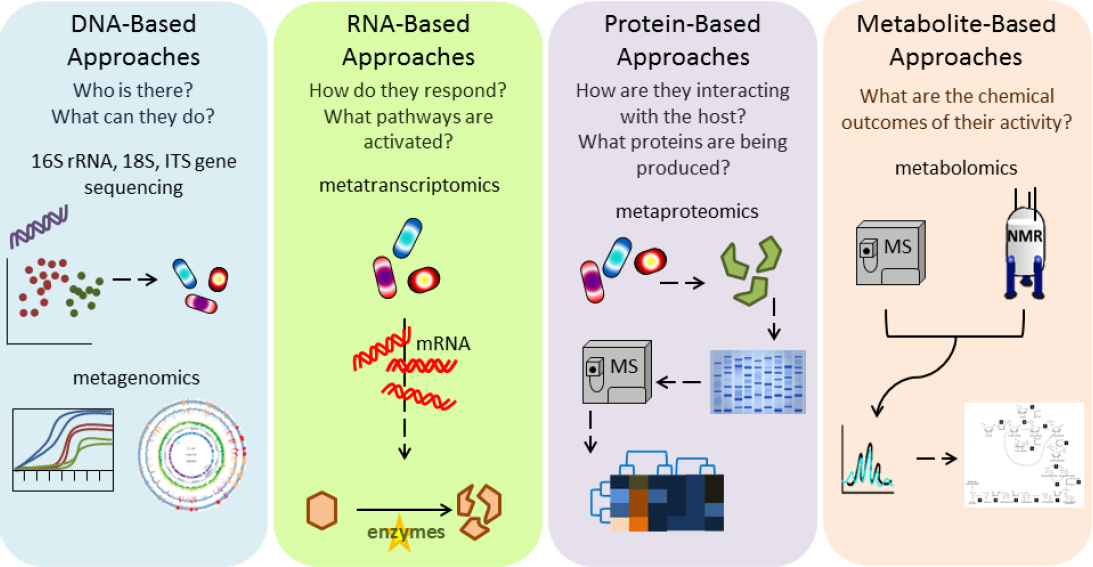
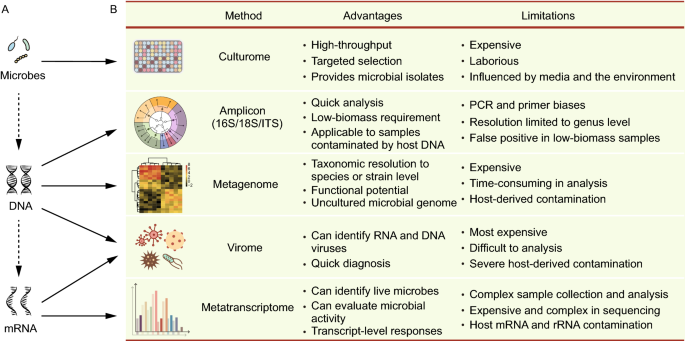
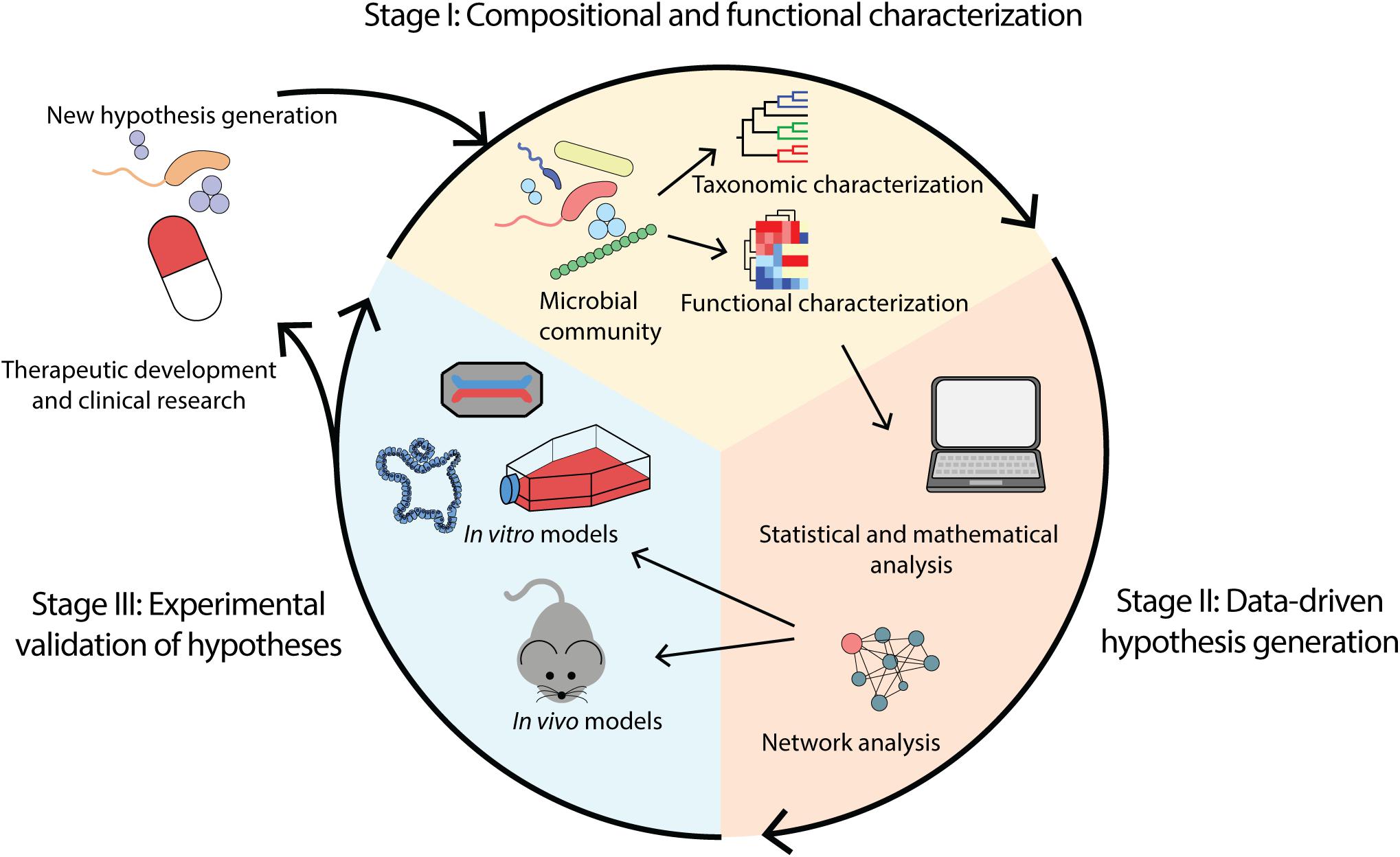
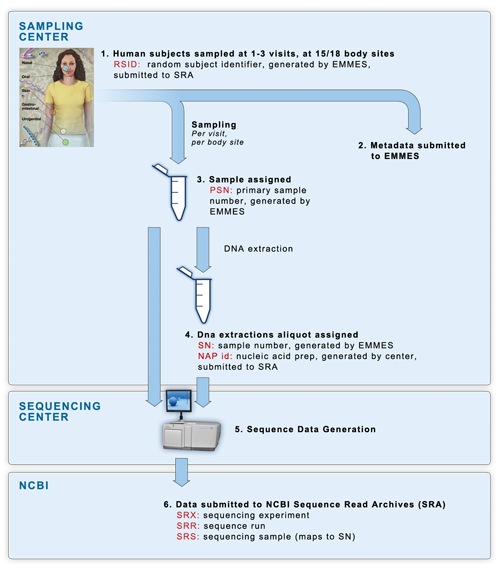
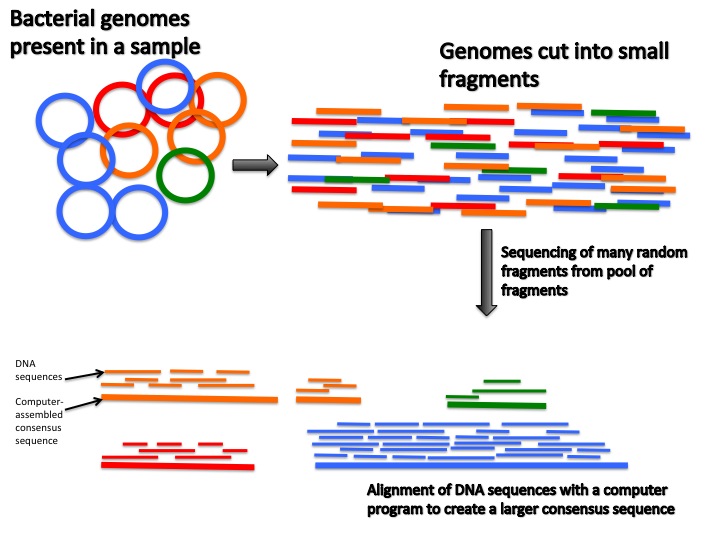
