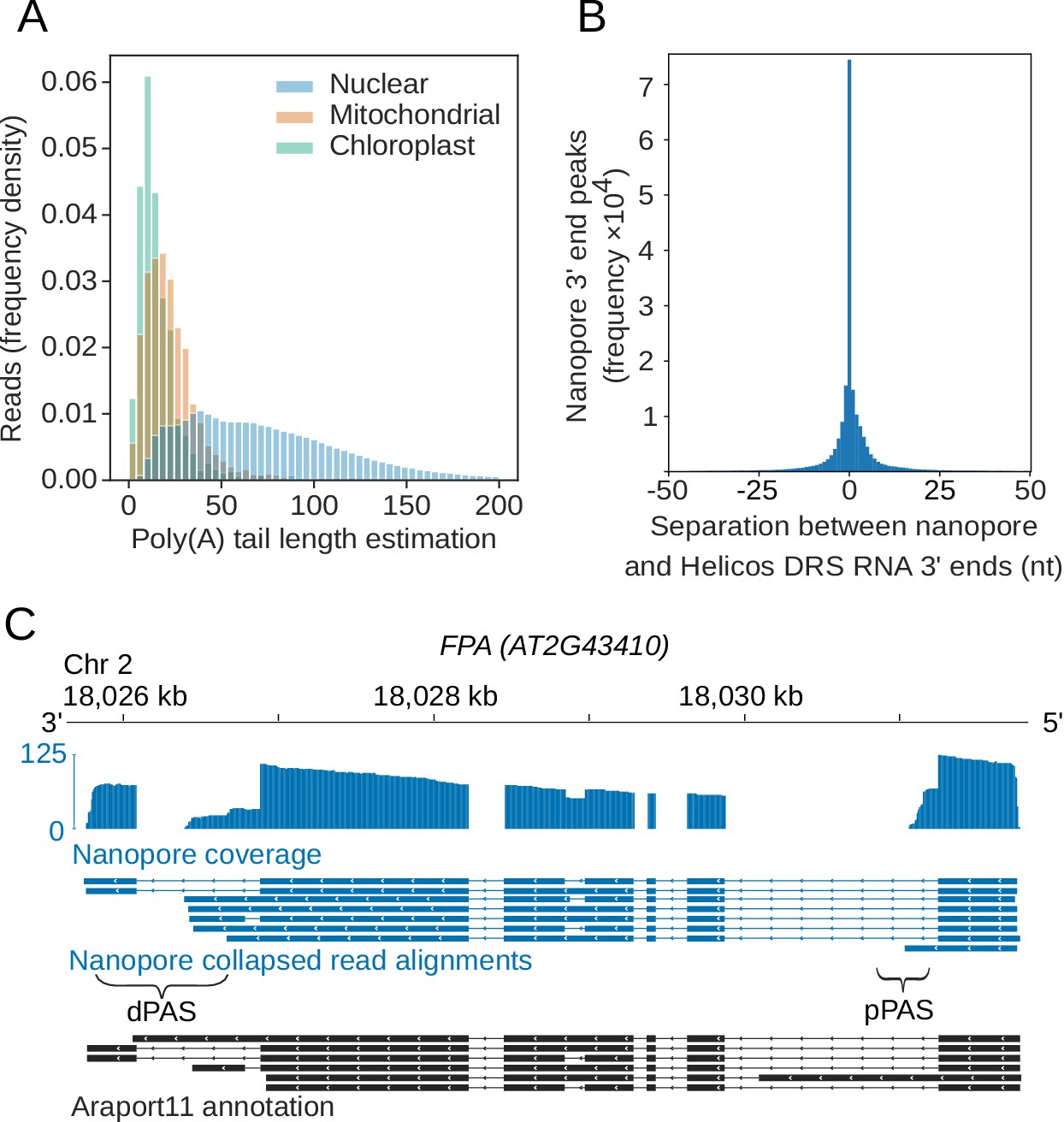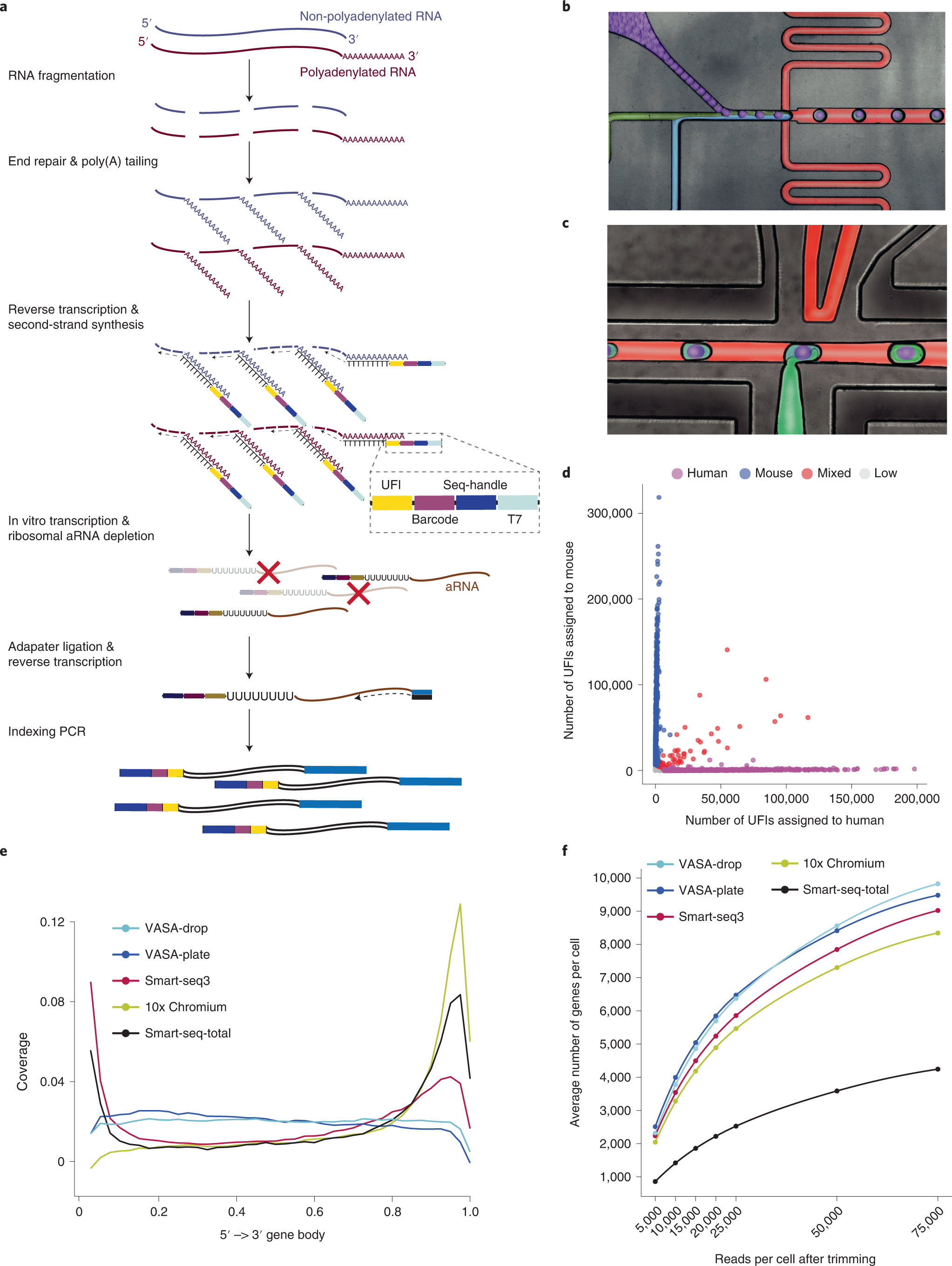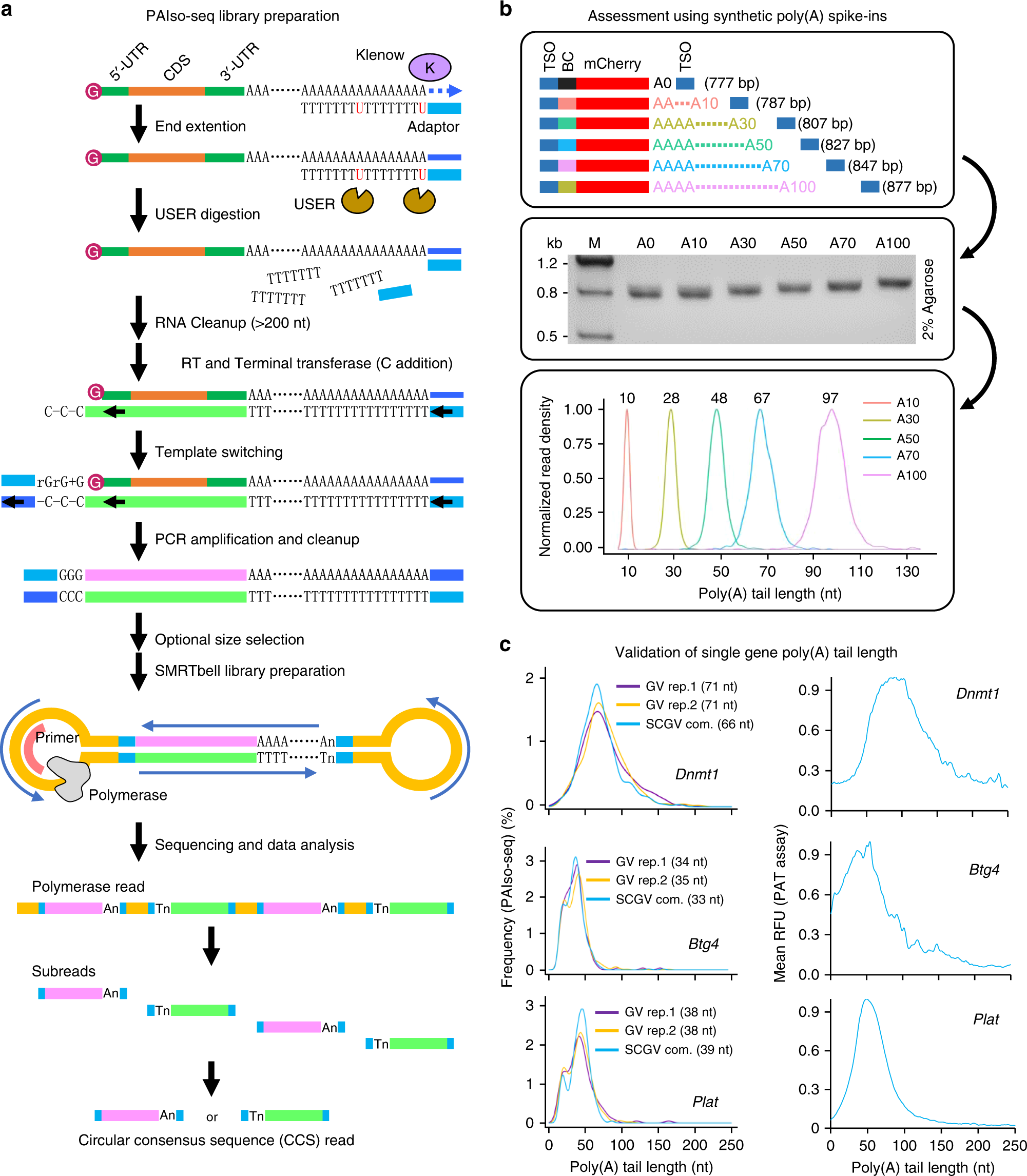
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails | Nature Communications
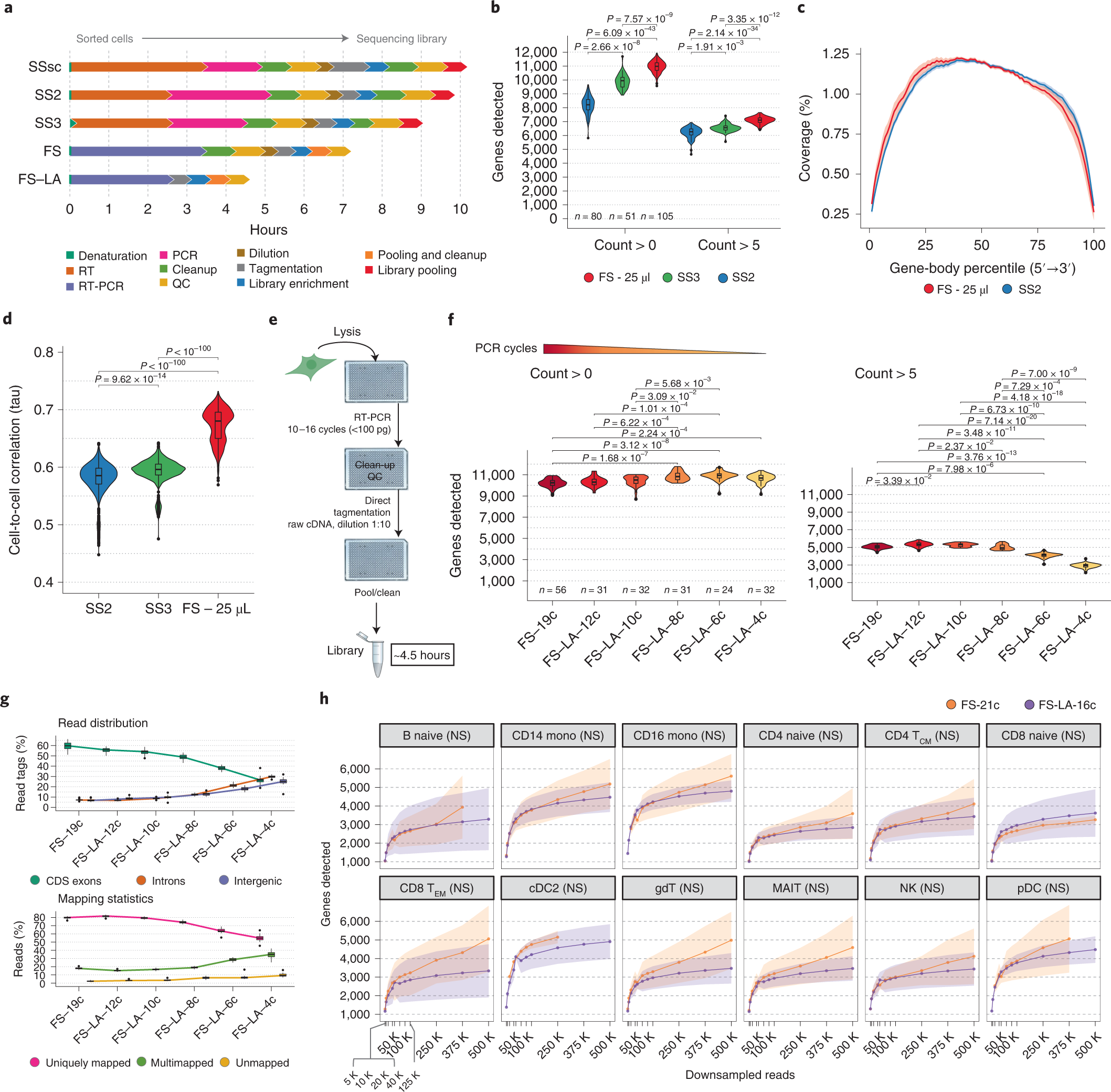
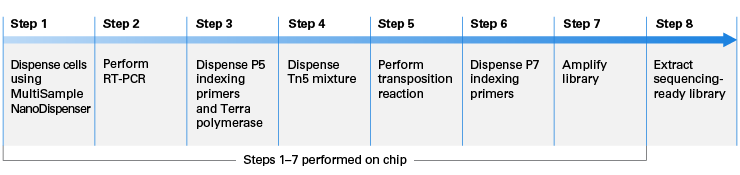
