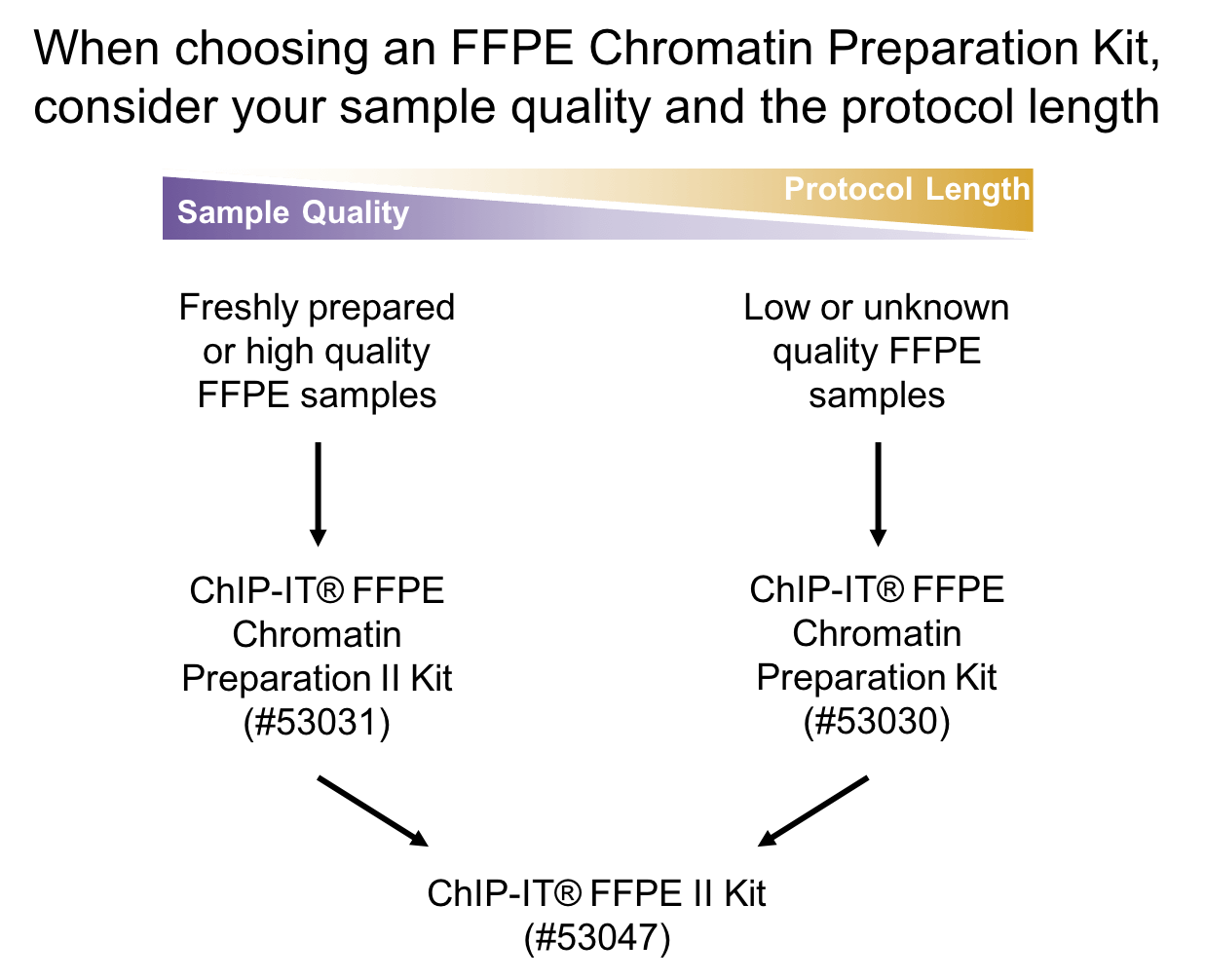
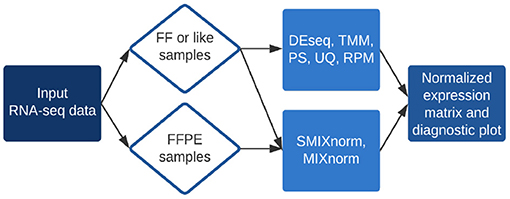
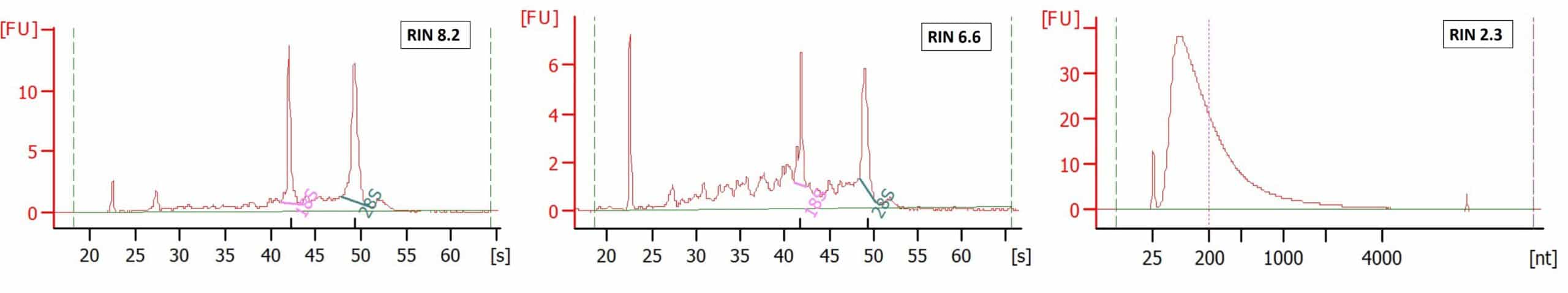
Processing FFPE samples for the TempO-Seq assay. (A) Examples of slides... | Download Scientific Diagram

Correlation of RNA-seq for FF and FFPE tumors. A) FFPE