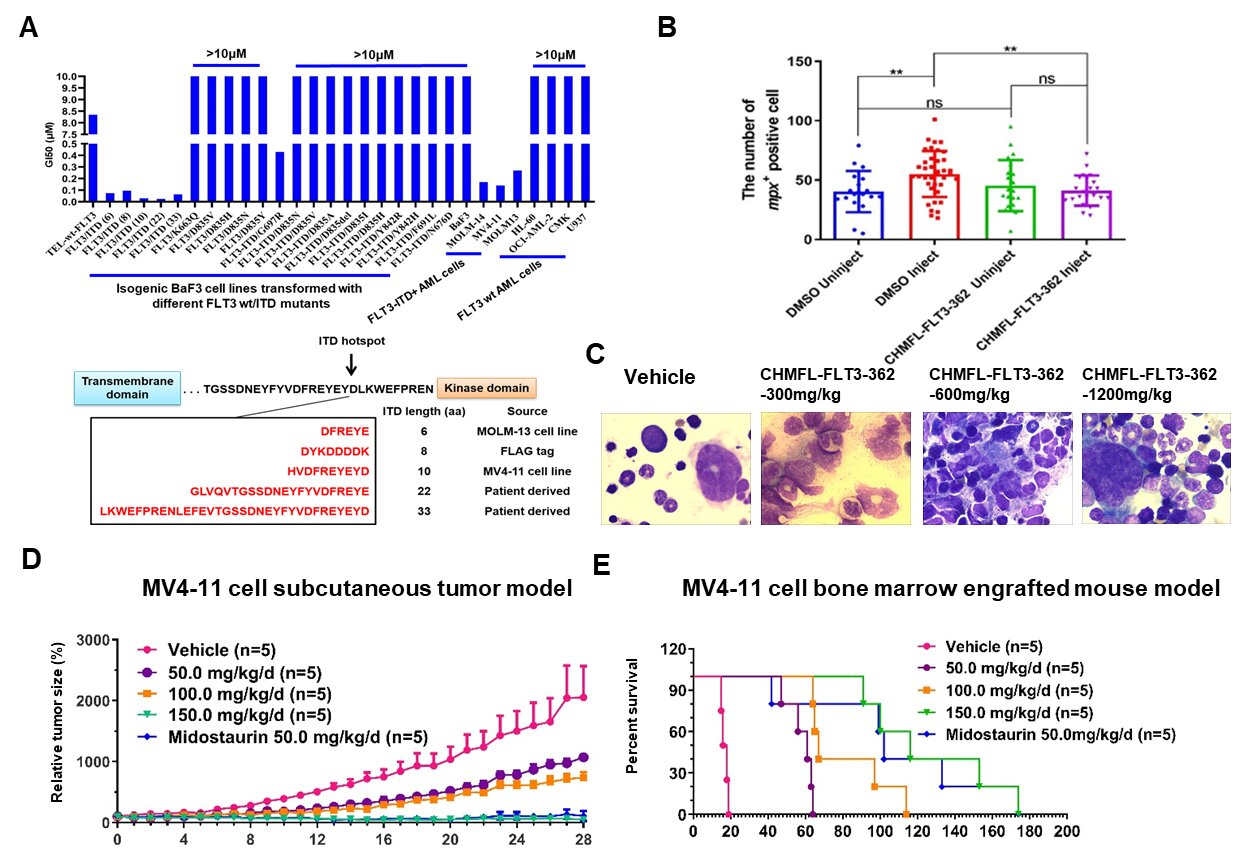
Clonal heterogeneity of FLT3-ITD detected by high-throughput amplicon sequencing correlates with adverse prognosis in acute myel

Detection of FLT3 Internal Tandem Duplication in Targeted, Short-Read-Length, Next-Generation Sequencing Data - ScienceDirect

Identification of emerging FLT3 ITD‐positive clones during clinical remission and kinetics of disease relapse in acute myeloid leukaemia with mutated nucleophosmin - Ottone - 2013 - British Journal of Haematology - Wiley Online Library

FLT3/D835Y mutation knock-in mice display less aggressive disease compared with FLT3/internal tandem duplication (ITD) mice | PNAS
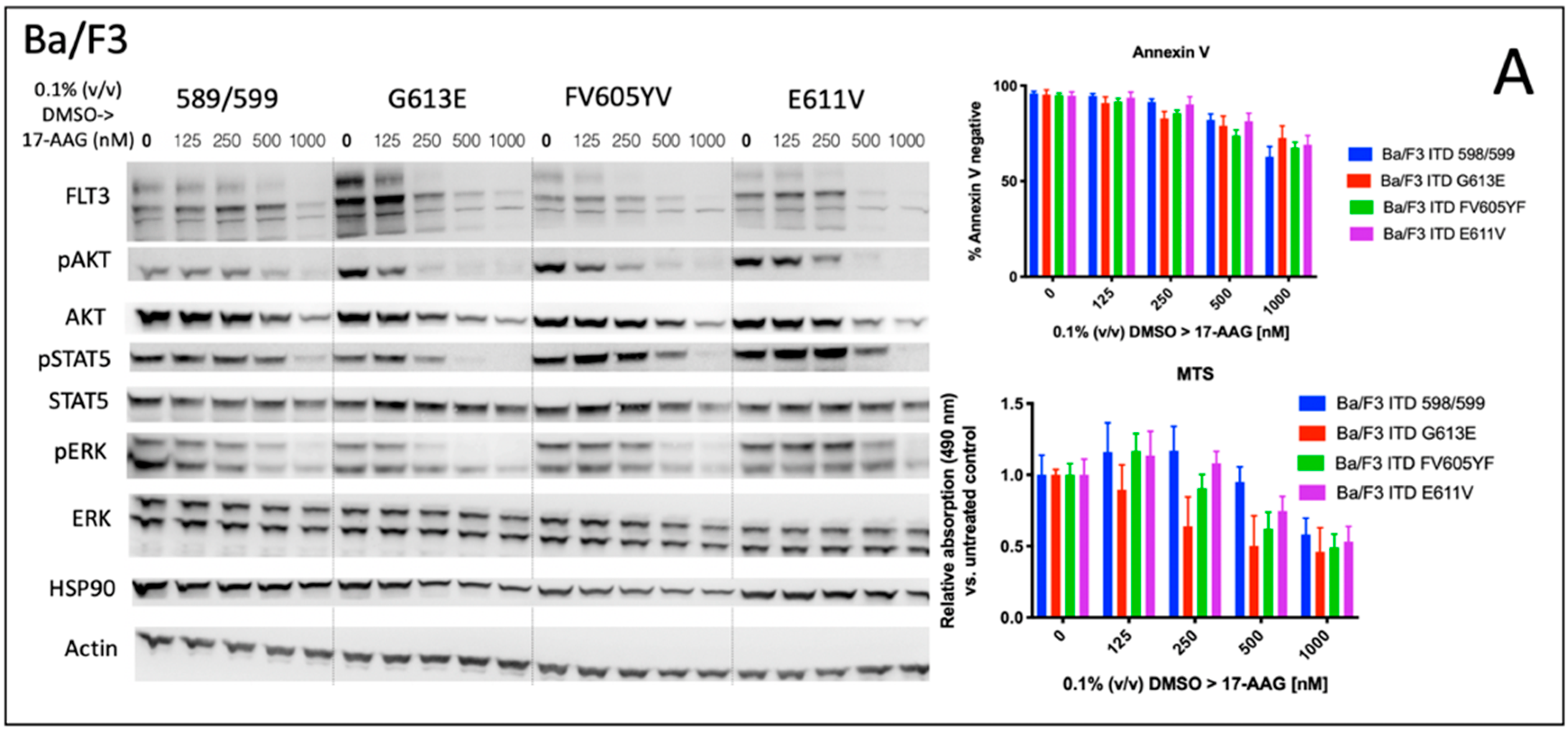