Fast and highly sensitive full-length single-cell RNA sequencing using FLASH-seq | Nature Biotechnology
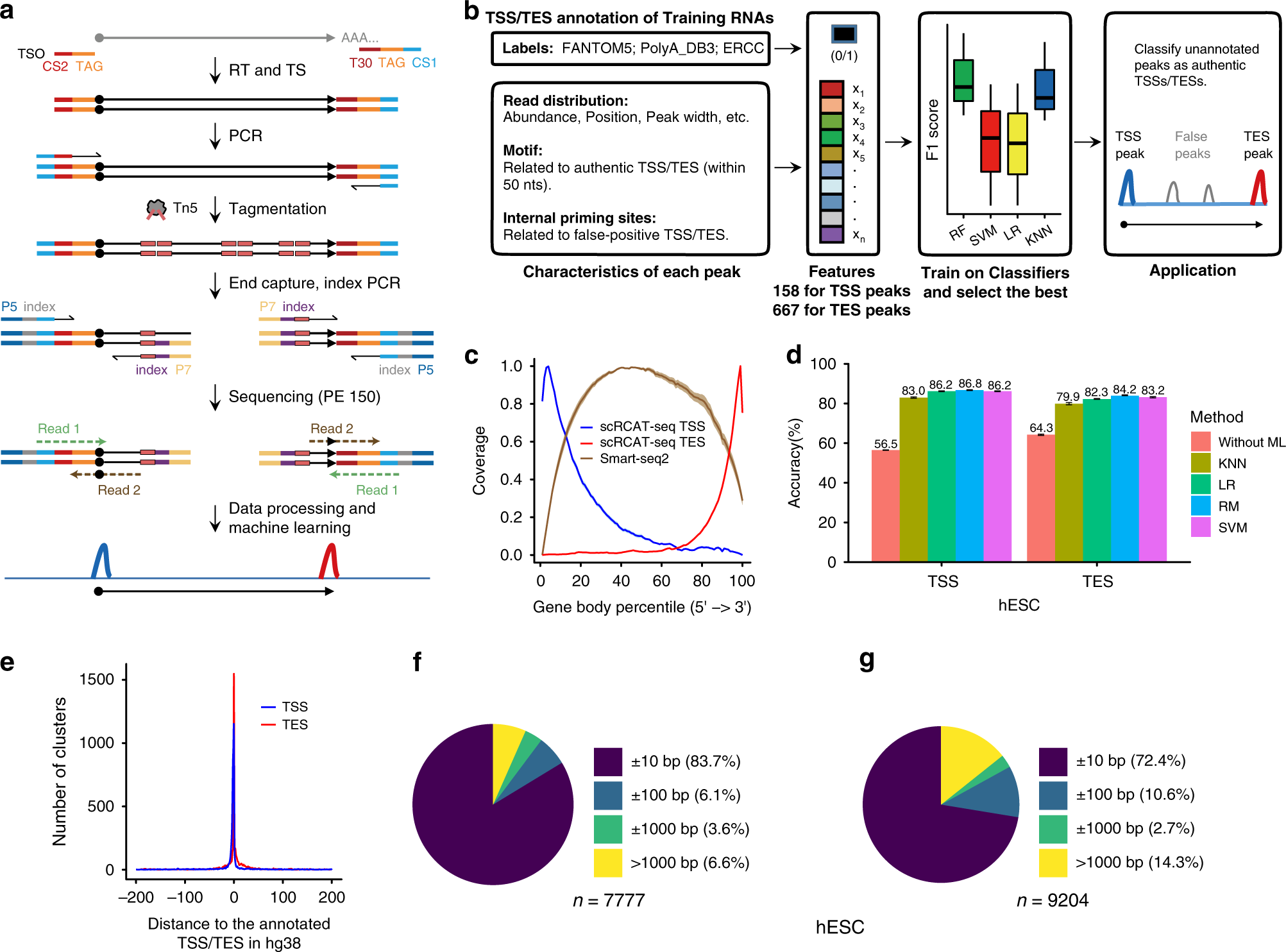
Single-cell RNA cap and tail sequencing (scRCAT-seq) reveals subtype-specific isoforms differing in transcript