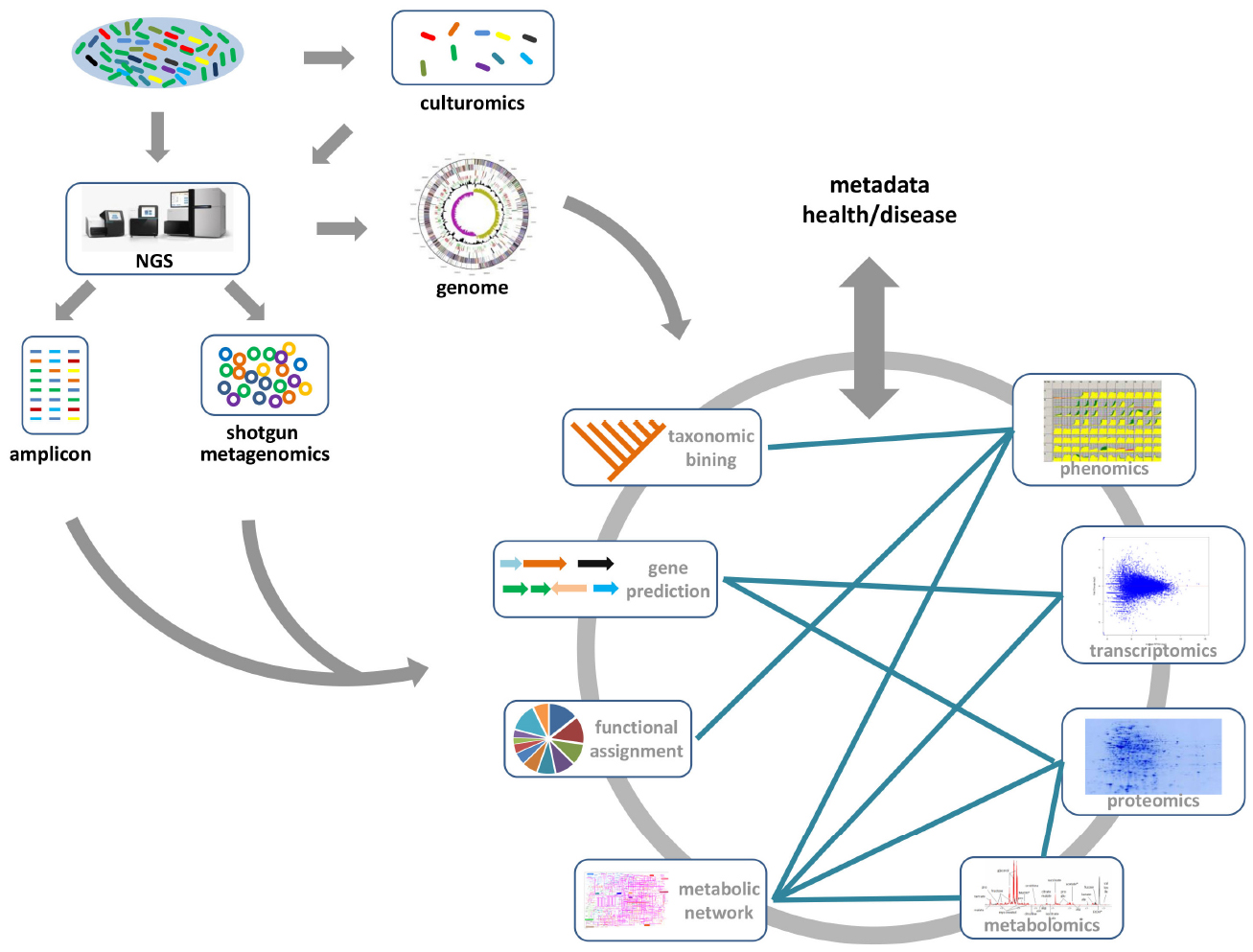
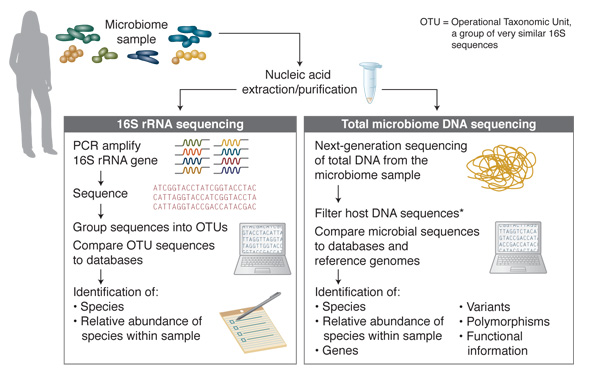
4 Current Methods for Studying the Human Microbiome | Environmental Chemicals, the Human Microbiome, and