vs. FF, tumor 1,... | Download Scientific Diagram
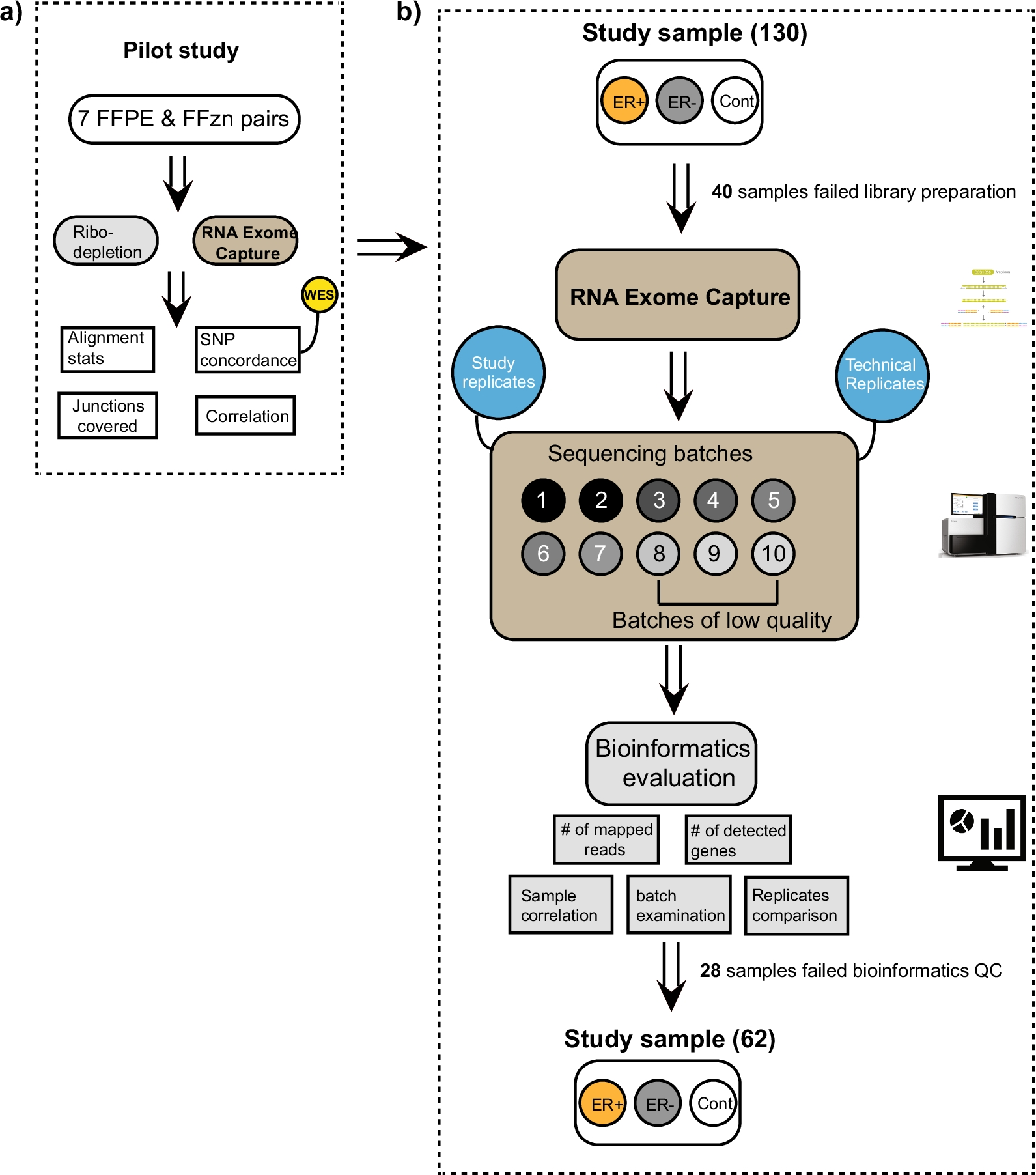
A novel method for RNA extraction from FFPE samples reveals significant differences in biomarker expression between orthotopic and subcutaneous pancreatic... | Oncotarget