demarcation | Nature Communications

Full Length Isoform Sequencing (Iso-Seq) Yields a More Comprehensive View of Gene Activity - Drug Discovery World (DDW)
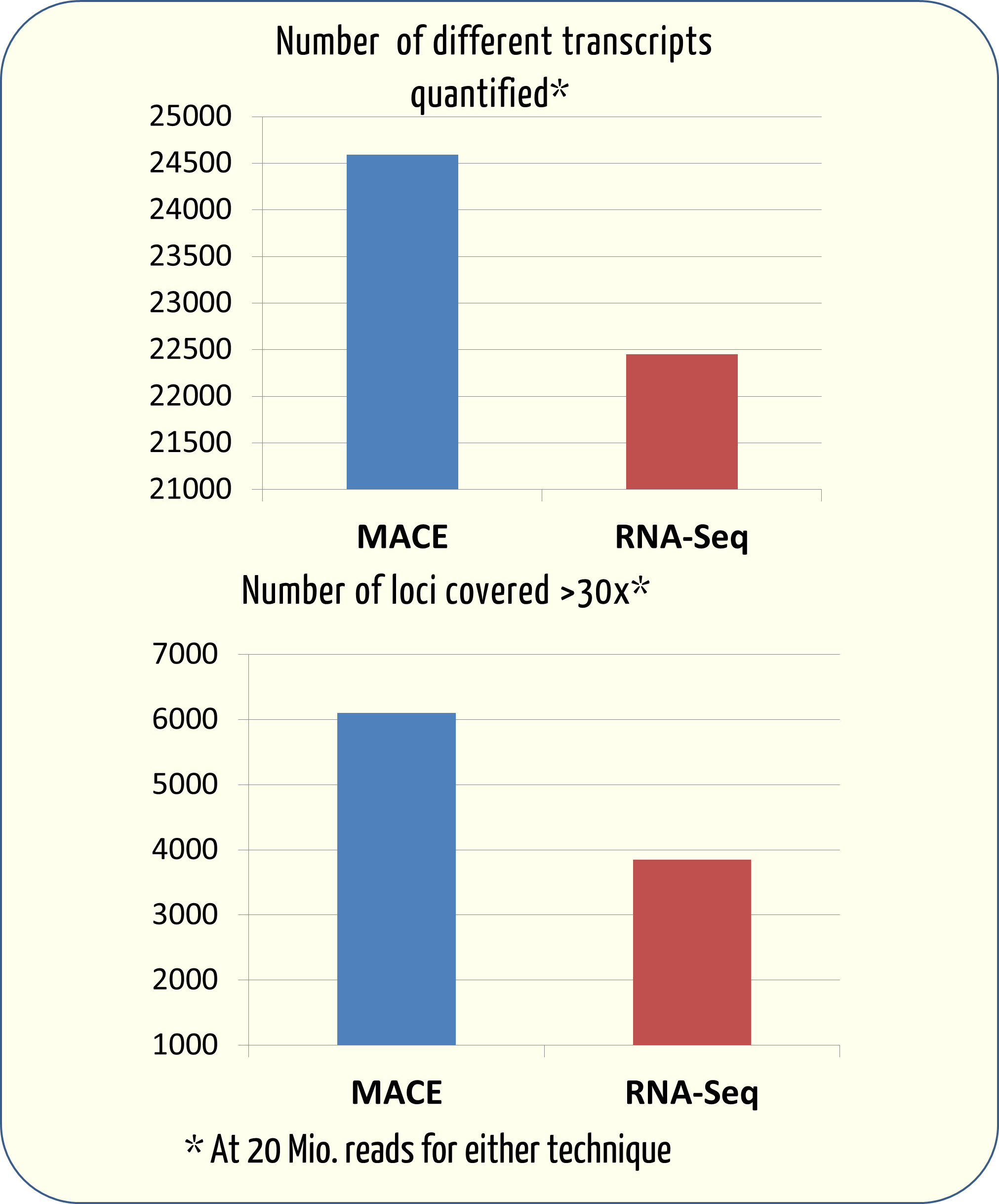
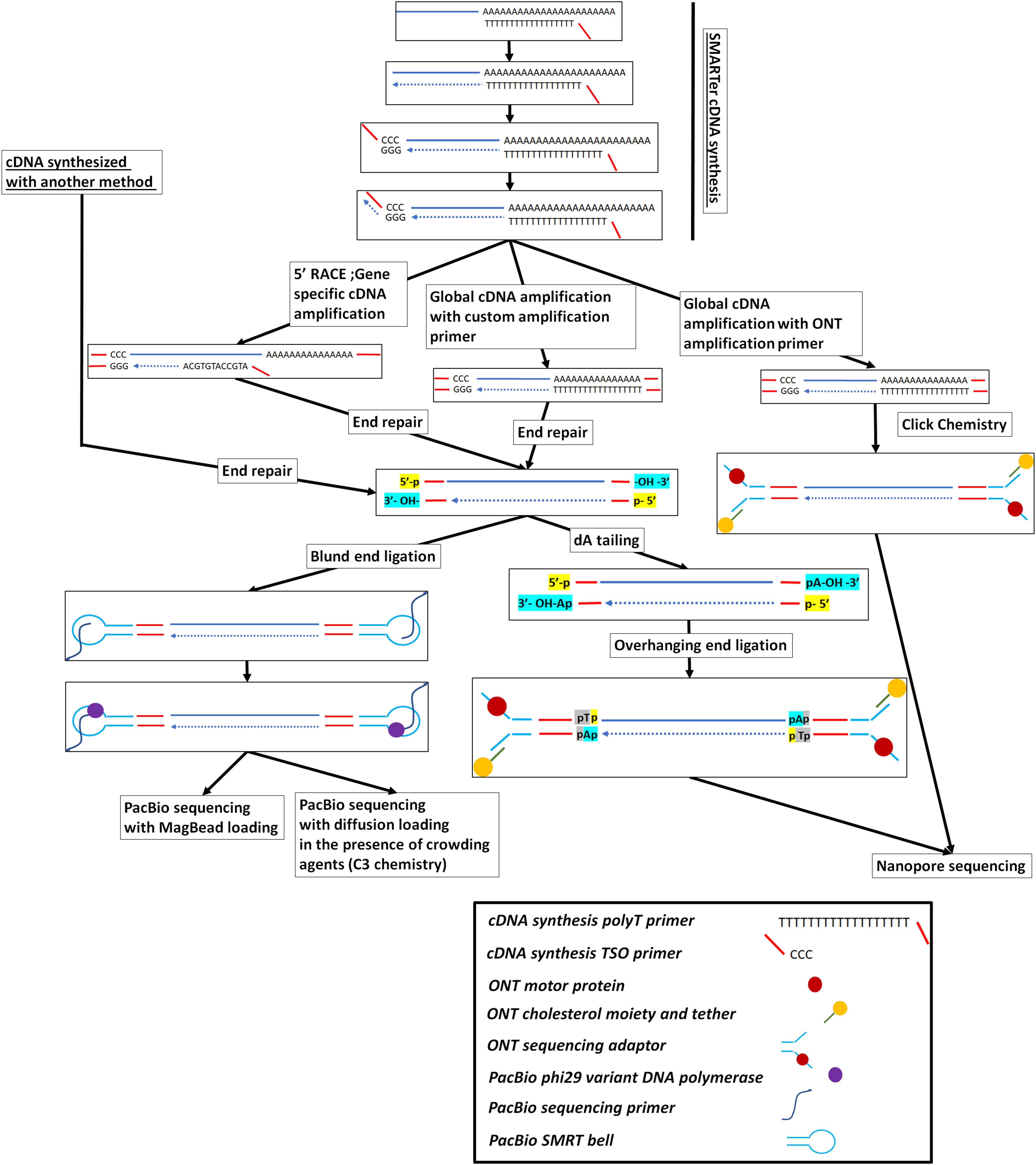