Health Risk: A Research Strategy |The National Academies Press

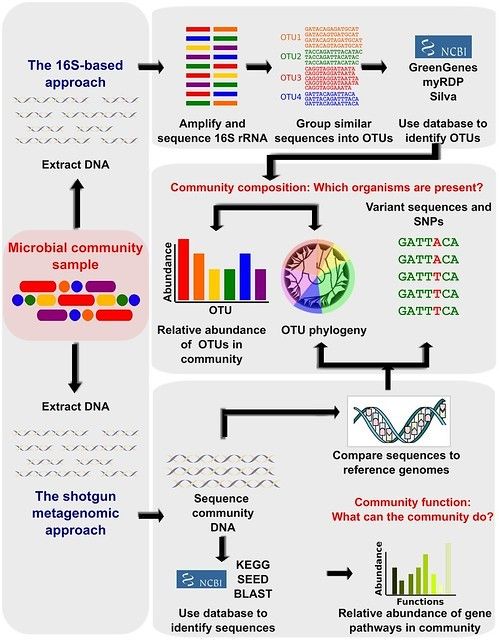
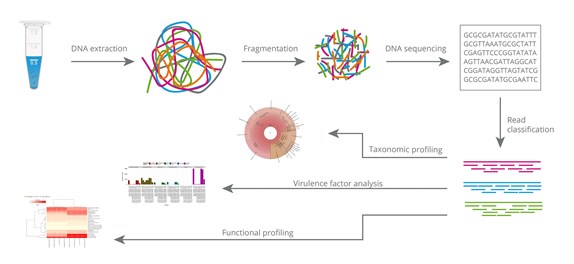
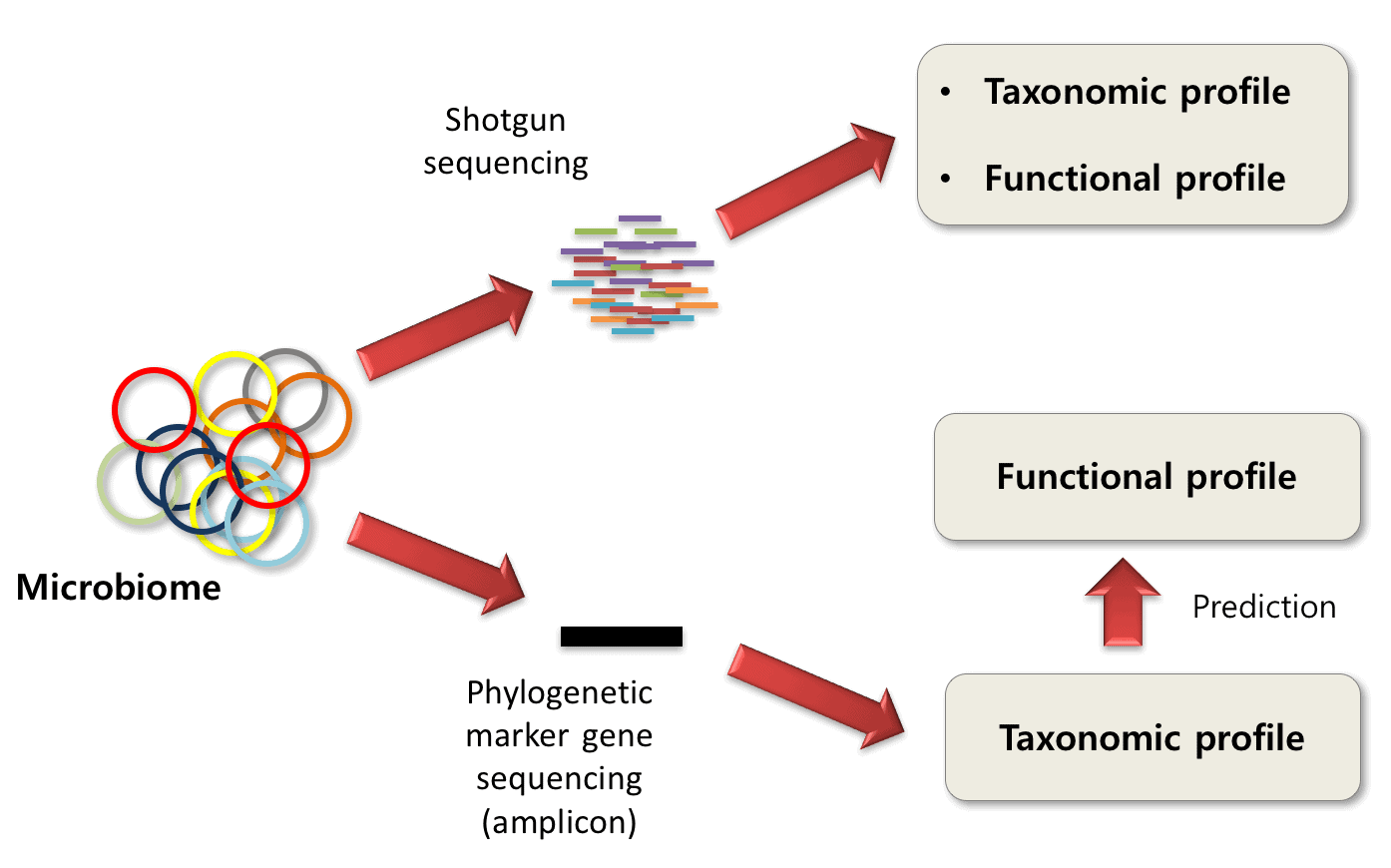
