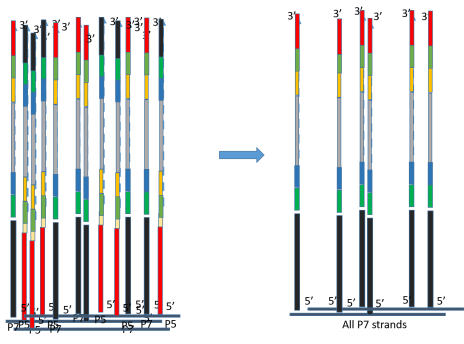
Addressing challenges in the production and analysis of illumina sequencing data | BMC Genomics | Full Text

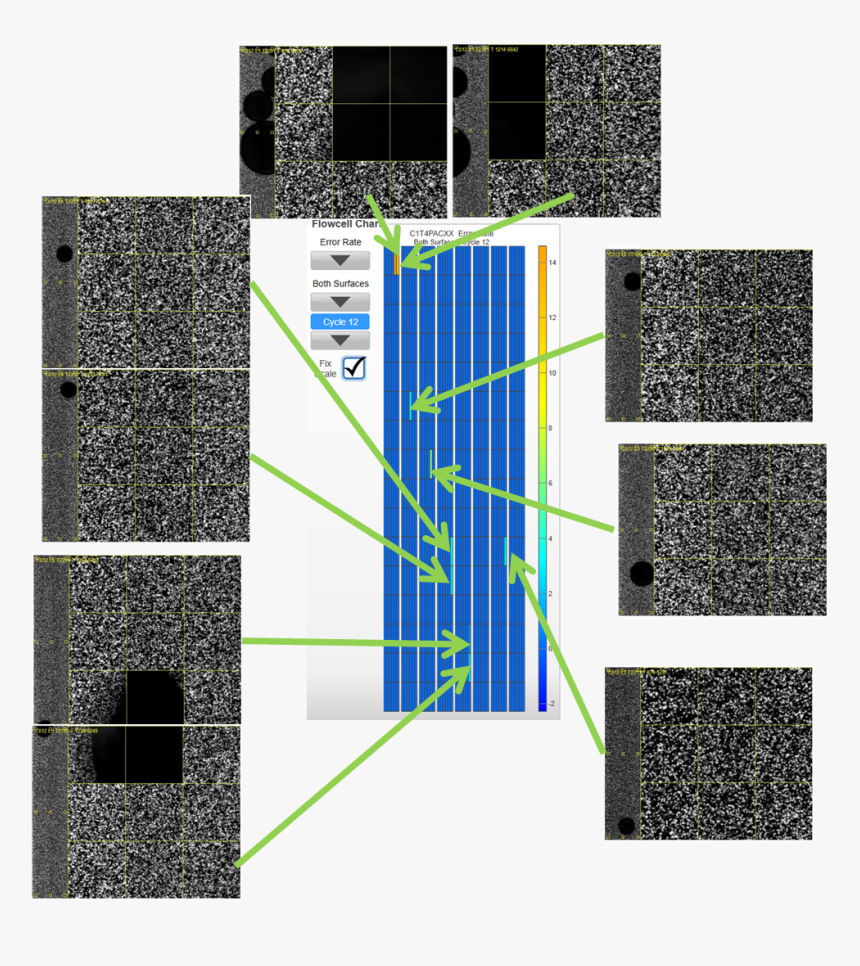
almost) everything you wanted to know about @illumina HiSeq 4000...and some stuff you didn't - Enseqlopedia

Flow-Cell-Based Technology for Massively Parallel Characterization of Base-Modified DNA Aptamers | Analytical Chemistry