![PDF] Two methods for full-length RNA sequencing for low quantities of cells and single cells | Semantic Scholar PDF] Two methods for full-length RNA sequencing for low quantities of cells and single cells | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d24fdc377b399fa9e6863f728d756fab1b9390db/2-Figure1-1.png)
PDF] Two methods for full-length RNA sequencing for low quantities of cells and single cells | Semantic Scholar
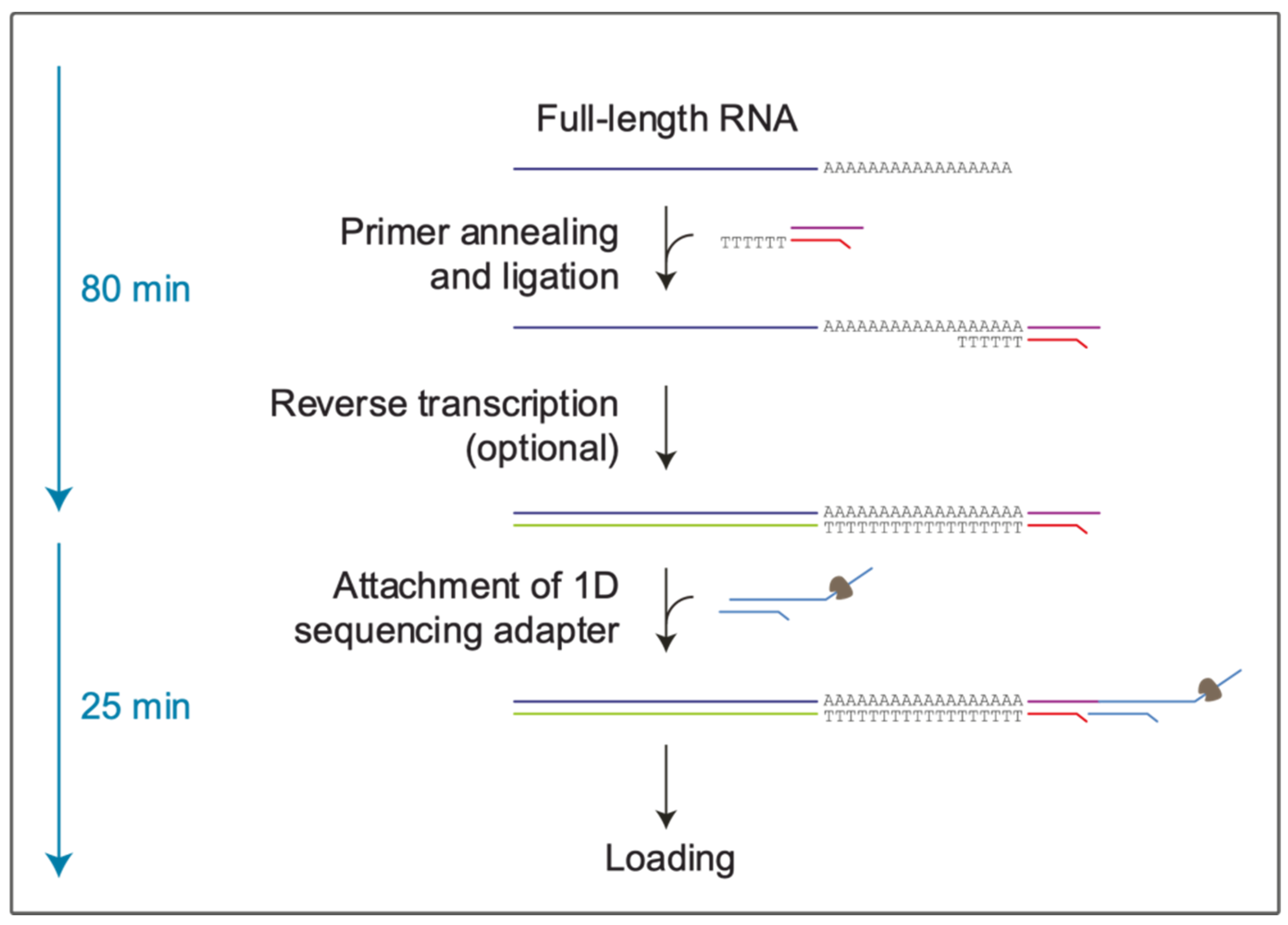
Life | Free Full-Text | Direct RNA Nanopore Sequencing of SARS-CoV-2 Extracted from Critical Material from Swabs
