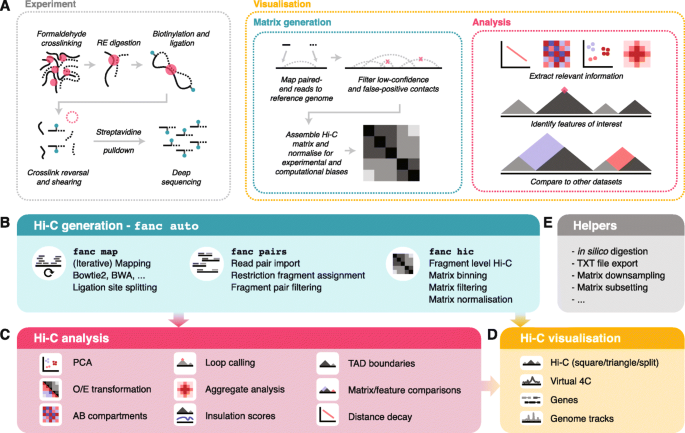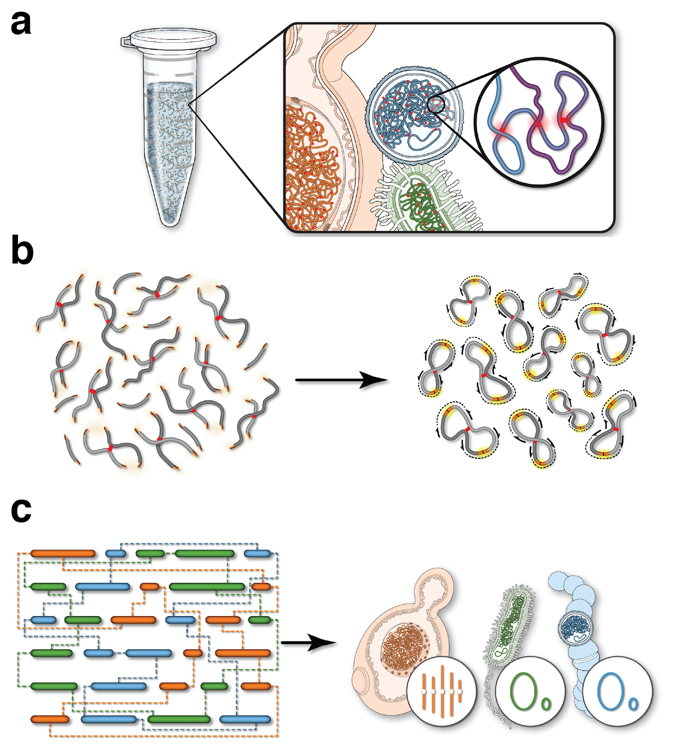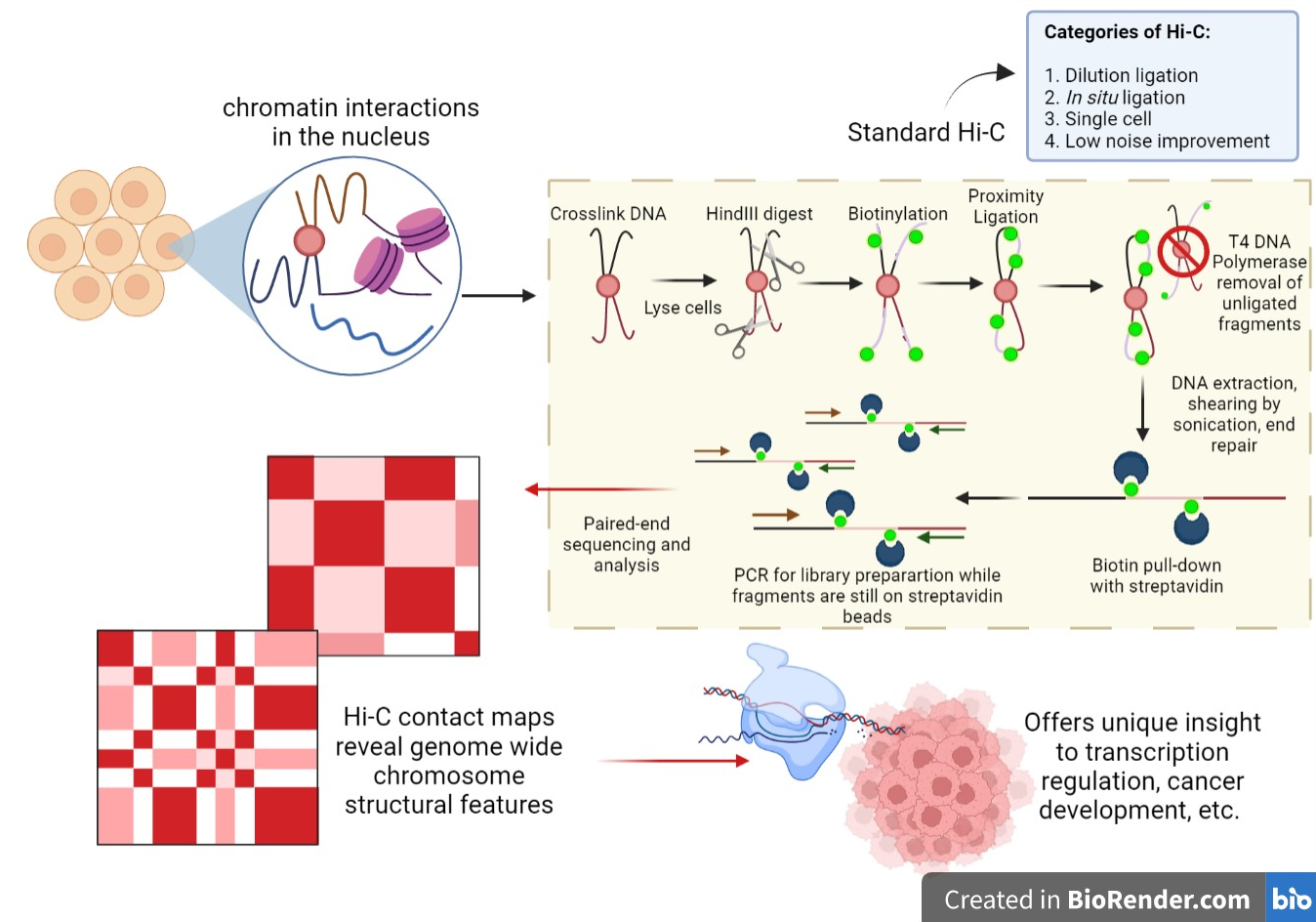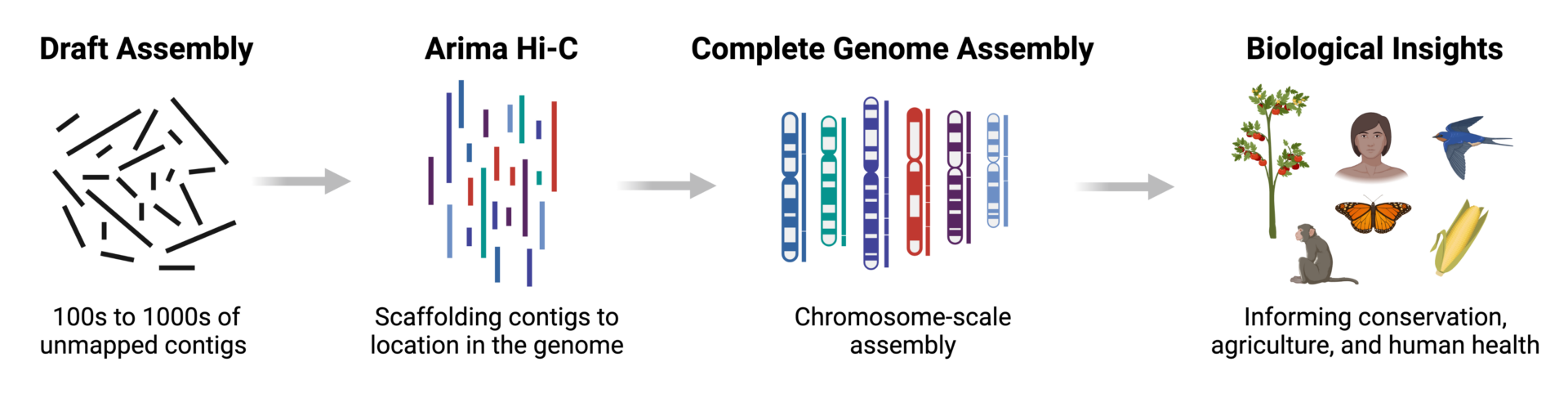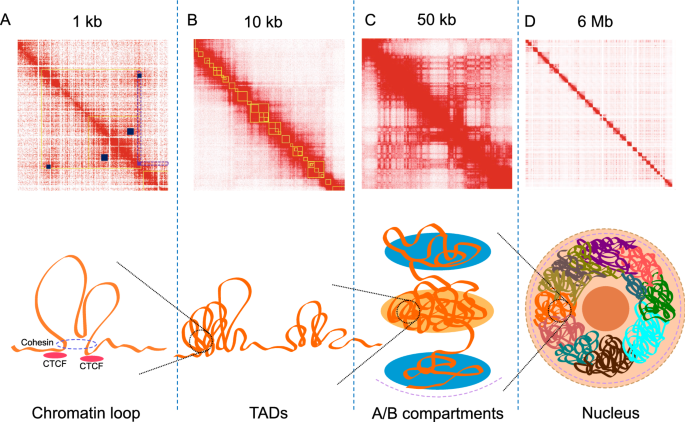
