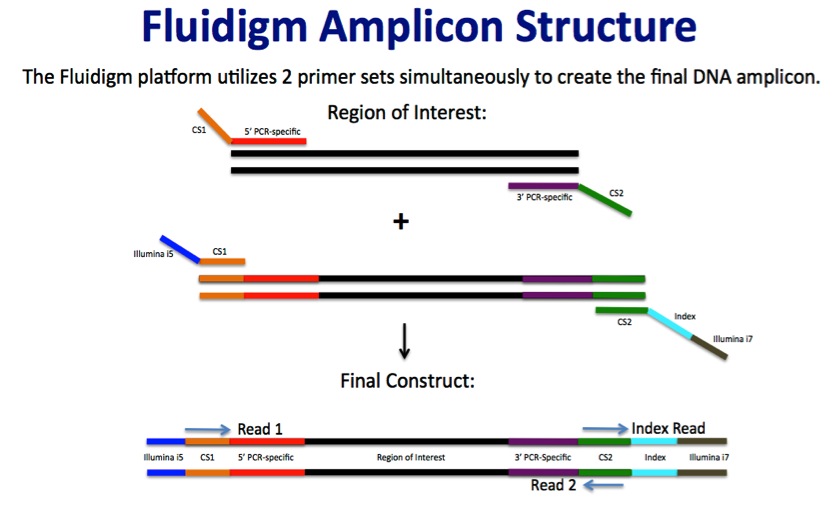
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech
![PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c5433f6643987939f74806d2f91db31fe7eac03a/4-Figure1-1.png)
PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar
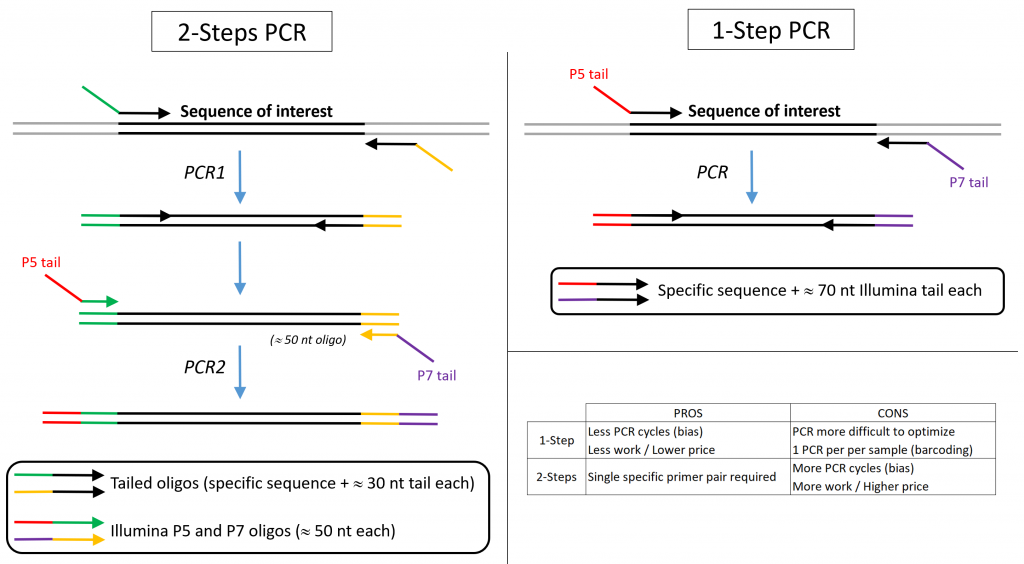
Illumina amplicon library preparation through 2-Step PCR amplification. | Download Scientific Diagram
![A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq [PeerJ] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2016/2492/1/fig-1-2x.jpg)
A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq [PeerJ]
![PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c5433f6643987939f74806d2f91db31fe7eac03a/8-Figure3-1.png)
PDF] A method for high precision sequencing of near full-length 16S rRNA genes on an Illumina MiSeq | Semantic Scholar

Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems

Reproducibility and repeatability of six high-throughput 16S rDNA sequencing protocols for microbiota profiling - ScienceDirect
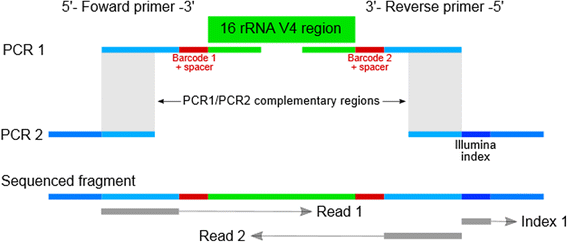
A novel ultra high-throughput 16S rRNA gene amplicon sequencing library preparation method for the Illumina HiSeq platform | Microbiome | Full Text

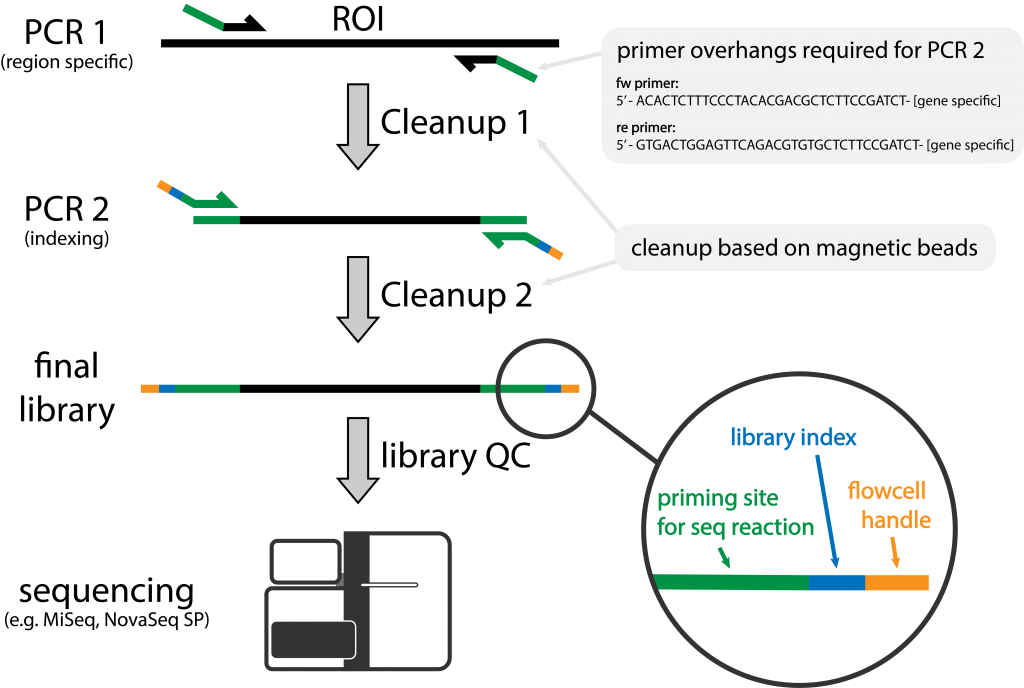
![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-3-full.png)


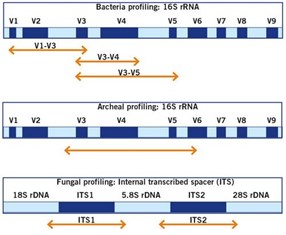

![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)