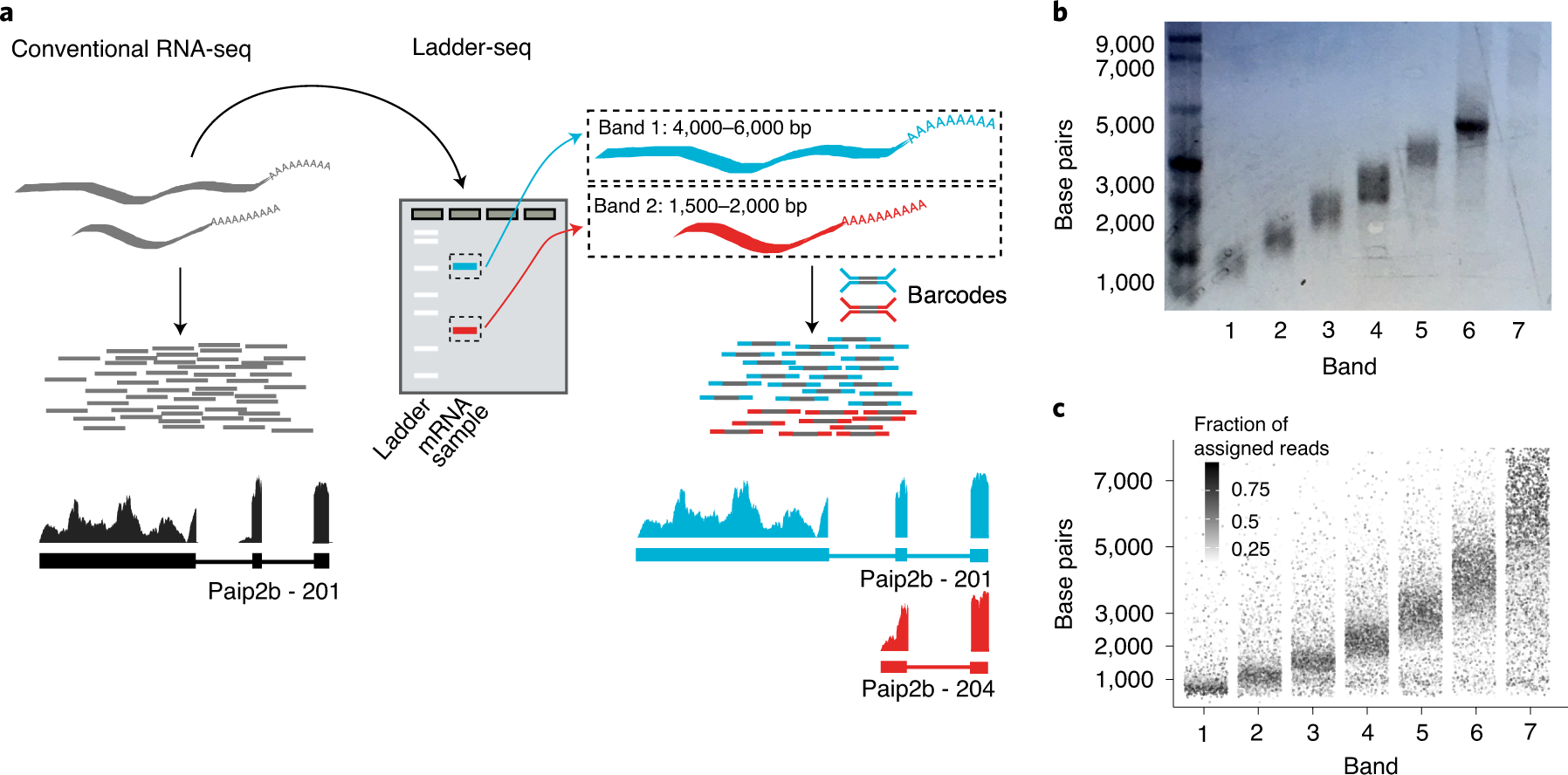
Partitioning RNAs by length improves transcriptome reconstruction from short-read RNA-seq data | Nature Biotechnology
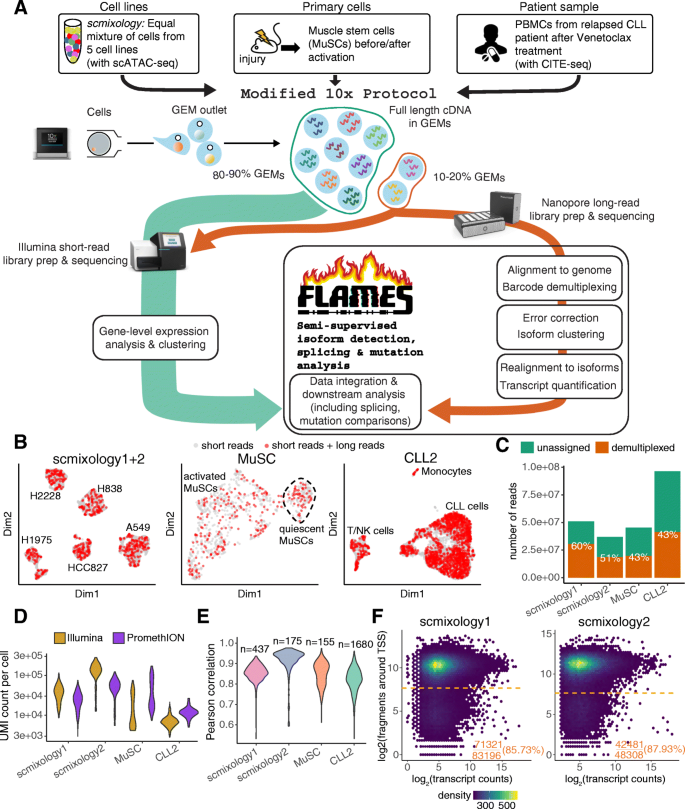
Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing | Genome Biology | Full Text
