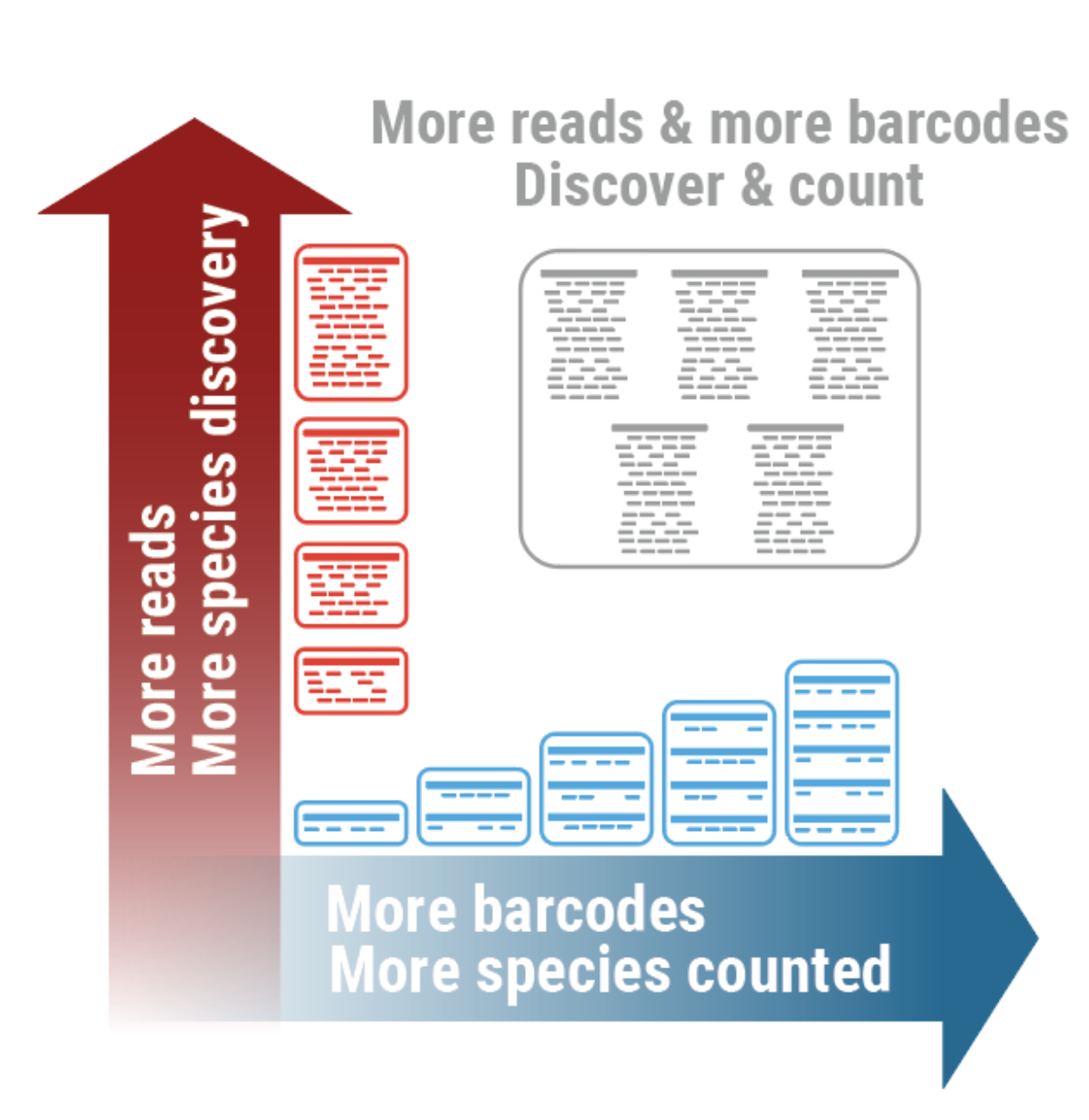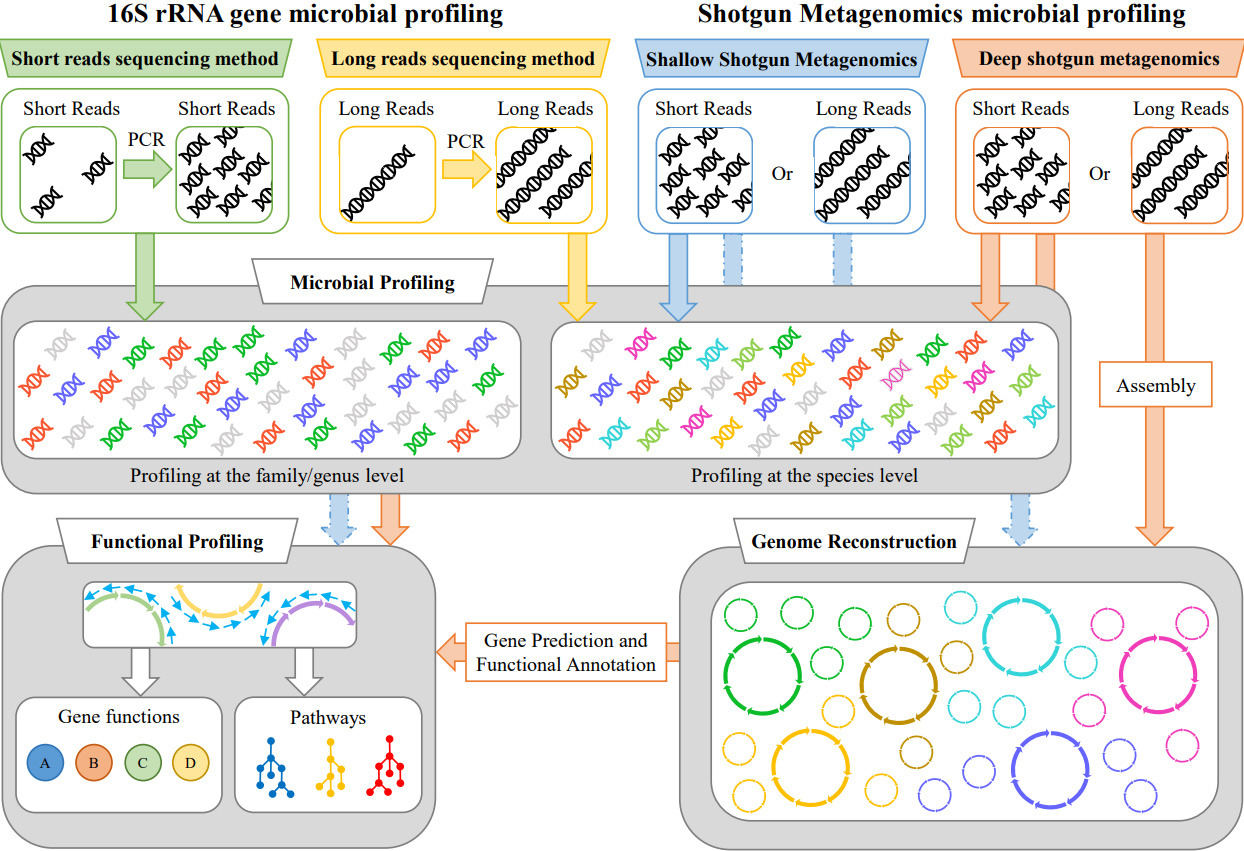
A breath of fresh air in microbiome science: shallow shotgun metagenomics for a reliable disentangling of microbial ecosystems
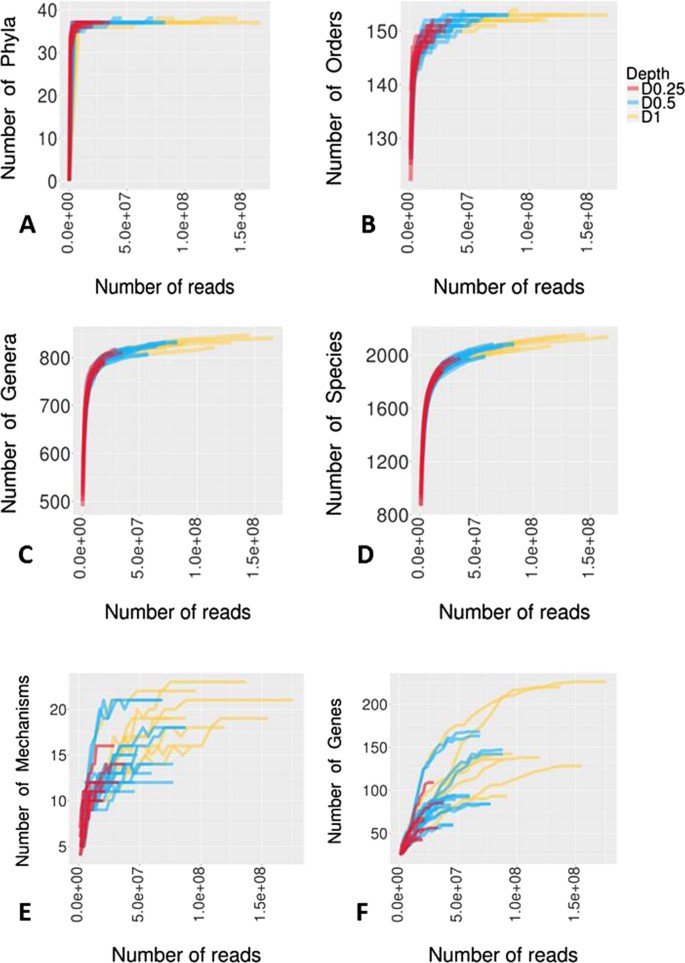
Impact of sequencing depth on the characterization of the