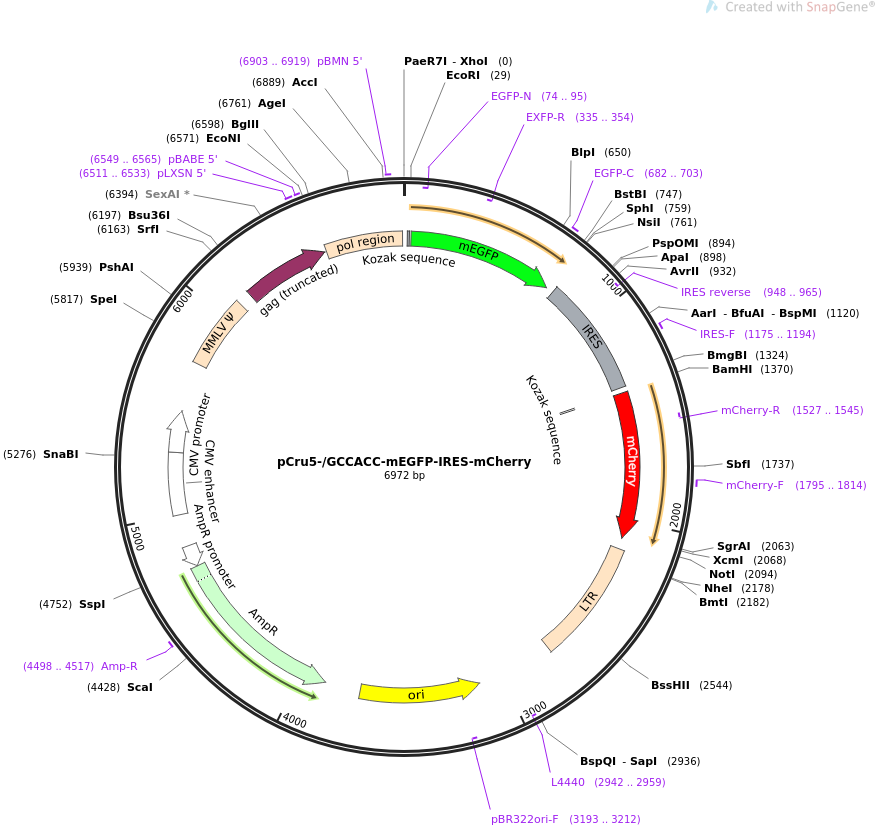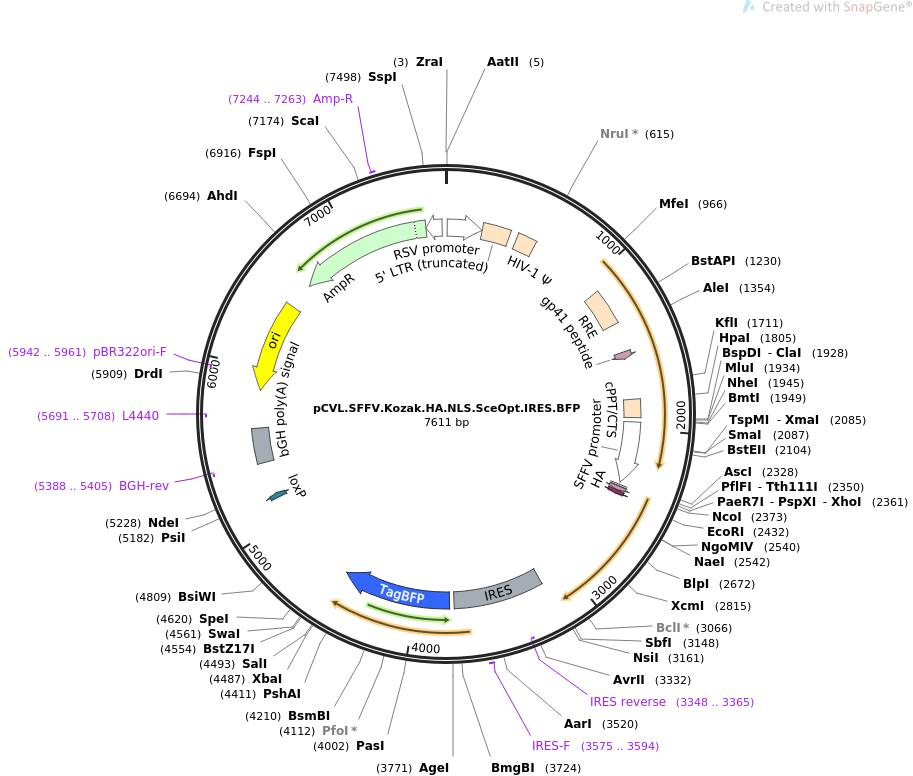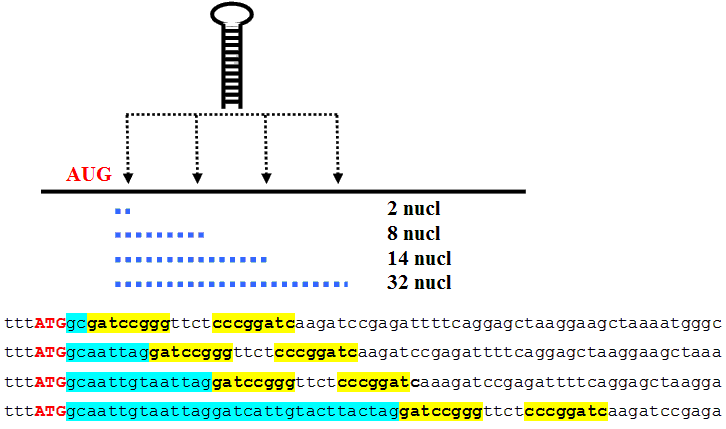
AUG_hairpin: program for prediction of a downstream hairpin potentially increasing initiation of translation at start AUG codon in a suboptimal context.