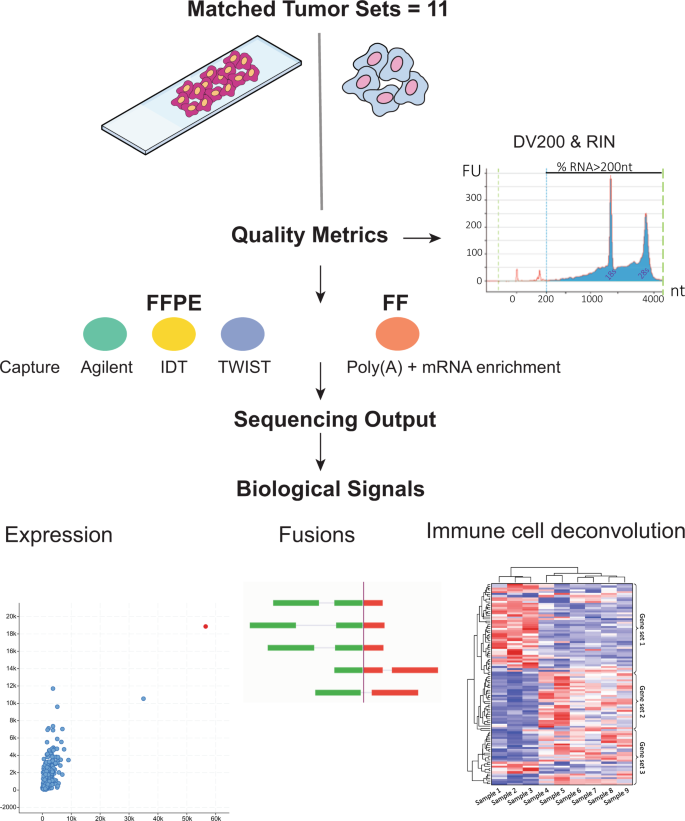
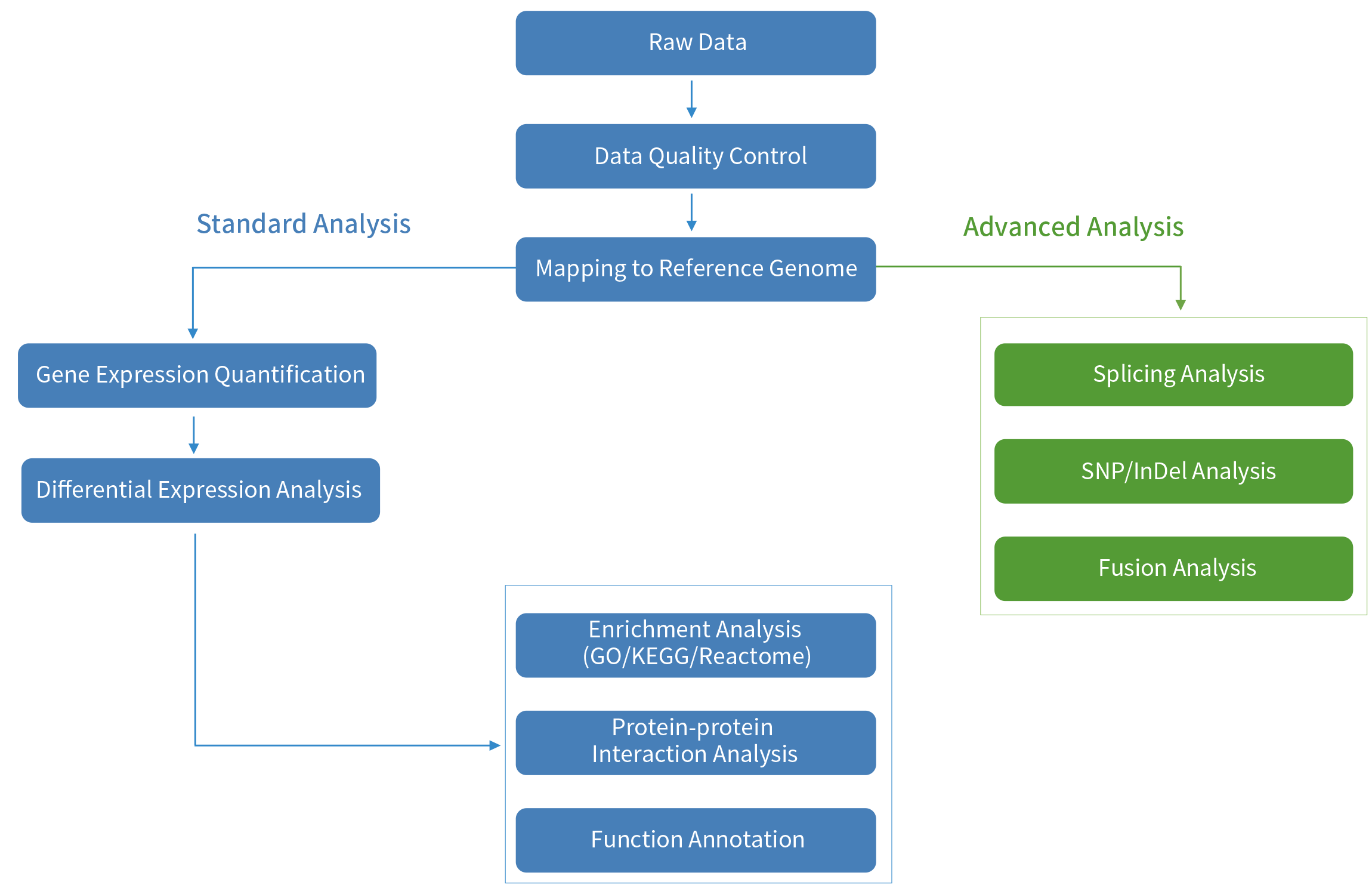
Functional comparison of exome capture-based methods for transcriptomic profiling of formalin-fixed paraffin-embedded tumors | npj Genomic Medicine
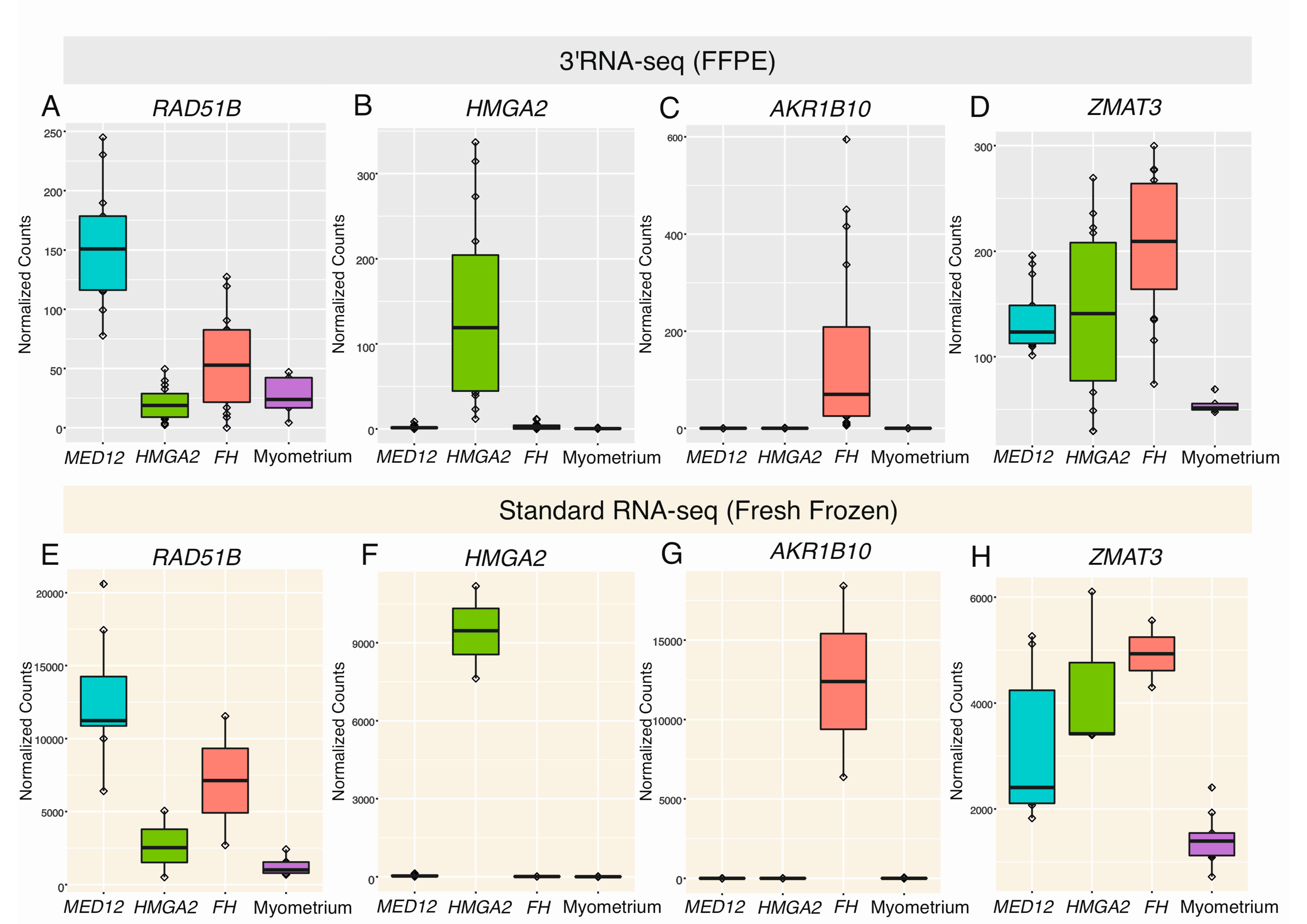