microbiome and resistome | Scientific Reports
Phylogenetic microbiota profiling in fecal samples depends on combination of sequencing depth and choice of NGS analysis method | PLOS ONE
Frontiers | Impact of Host DNA and Sequencing Depth on the Taxonomic Resolution of Whole Metagenome Sequencing for Microbiome Analysis
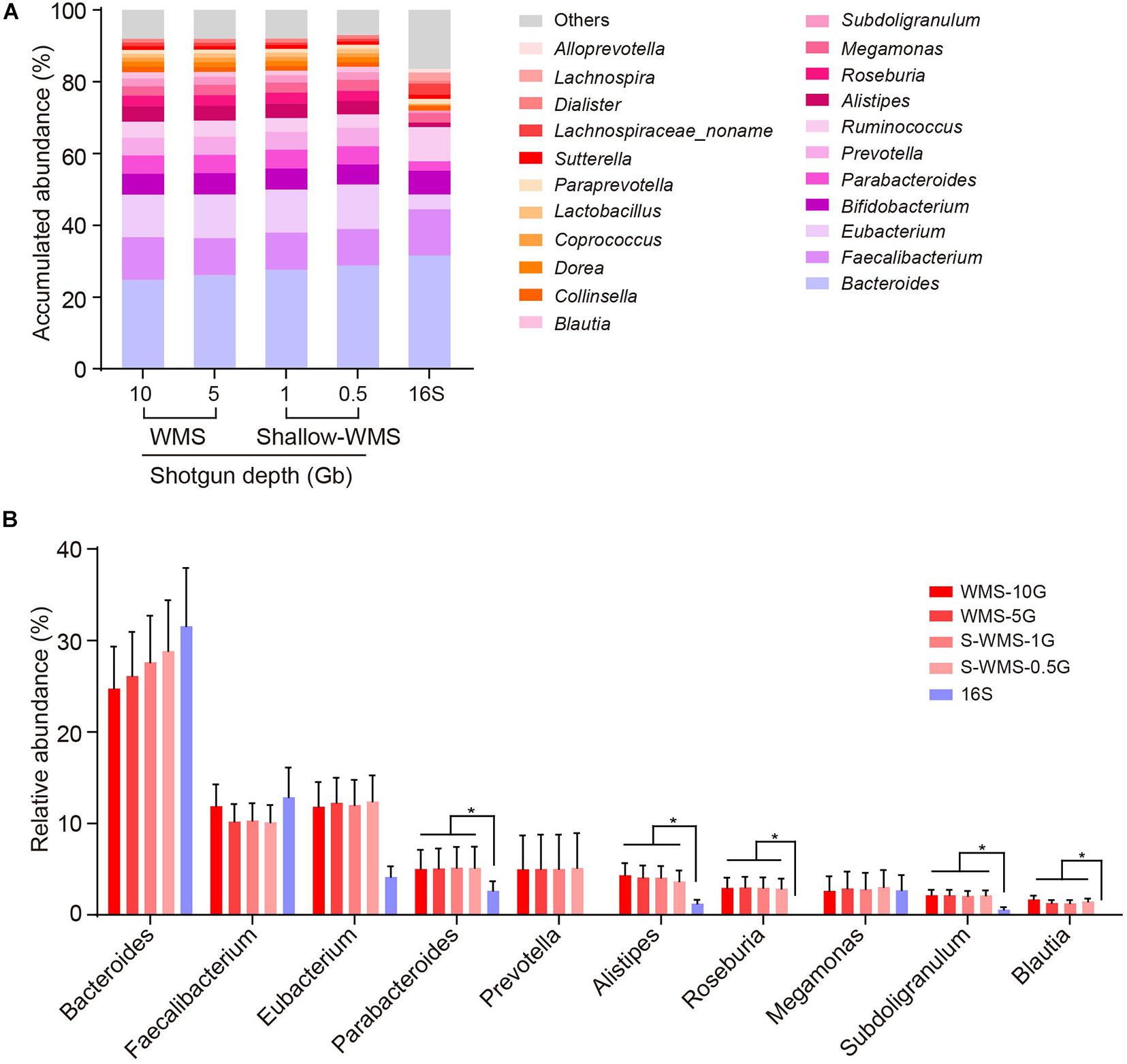
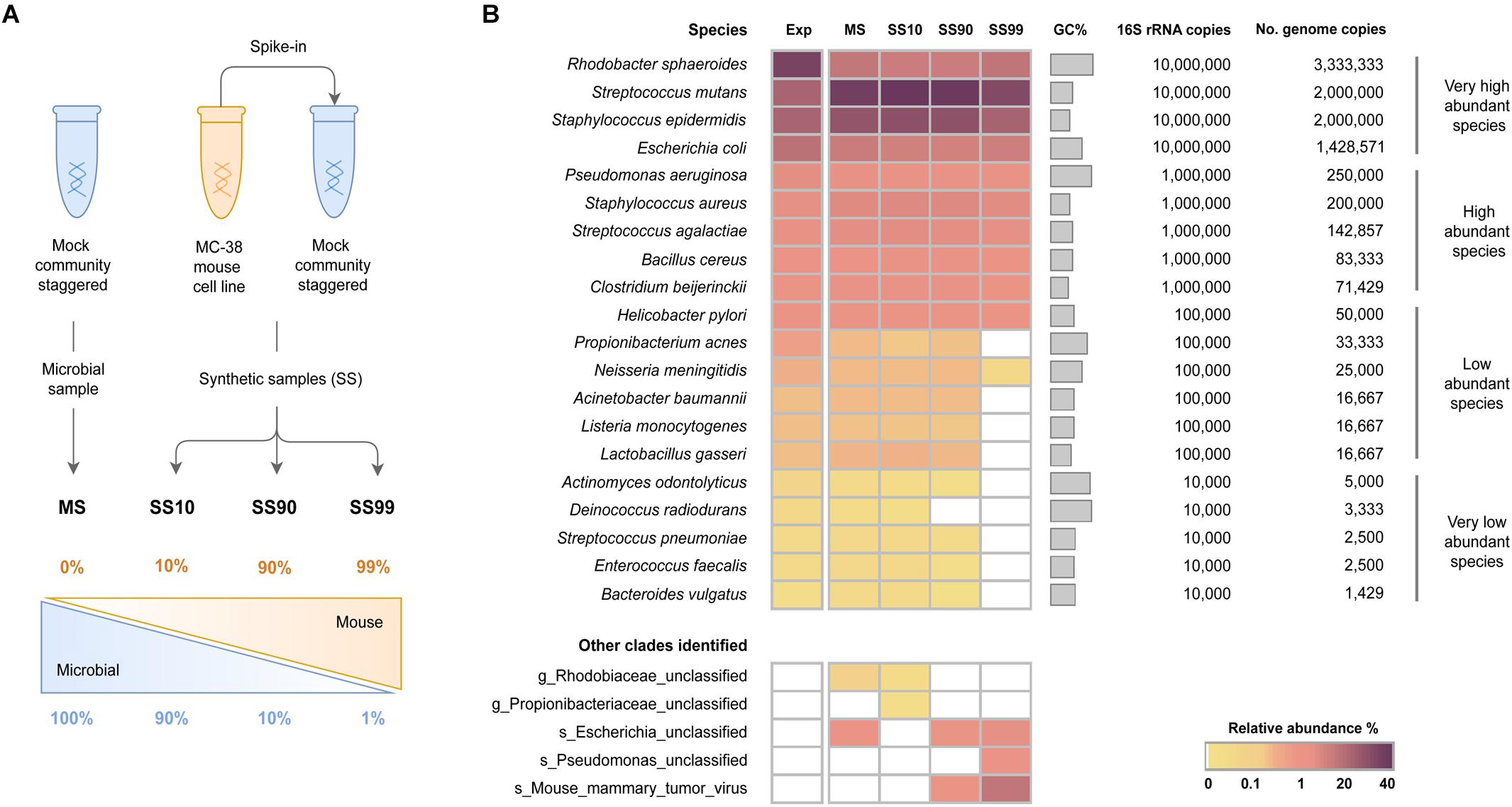
Frontiers | Characterization of Shallow Whole-Metagenome Shotgun Sequencing as a High-Accuracy and Low-Cost Method by Complicated Mock Microbiomes

STROBE-metagenomics: a STROBE extension statement to guide the