Seeing the forest through the trees: prioritising potentially functional interactions from Hi-C | Epigenetics & Chromatin | Full Text

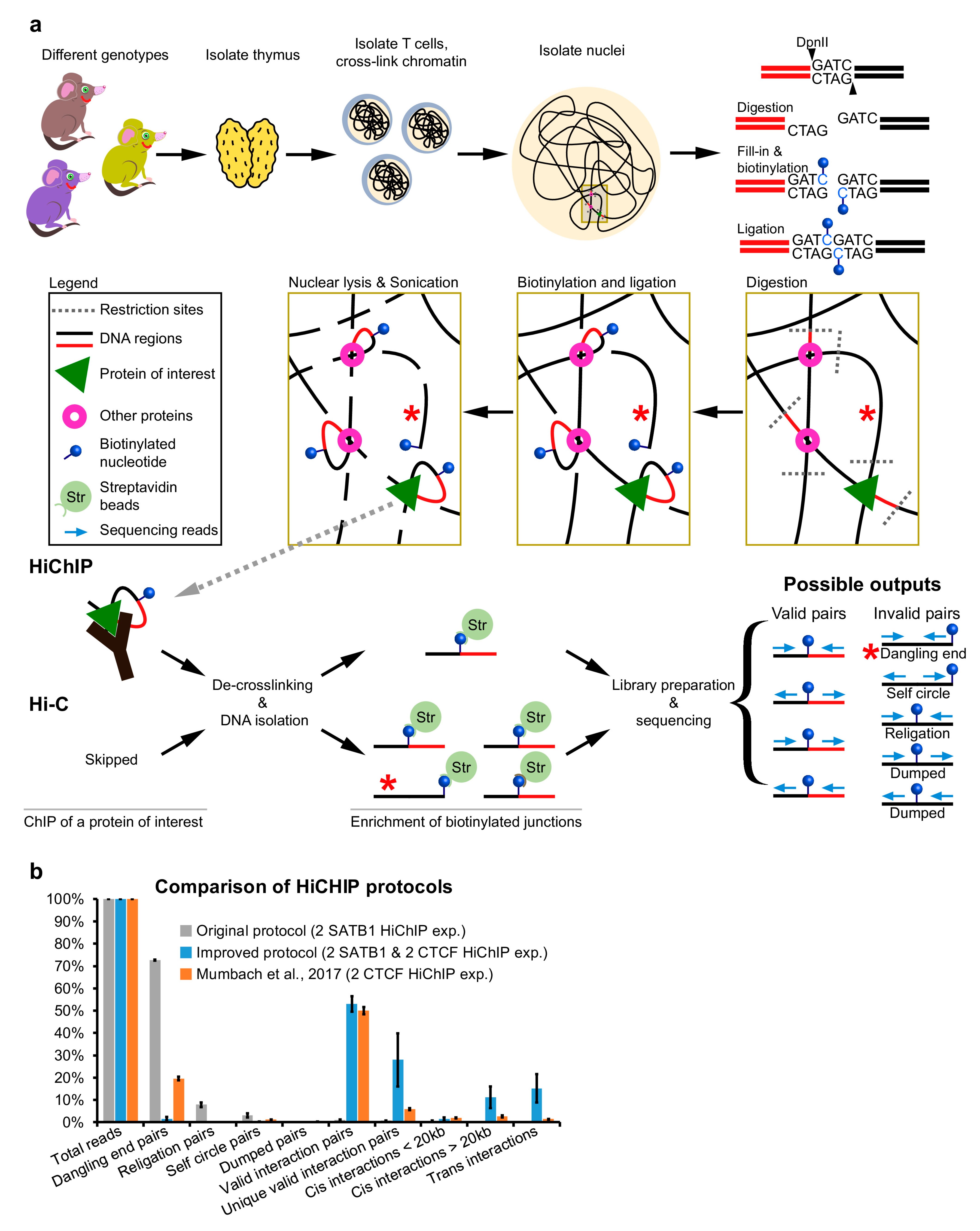
Hi‐C 3.0: Improved Protocol for Genome‐Wide Chromosome Conformation Capture - Lafontaine - 2021 - Current Protocols - Wiley Online Library
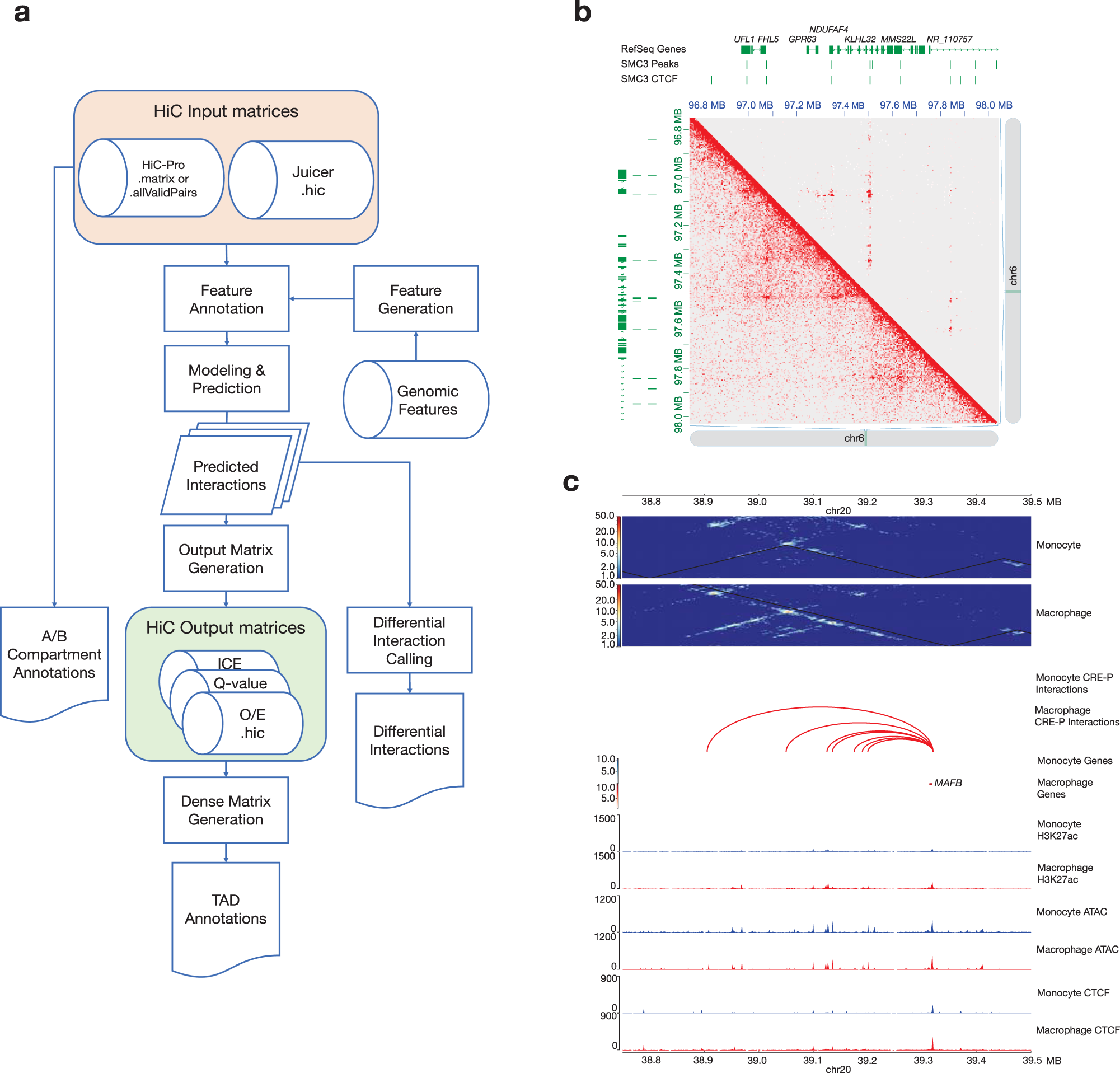
HiC-DC+ enables systematic 3D interaction calls and differential analysis for Hi-C and HiChIP | Nature Communications

Hi-C 2.0: An optimized Hi-C procedure for high-resolution genome-wide mapping of chromosome conformation - ScienceDirect

Liquid chromatin Hi-C characterizes compartment-dependent chromatin interaction dynamics | Nature Genetics