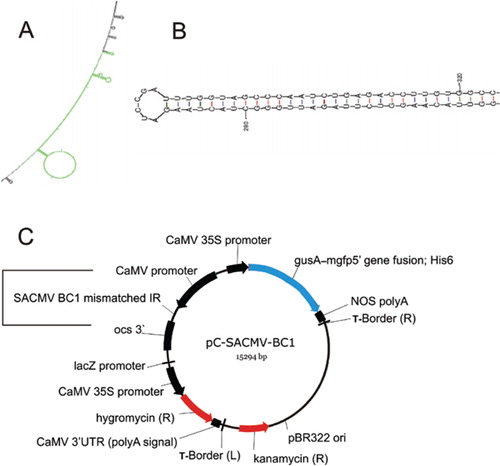
Construction of effective inverted repeat silencing constructs using sodium bisulfite treatment coupled with strand-specific PCR | BioTechniques
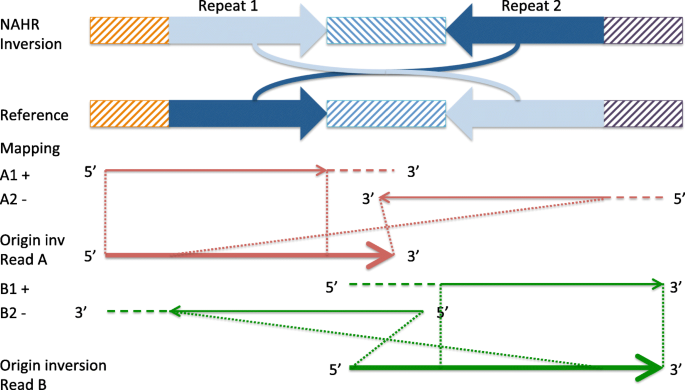
npInv: accurate detection and genotyping of inversions using long read sub-alignment | BMC Bioinformatics | Full Text

Global analysis of inverted repeat sequences in human gene promoters reveals their