reporting of metagenomics studies - The Lancet Infectious Diseases
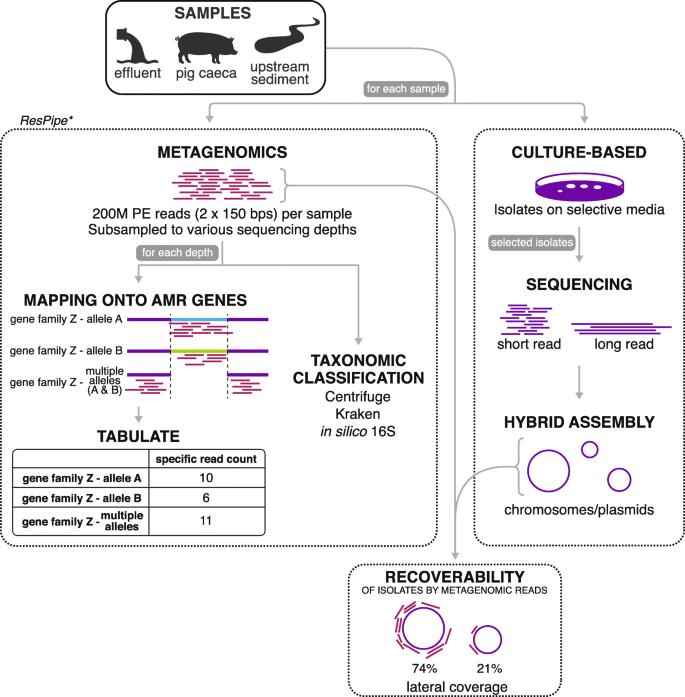
The impact of sequencing depth on the inferred taxonomic composition and AMR gene content of metagenomic samples | Environmental Microbiome | Full Text

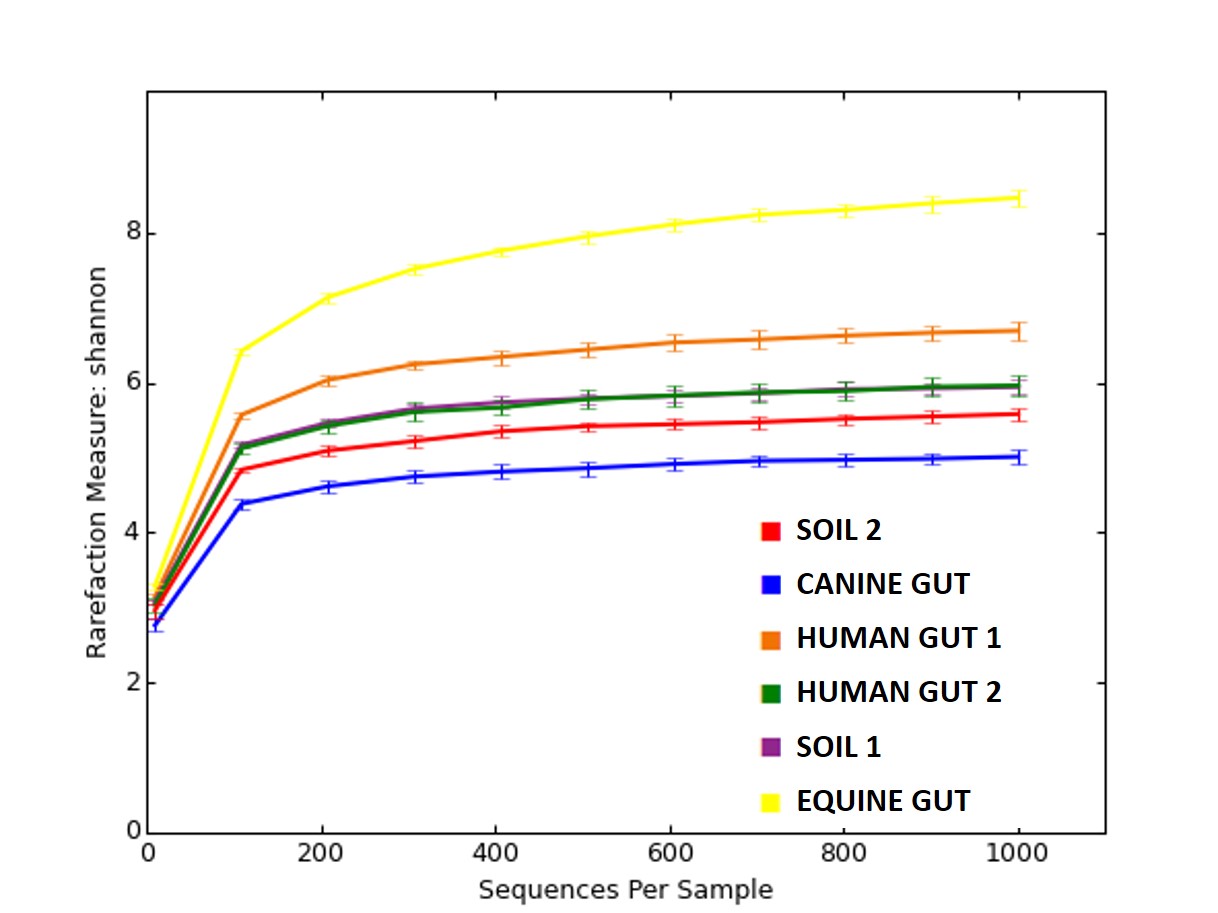