Cancers | Free Full-Text | 3′RNA Sequencing Accurately Classifies Formalin-Fixed Paraffin-Embedded Uterine Leiomyomas

Isolation of RNA from FF and FFPE tissues and generation of RNA-Seq... | Download Scientific Diagram
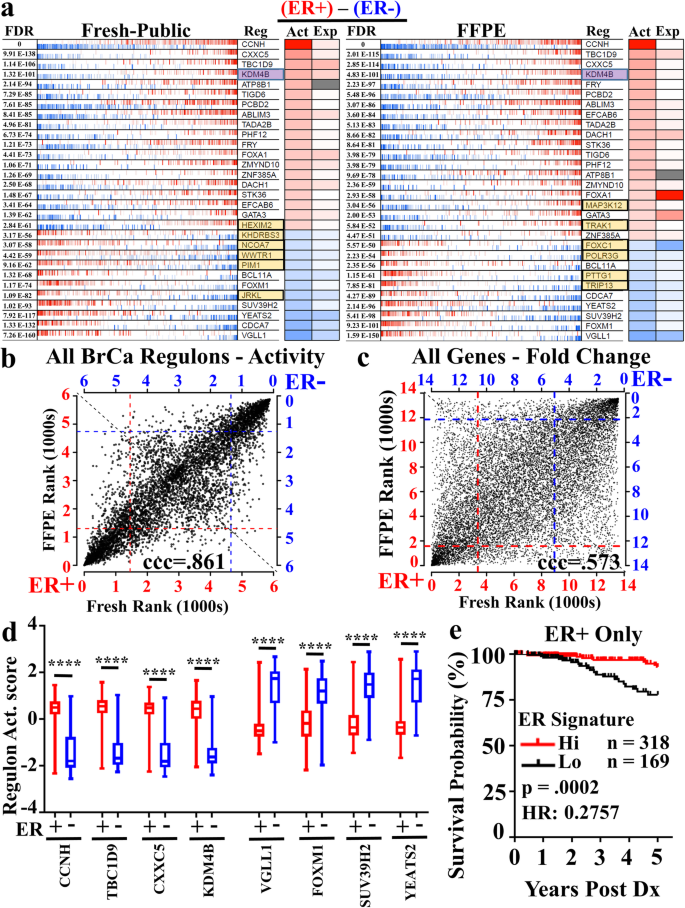
RNA-seq from archival FFPE breast cancer samples: molecular pathway fidelity and novel discovery | BMC Medical Genomics | Full Text

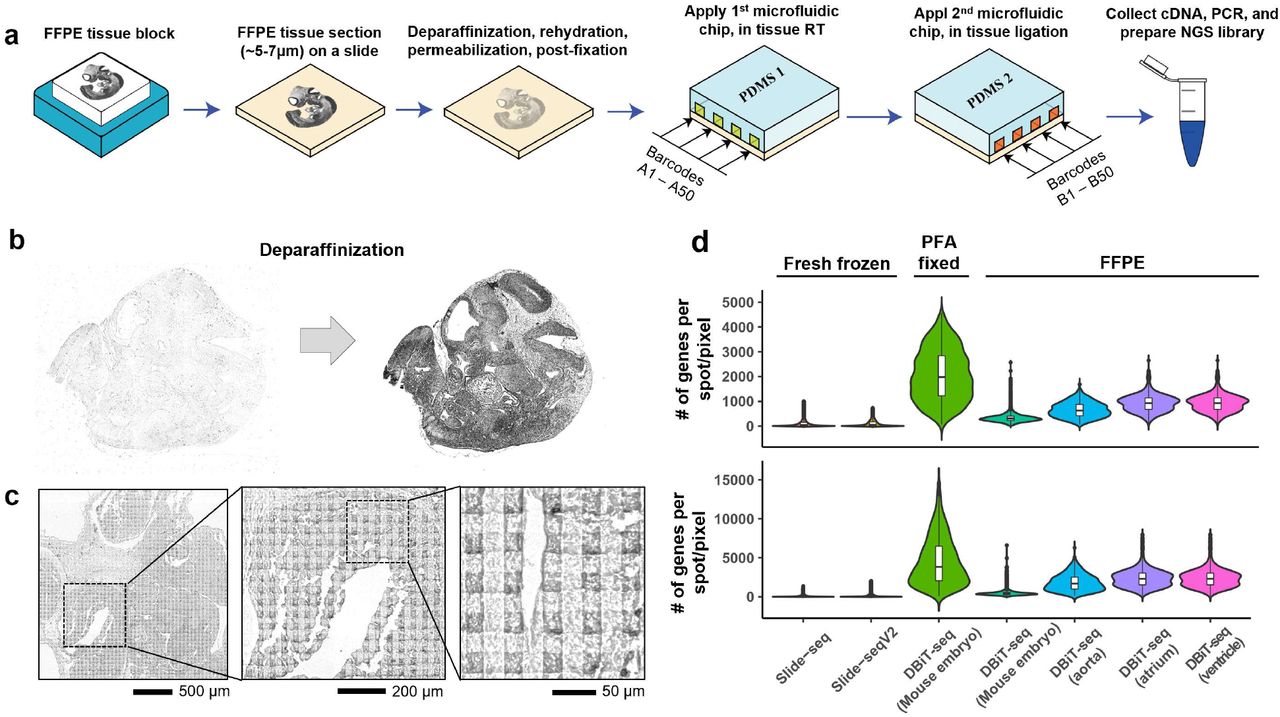
Reproducible and sensitive micro-tissue RNA sequencing from formalin-fixed paraffin-embedded tissues for spatial gene expression analysis | Scientific Reports