Cells | Free Full-Text | Modulation of FLT3-ITD Localization and Targeting of Distinct Downstream Signaling Pathways as Potential Strategies to Overcome FLT3-Inhibitor Resistance
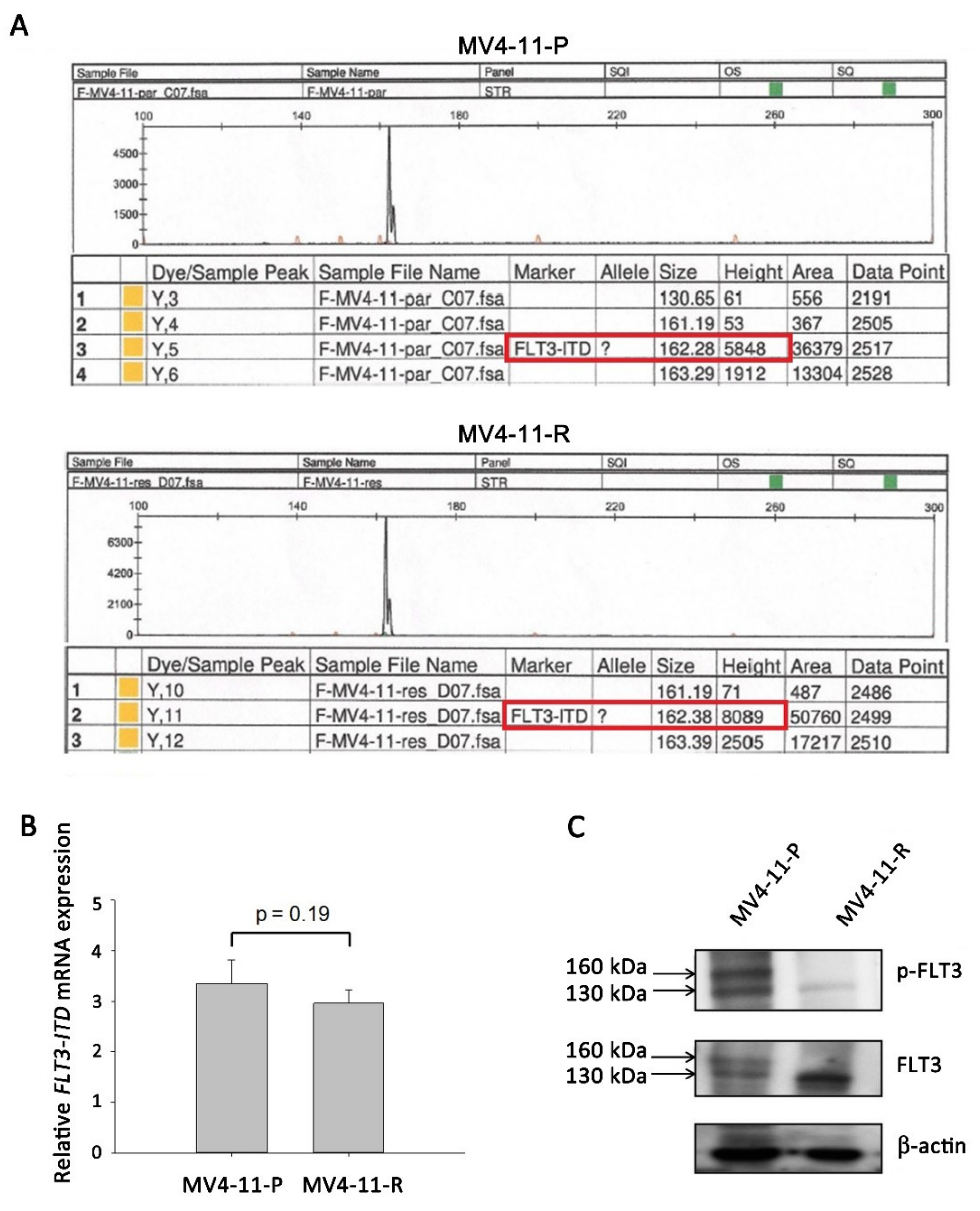
IJMS | Free Full-Text | Cytarabine-Resistant FLT3-ITD Leukemia Cells are Associated with TP53 Mutation and Multiple Pathway Alterations—Possible Therapeutic Efficacy of Cabozantinib
Clonal heterogeneity of FLT3 -ITD detected by high-throughput amplicon sequencing correlates with adverse prognosis in acute myeloid leukemia | Oncotarget
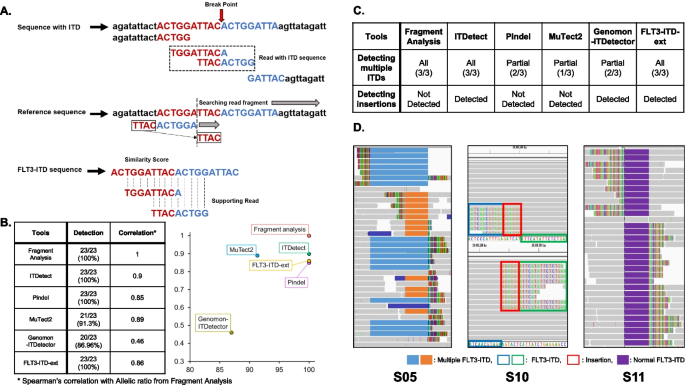