Impact of sequencing depth on genus-level richness. Three methods are... | Download Scientific Diagram
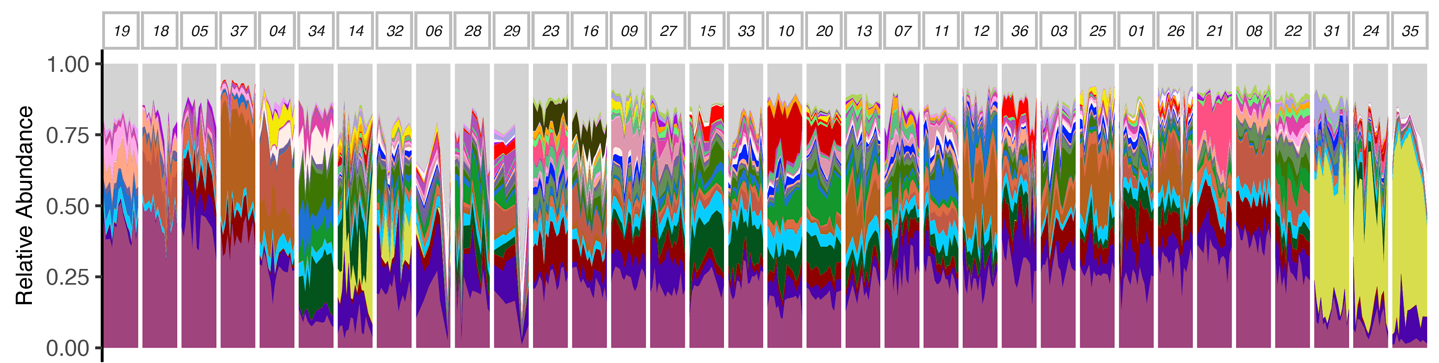
The relative abundances (metagenome, average sequencing depth, blue)... | Download Scientific Diagram
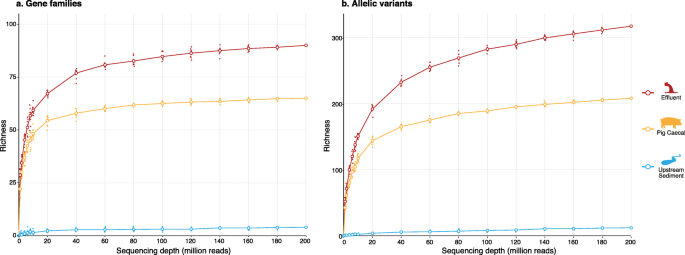