Multicenter assessment of microbial community profiling using 16S rRNA gene sequencing and shotgun metagenomic sequencing - ScienceDirect

Comprehensive 16S rRNA and metagenomic data from the gut microbiome of aging and rejuvenation mouse models | Scientific Data

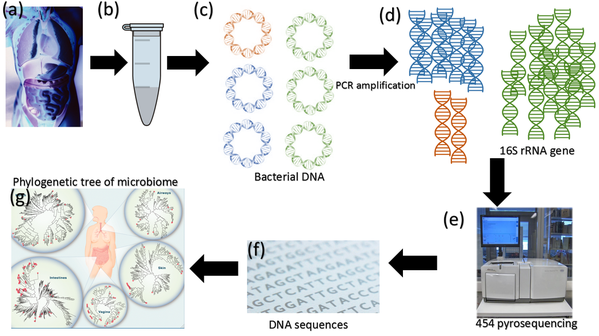
Impact of DNA extraction method and targeted 16S-rRNA hypervariable region on oral microbiota profiling | Scientific Reports
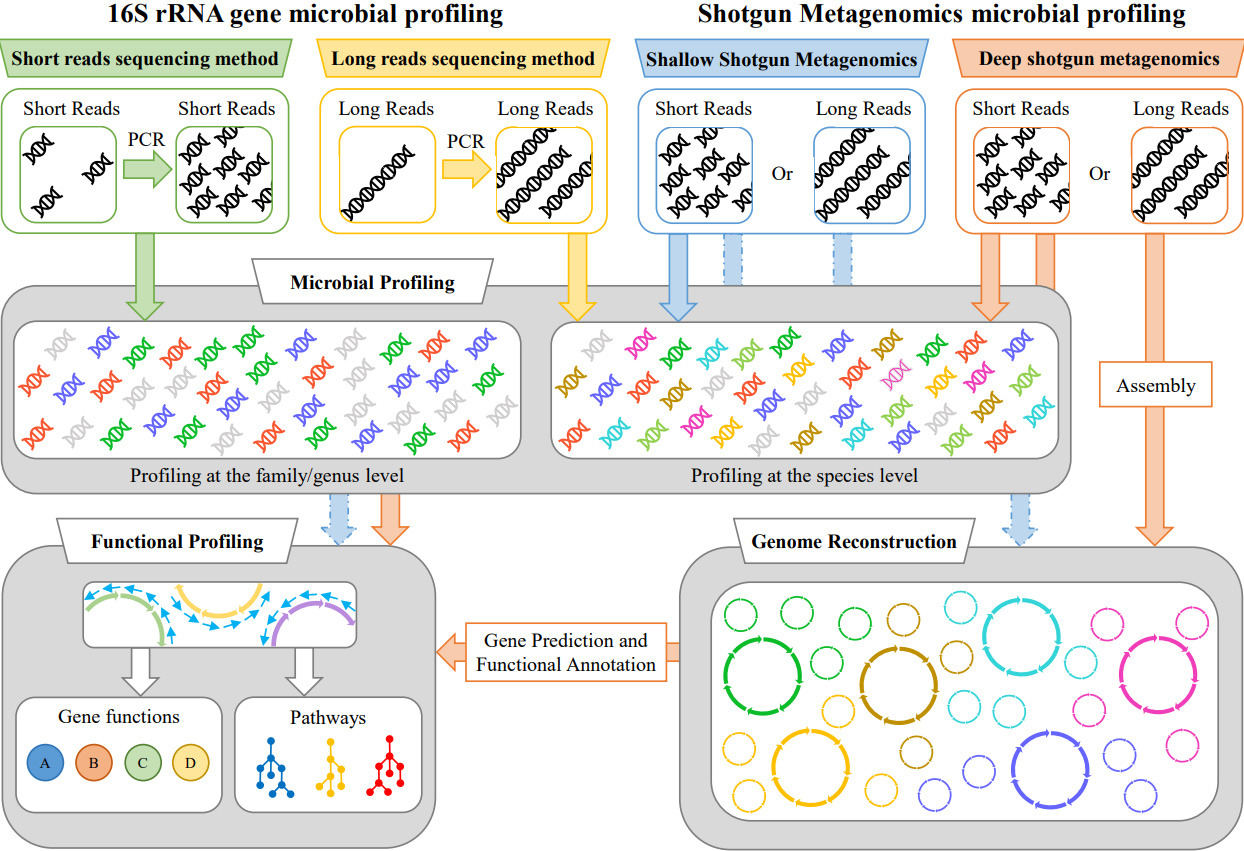
A breath of fresh air in microbiome science: shallow shotgun metagenomics for a reliable disentangling of microbial ecosystems