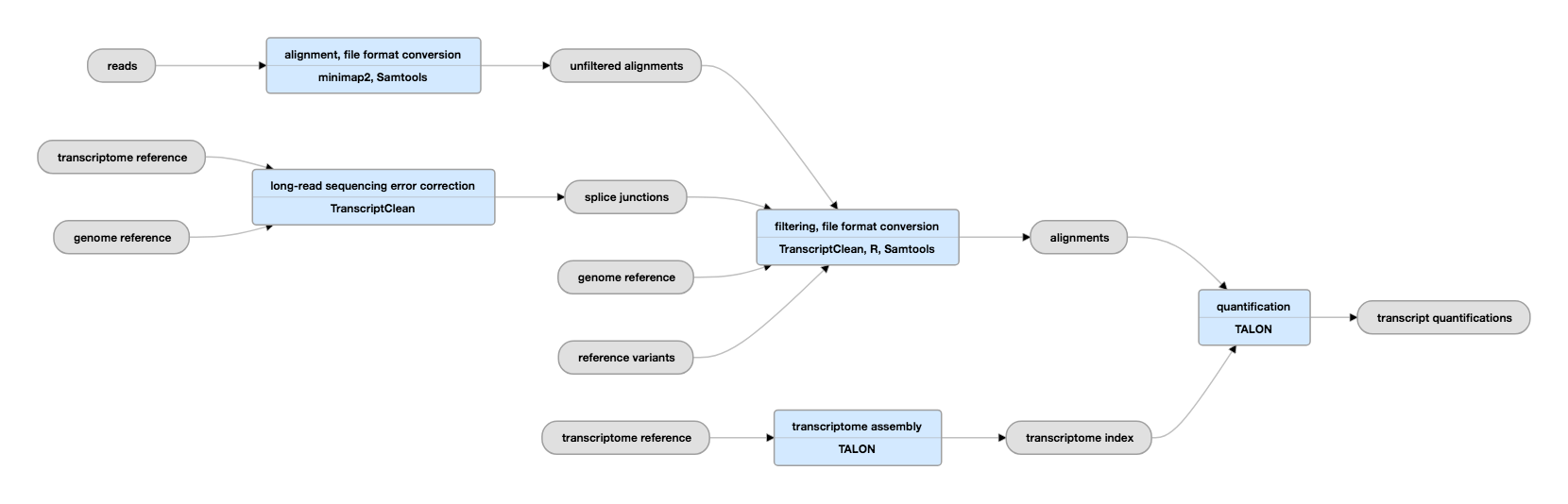
A technology-agnostic long-read analysis pipeline for transcriptome discovery and quantification | bioRxiv

Nanopore long-read RNA-seq and absolute quantification delineate transcription dynamics in early embryo development of an insect pest | Scientific Reports
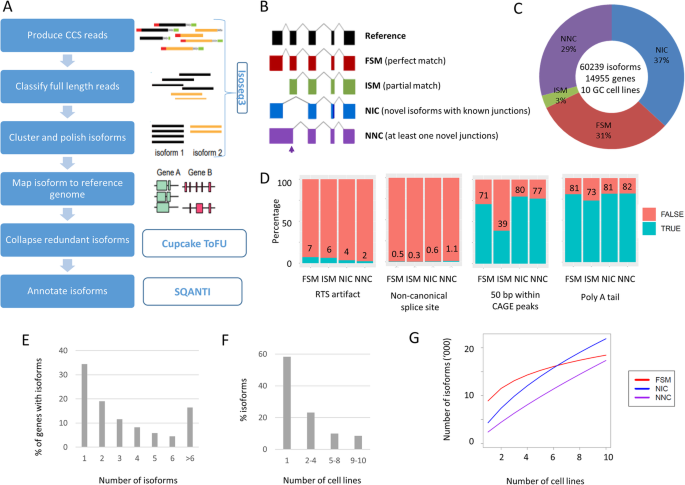
Long-read transcriptome sequencing reveals abundant promoter diversity in distinct molecular subtypes of gastric cancer | Genome Biology | Full Text

ESPRESSO: Robust discovery and quantification of transcript isoforms from error-prone long-read RNA-seq data | Science Advances

Researchers use a combination of third generation long-read RNA sequencing technology and short-read RNAseq to characterize the splicing landscape of prostate cancer | RNA-Seq Blog

A systematic benchmark of Nanopore long read RNA sequencing for transcript level analysis in human cell lines | bioRxiv

IJMS | Free Full-Text | L-RAPiT: A Cloud-Based Computing Pipeline for the Analysis of Long-Read RNA Sequencing Data

Figure 1 from LONG-READ RNA SEQUENCING ANALYSIS OF THE LYTIC HUMAN CYTOMEGALOVIRUS TRANSCRIPTOME | Semantic Scholar

A systematic benchmark of Nanopore long read RNA sequencing for transcript level analysis in human cell lines | bioRxiv