The impact of sequencing depth on the inferred taxonomic composition and AMR gene content of metagenomic samples | Environmental Microbiome | Full Text
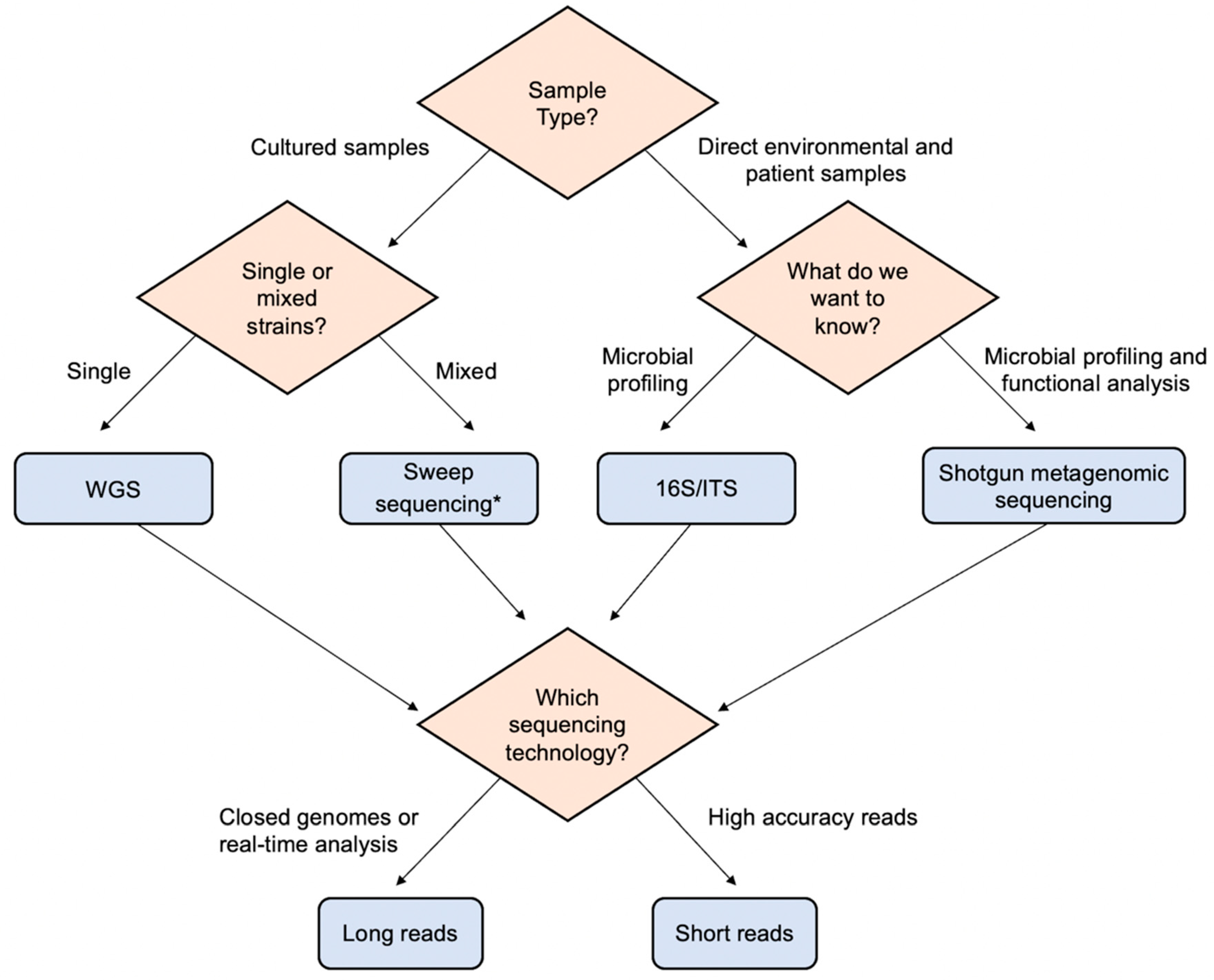
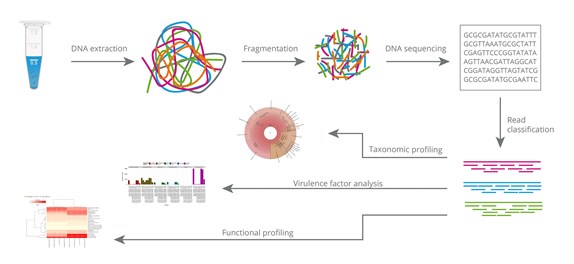
IJMS | Free Full-Text | Combination of Whole Genome Sequencing and Metagenomics for Microbiological Diagnostics