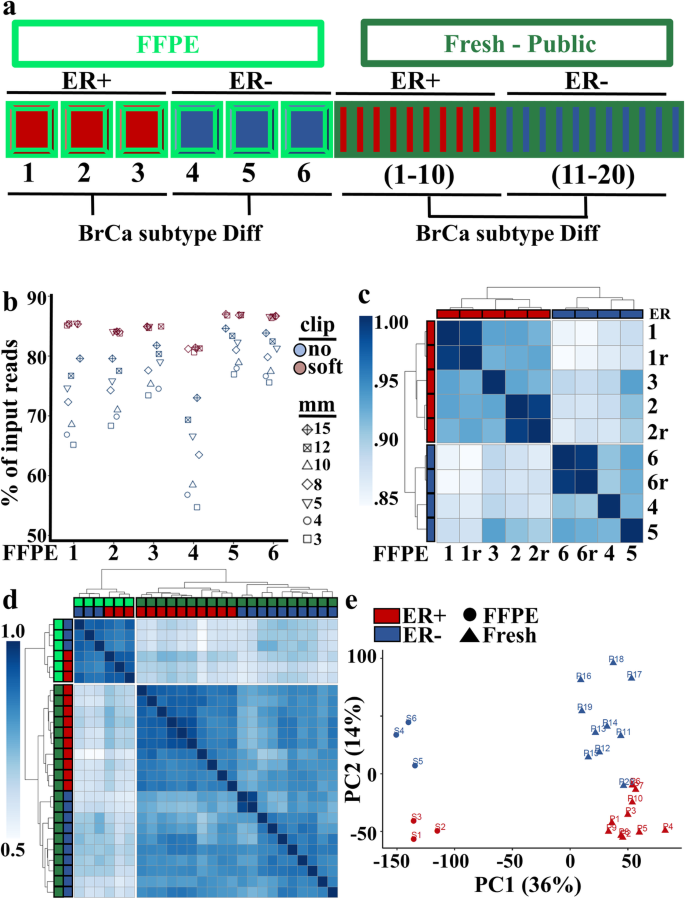
RNA-seq from archival FFPE breast cancer samples: molecular pathway fidelity and novel discovery | BMC Medical Genomics | Full Text

Enriched transcriptome analysis of laser capture microdissected populations of single cells to investigate