ITDetect: a method to detect internal tandem duplication of FMS-like tyrosine kinase (FLT3) from next-generation sequencing data with high sensitivity and clinical application | BMC Bioinformatics | Full Text
Clonal heterogeneity of FLT3 -ITD detected by high-throughput amplicon sequencing correlates with adverse prognosis in acute myeloid leukemia | Oncotarget

Prediction of HLA-binding peptides encoded by the FLT3-ITD sequence... | Download Scientific Diagram

FLT3-ITD, NPM1, and DNMT3A Gene Mutations and Risk Factors in Normal Karyotype Acute Myeloid Leukemia and Myelodysplastic Syndrome Patients in Upper Northern Thailand - Abstract - Europe PMC
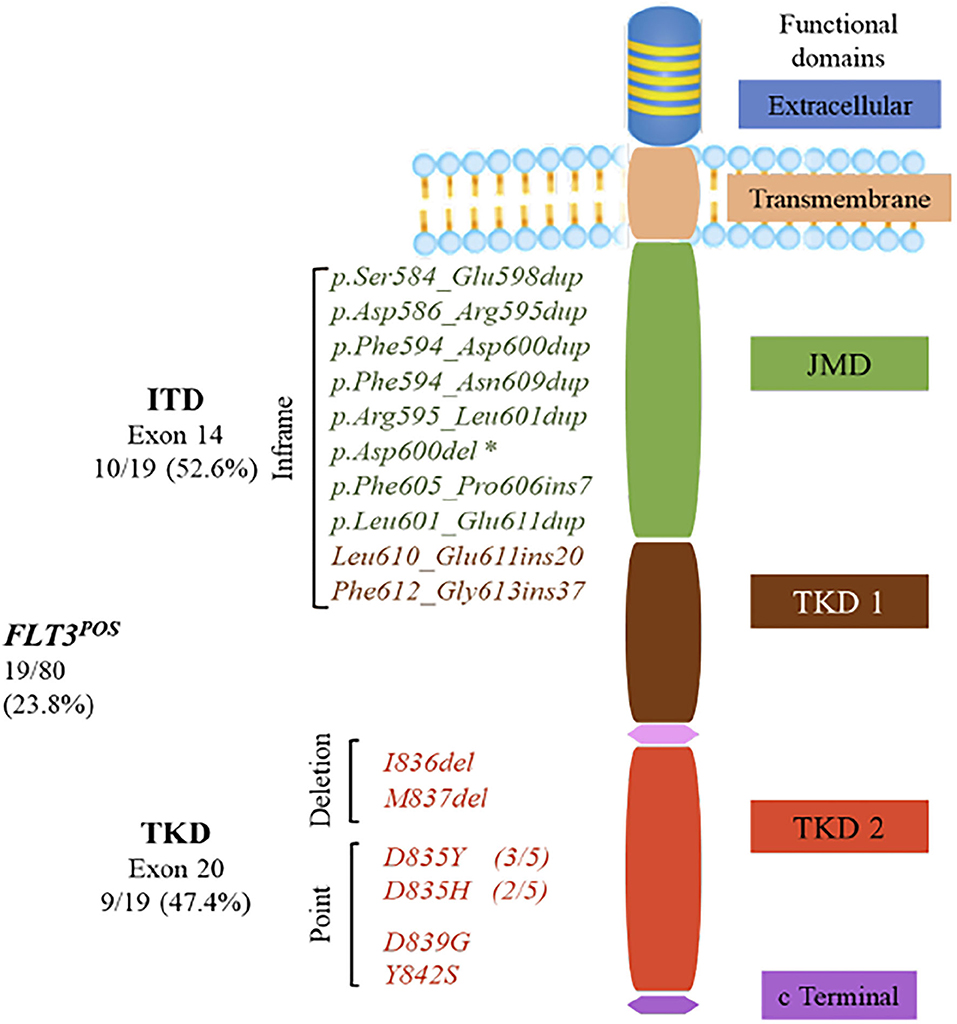
Frontiers | Profiling FLT3 Mutations in Mexican Acute Myeloid Leukemia Pediatric Patients: Impact on Overall Survival

Figure 4 from Detection of FLT3 (FMS-like Tyrosine Kinase) Internal Tandem Duplication (ITD) Mutation using Next Generation Sequencing Technology and Nested PCR | Semantic Scholar
![PDF] FLT3-ITD, NPM1, and DNMT3A Gene Mutations and Risk Factors in Normal Karyotype Acute Myeloid Leukemia and Myelodysplastic Syndrome Patients in Upper Northern Thailand | Semantic Scholar PDF] FLT3-ITD, NPM1, and DNMT3A Gene Mutations and Risk Factors in Normal Karyotype Acute Myeloid Leukemia and Myelodysplastic Syndrome Patients in Upper Northern Thailand | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c42682134066e9ecb7be4d79b2d585c516877b2d/3-Figure1-1.png)