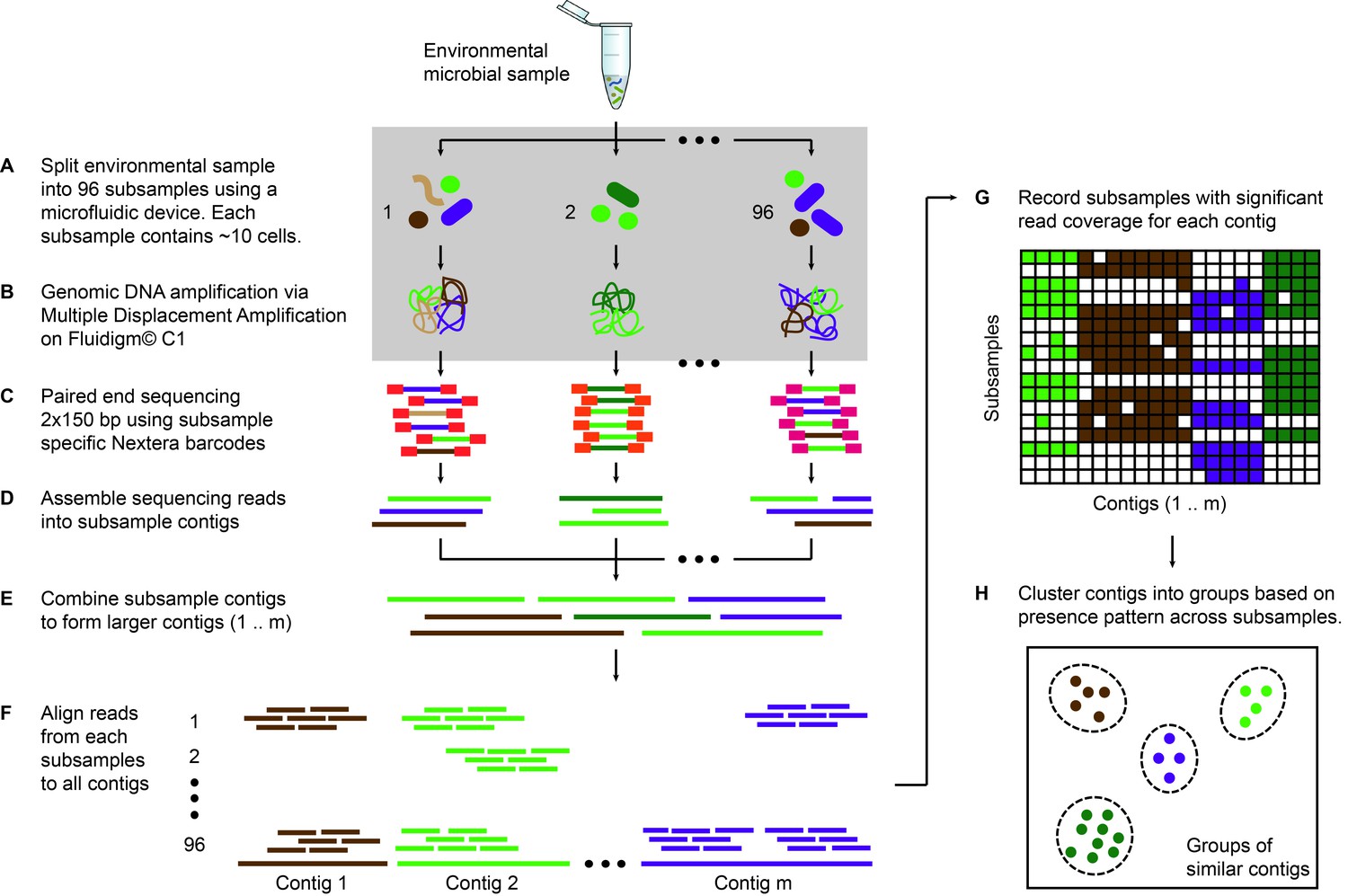
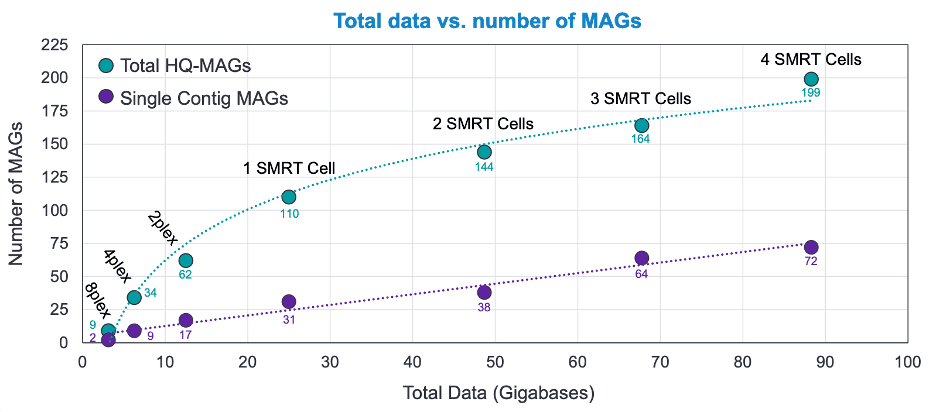
Microfluidic-based mini-metagenomics enables discovery of novel microbial lineages from complex environmental samples | eLife
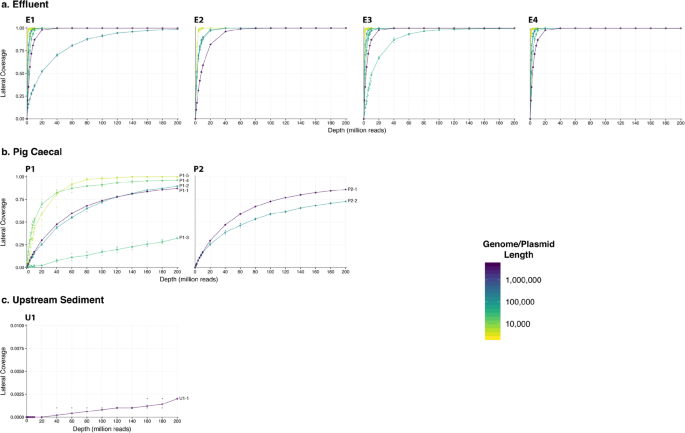
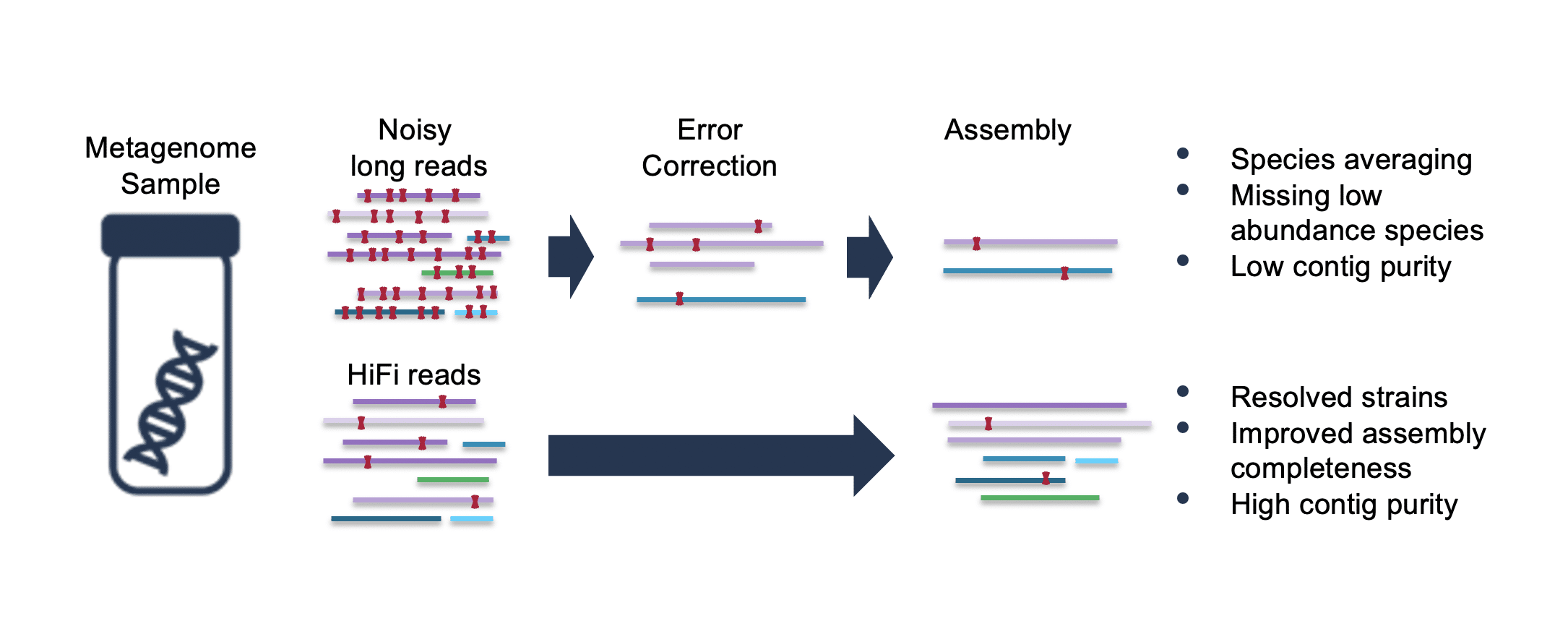