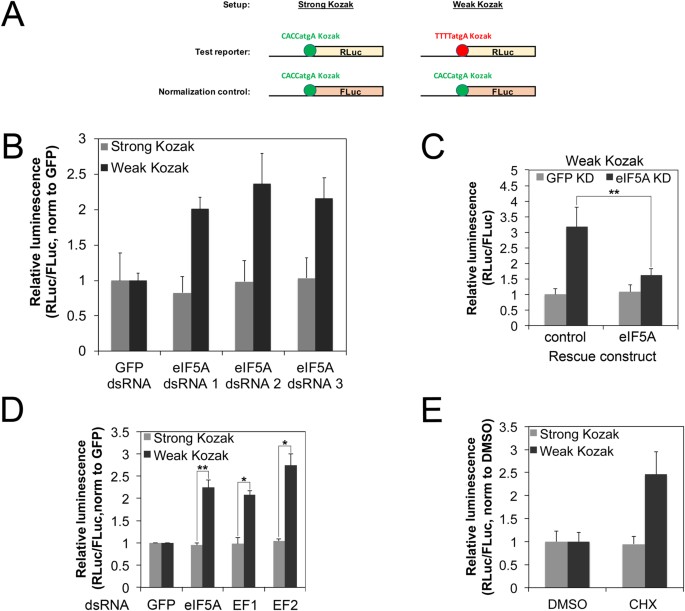
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports
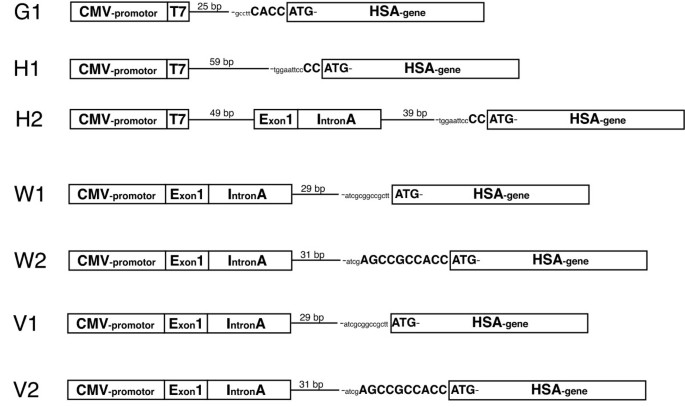
In vitro analysis of expression vectors for DNA vaccination of horses: the effect of a Kozak sequence | Acta Veterinaria Scandinavica | Full Text