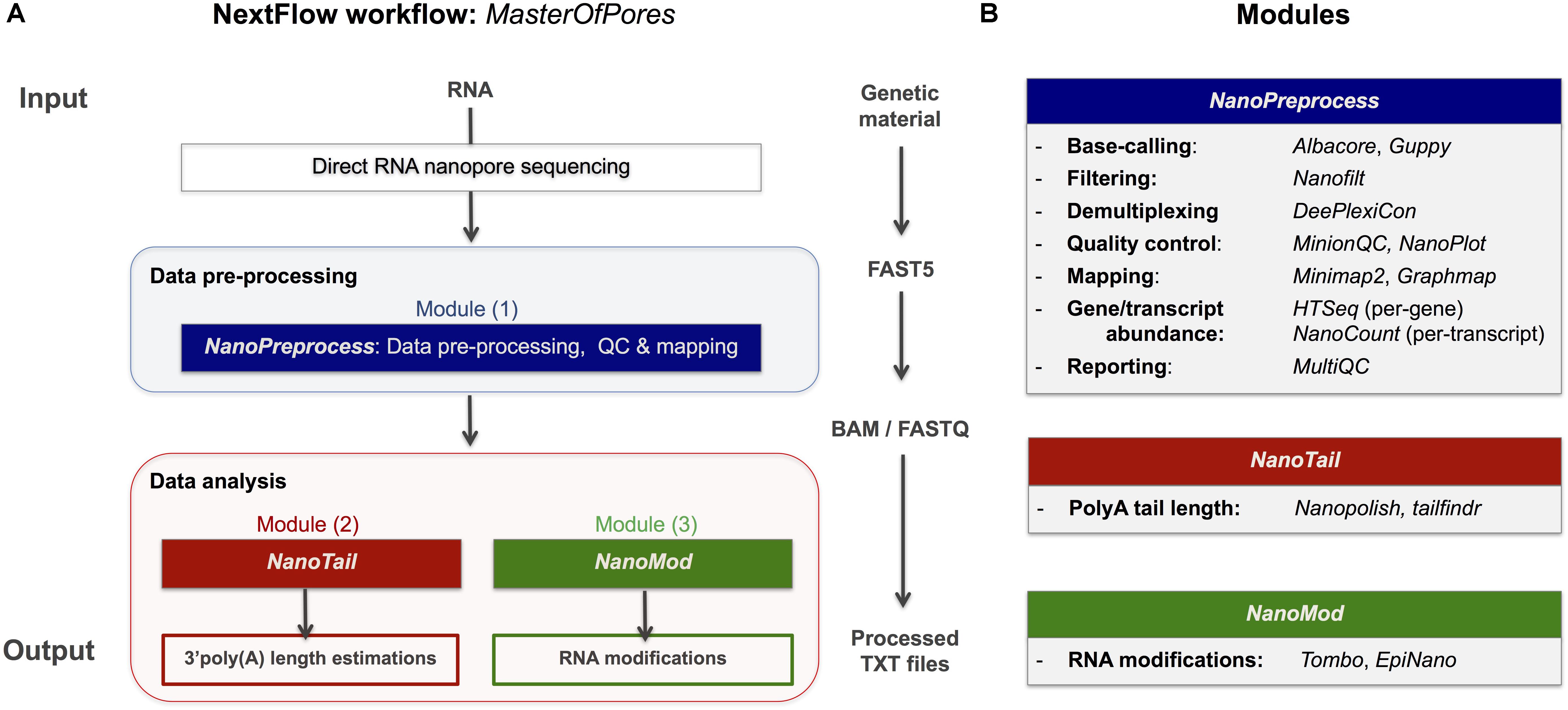
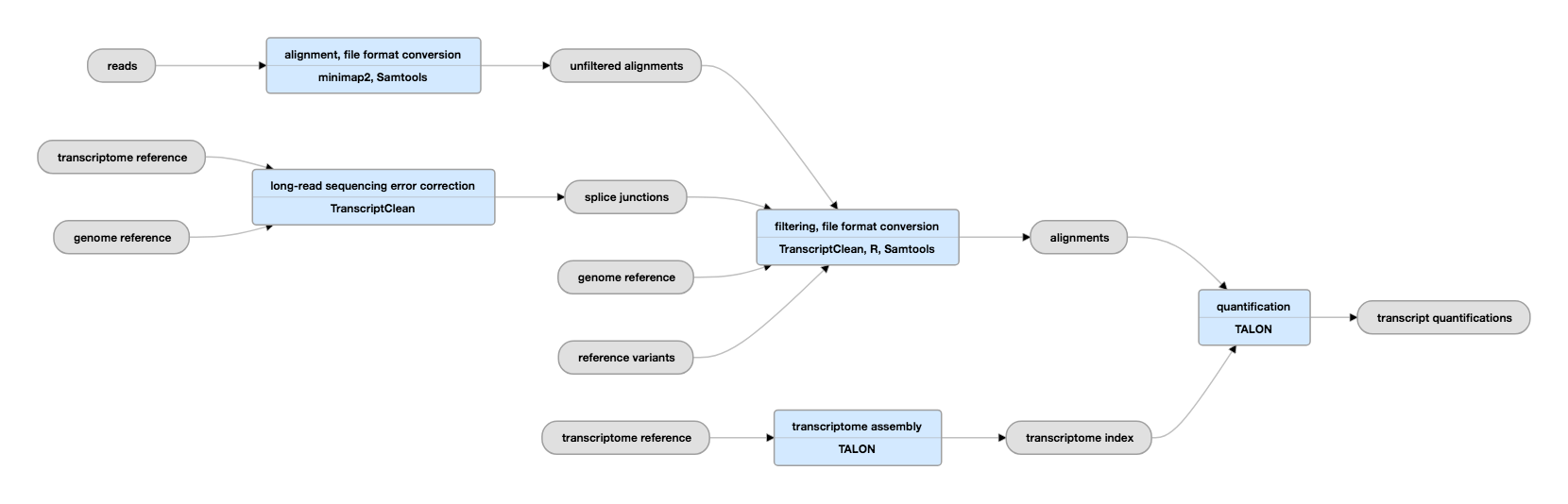
Frontiers | MasterOfPores: A Workflow for the Analysis of Oxford Nanopore Direct RNA Sequencing Datasets

cDNA approaches to measuring the