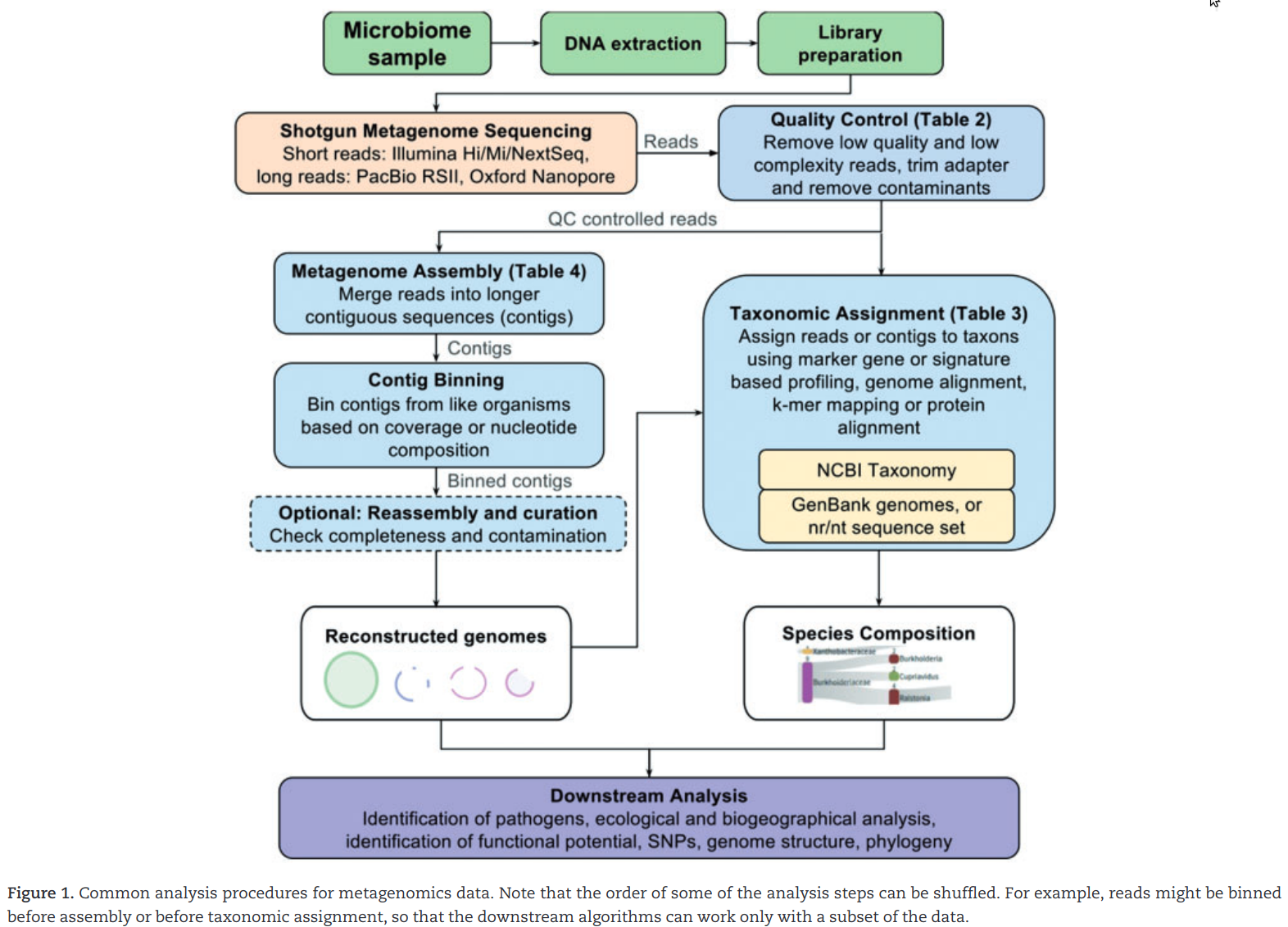
![PDF] Metagenomic Assembly: Overview, Challenges and Applications | Semantic Scholar PDF] Metagenomic Assembly: Overview, Challenges and Applications | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/488bffc68f690e33a65b472fc69914721b87d879/3-Figure2-1.png)