non-random distribution and association with specific biological pathways - ScienceDirect
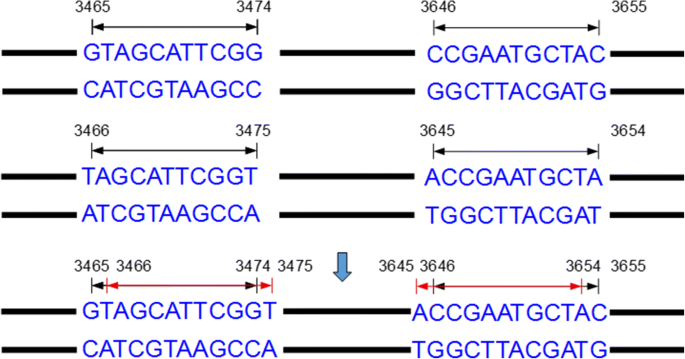
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text
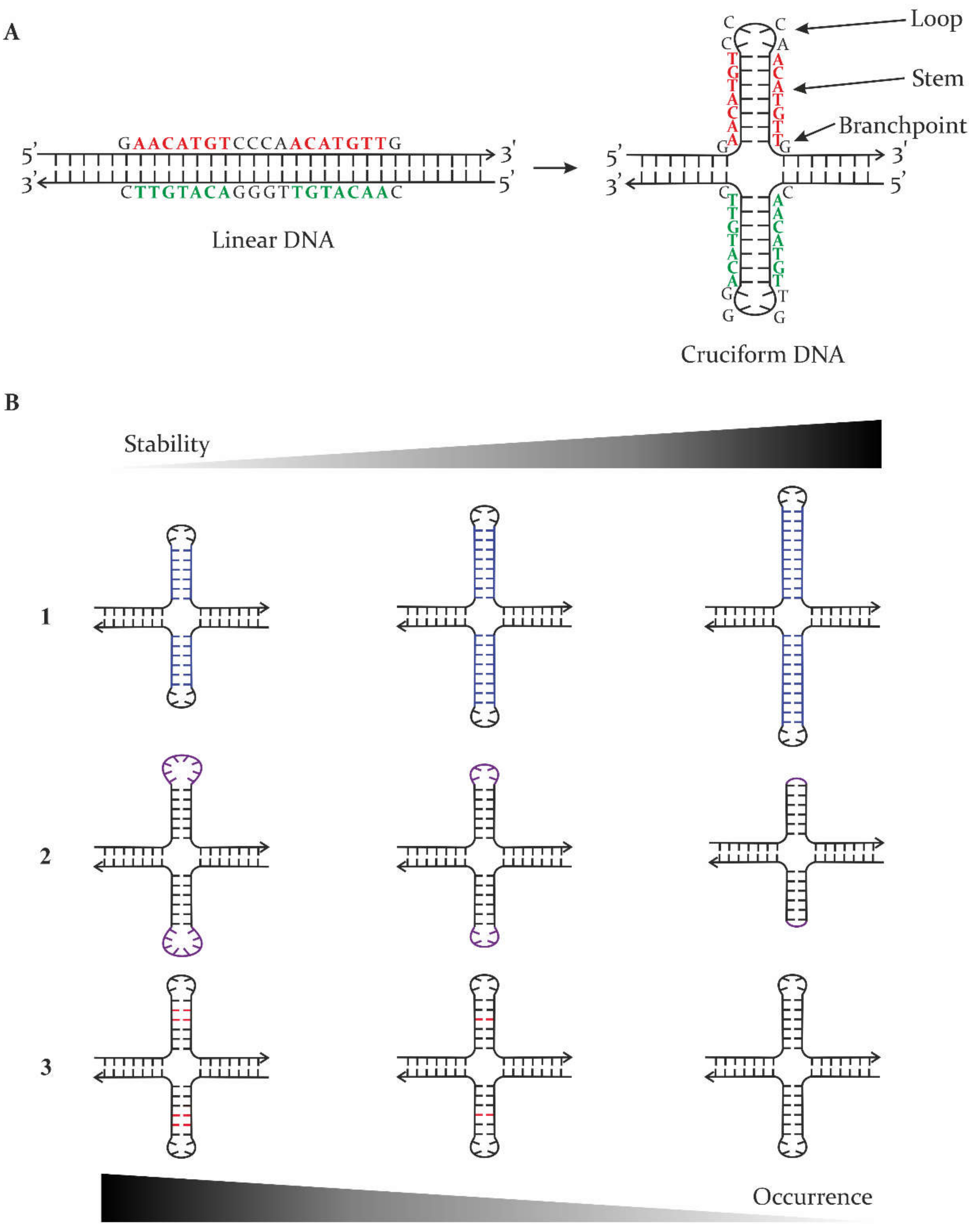
IJMS | Free Full-Text | Interaction of Proteins with Inverted Repeats and Cruciform Structures in Nucleic Acids
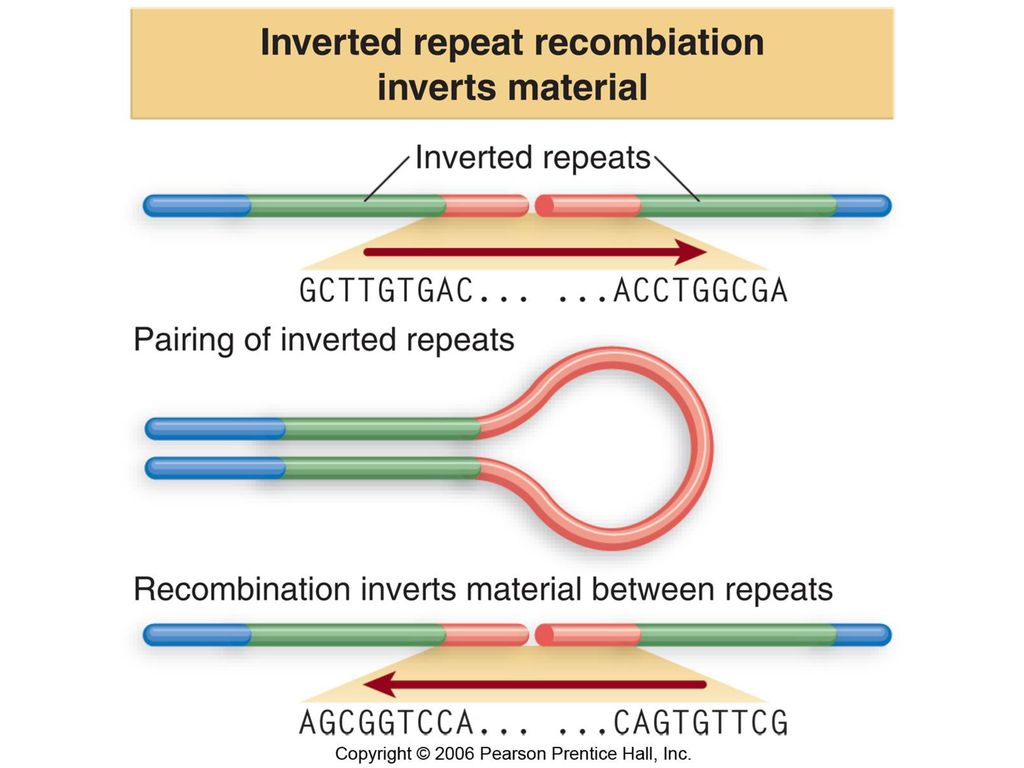