Transcripts with weak Kozak sequences are spared when mRNA-specific... | Download Scientific Diagram

Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv
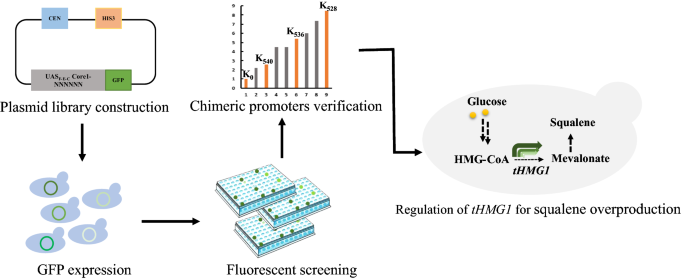
Fine-tuning the expression of pathway gene in yeast using a regulatory library formed by fusing a synthetic minimal promoter with different Kozak variants | Microbial Cell Factories | Full Text

Structural insights into the Kozak sequence interactions within the mammalian late-stage initiation complexes | bioRxiv

Bioinformatic analyses of mammalian 5'-UTR sequence properties of mRNAs predicts alternative translation initiation sites | BMC Bioinformatics | Full Text

Combination of Kozak sequence and optimized gene arrangement further... | Download Scientific Diagram
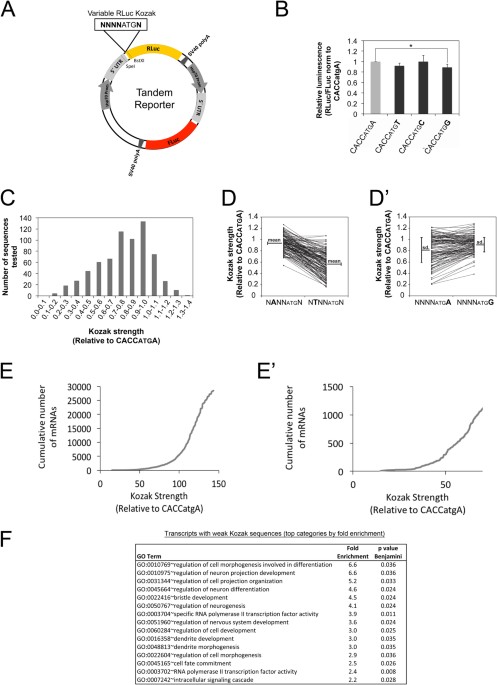
Changes in global translation elongation or initiation rates shape the proteome via the Kozak sequence | Scientific Reports

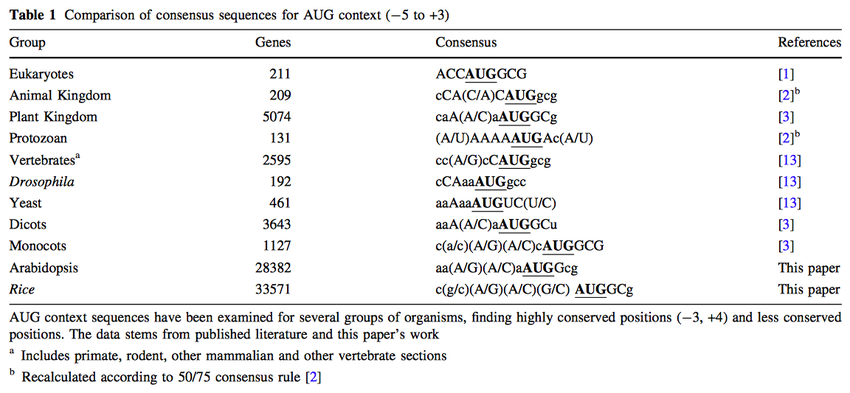
Conservation and Variability of the AUG Initiation Codon Context in Eukaryotes: Trends in Biochemical Sciences

Quantitative analysis of mammalian translation initiation sites by FACS‐seq | Molecular Systems Biology