transcriptome. Shown is full-length... | Download Scientific Diagram
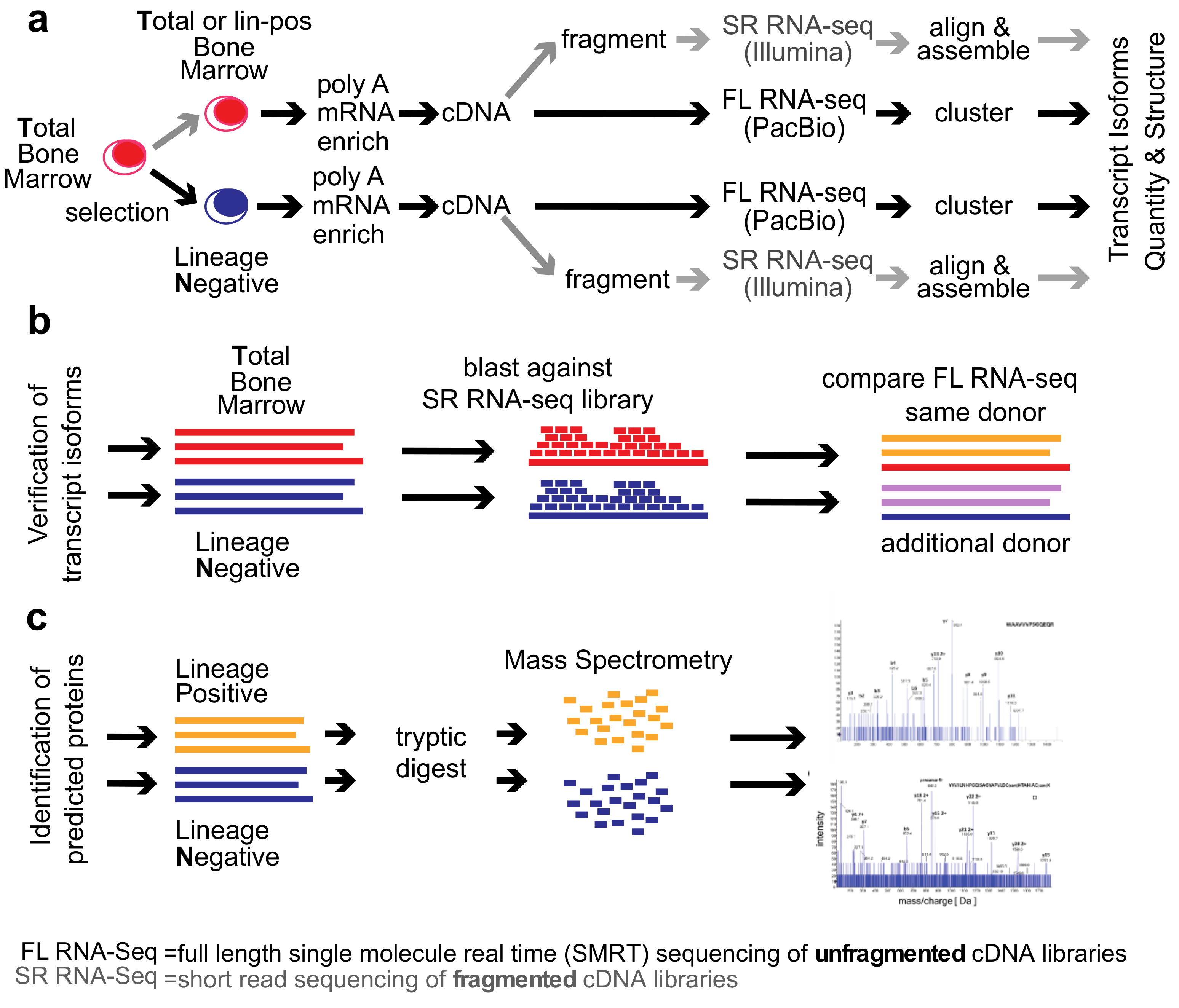
Genes | Free Full-Text | Single-Molecule Real-Time (SMRT) Full-Length RNA- Sequencing Reveals Novel and Distinct mRNA Isoforms in Human Bone Marrow Cell Subpopulations