Figure: Title: Transposons have inverted repeats and generate target repeats Caption: Transposons have inverted terminal repeats and generate direct. - ppt download
Unusually Long Palindromes Are Abundant in Mitochondrial Control Regions of Insects and Nematodes | PLOS ONE
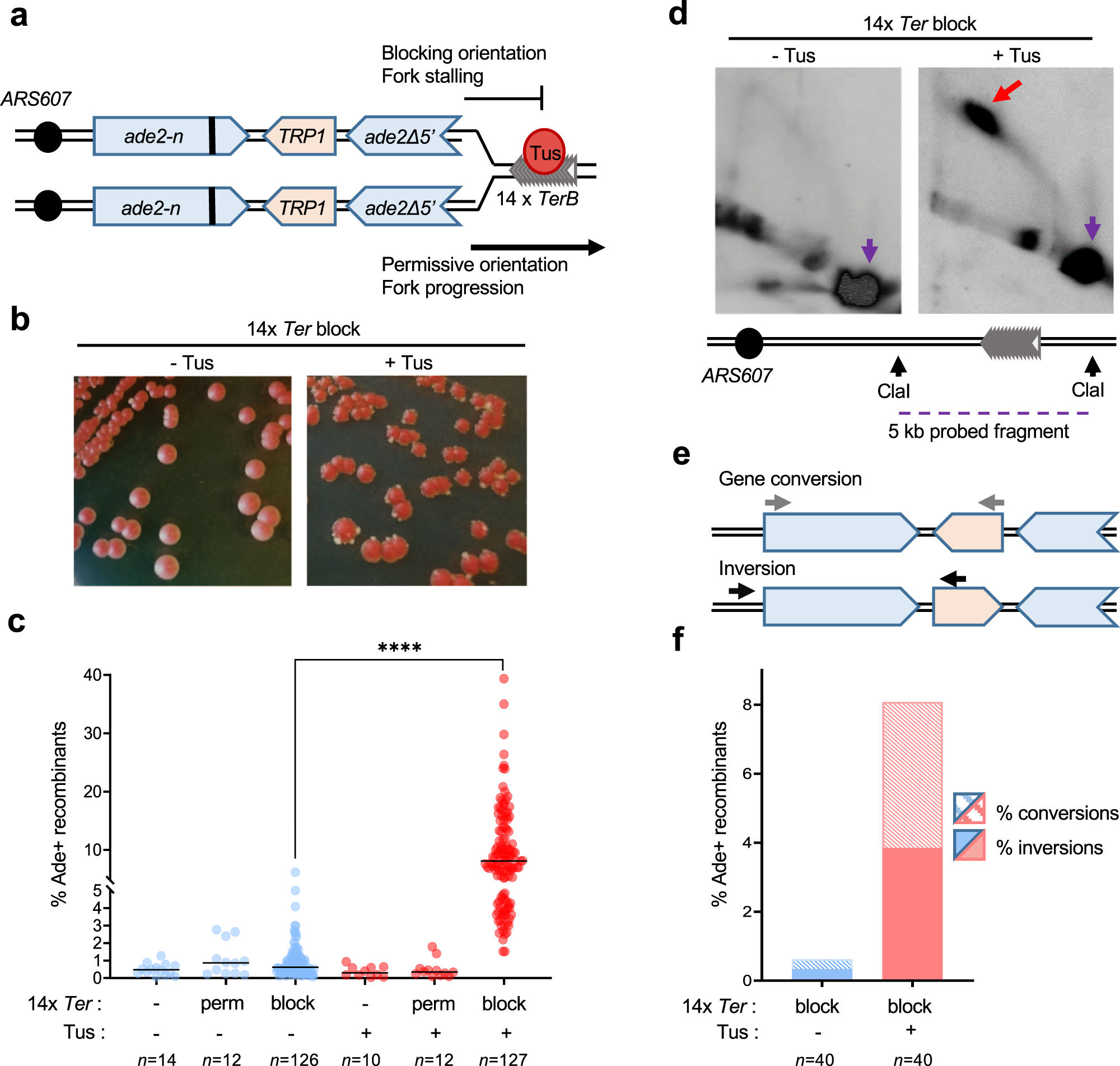
Mechanism for inverted-repeat recombination induced by a replication fork barrier | Nature Communications
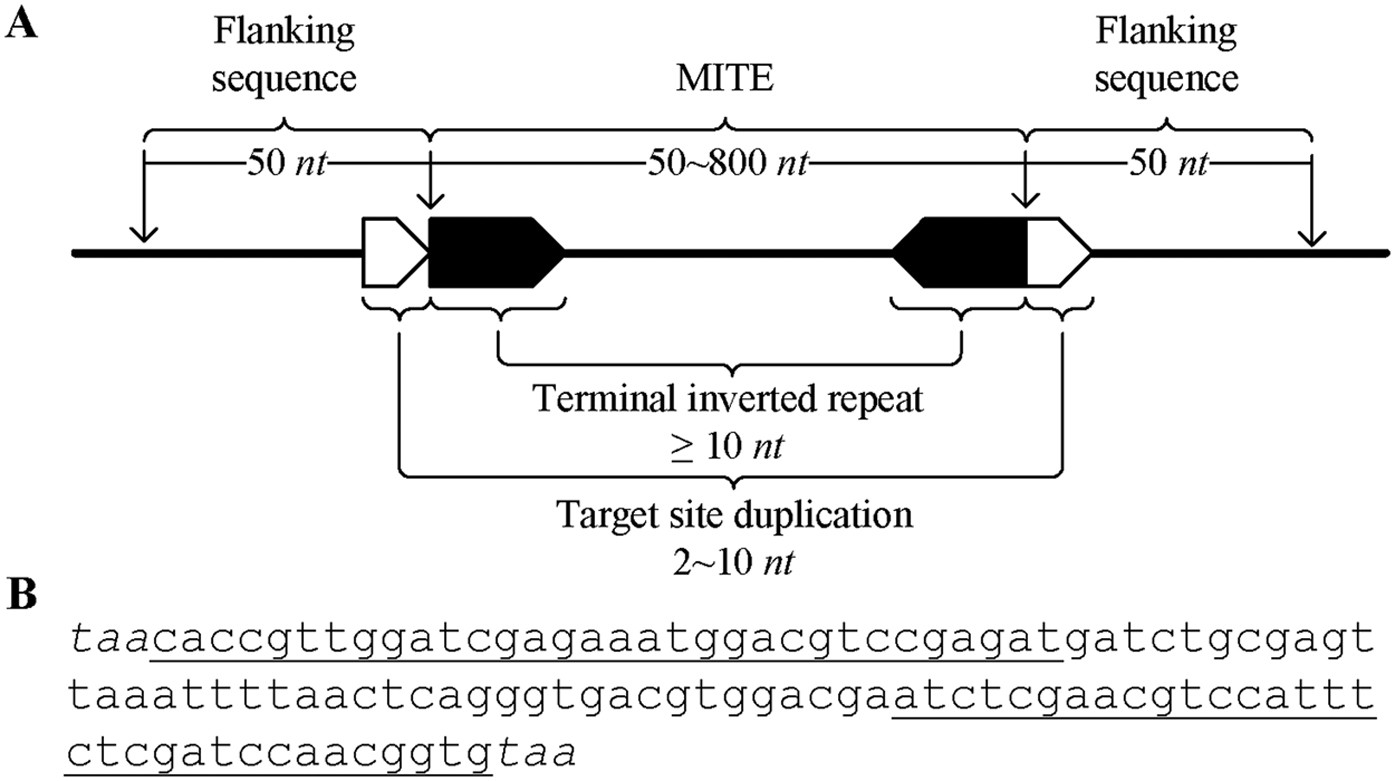
detectMITE: A novel approach to detect miniature inverted repeat transposable elements in genomes | Scientific Reports
![PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c29fd938f405ed8516c10b9bdaa27f988b29dd31/2-Figure1-1.png)
PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar

Fusion of nearby inverted repeats by a replication-based mechanism leads to formation of dicentric and acentric chromosomes that cause genome instability in budding yeast