intracellular heterogeneity in immunostained FFPE sections - ScienceDirect
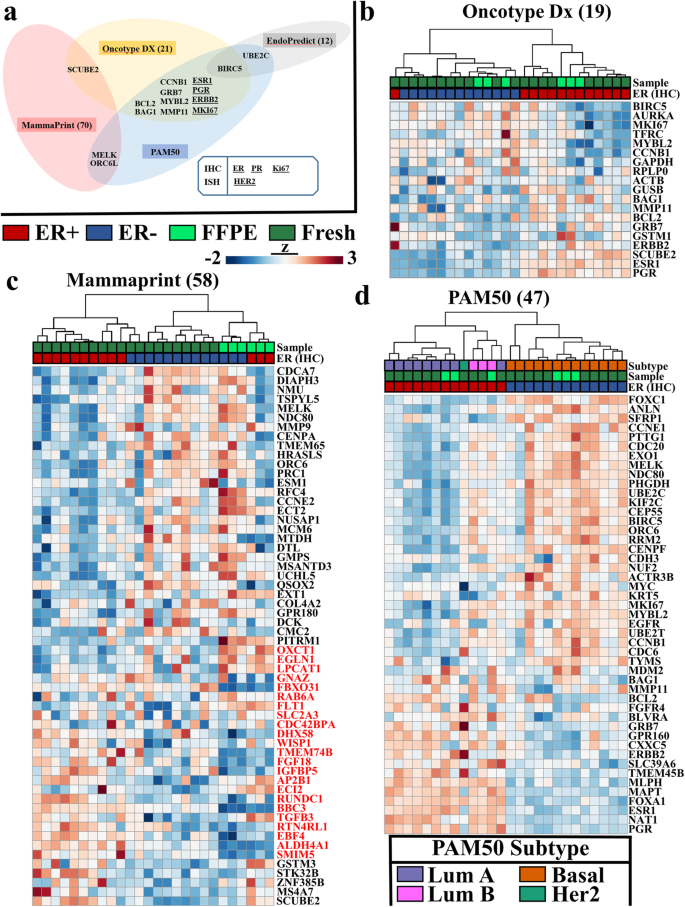
RNA-seq from archival FFPE breast cancer samples: molecular pathway fidelity and novel discovery | BMC Medical Genomics | Full Text
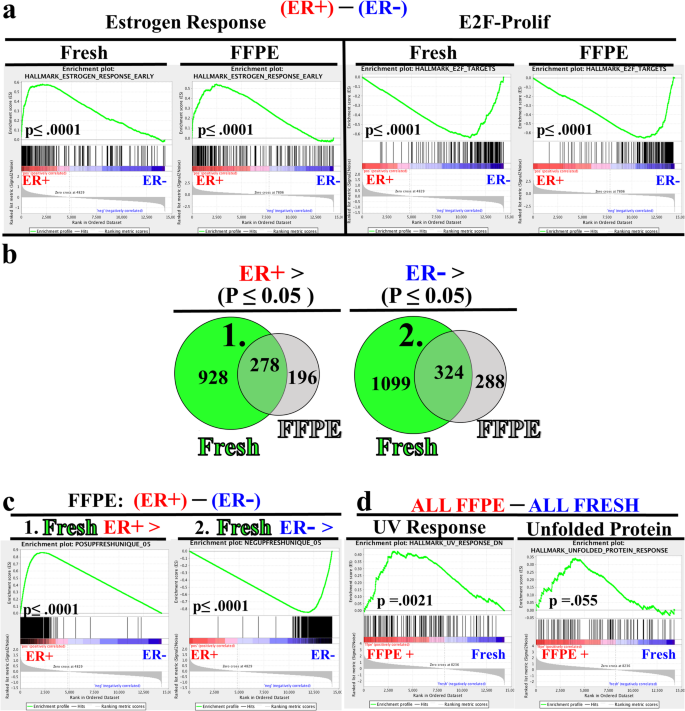
RNA-seq from archival FFPE breast cancer samples: molecular pathway fidelity and novel discovery | BMC Medical Genomics | Full Text

Large scale, robust, and accurate whole transcriptome profiling from clinical formalin-fixed paraffin-embedded samples | Scientific Reports
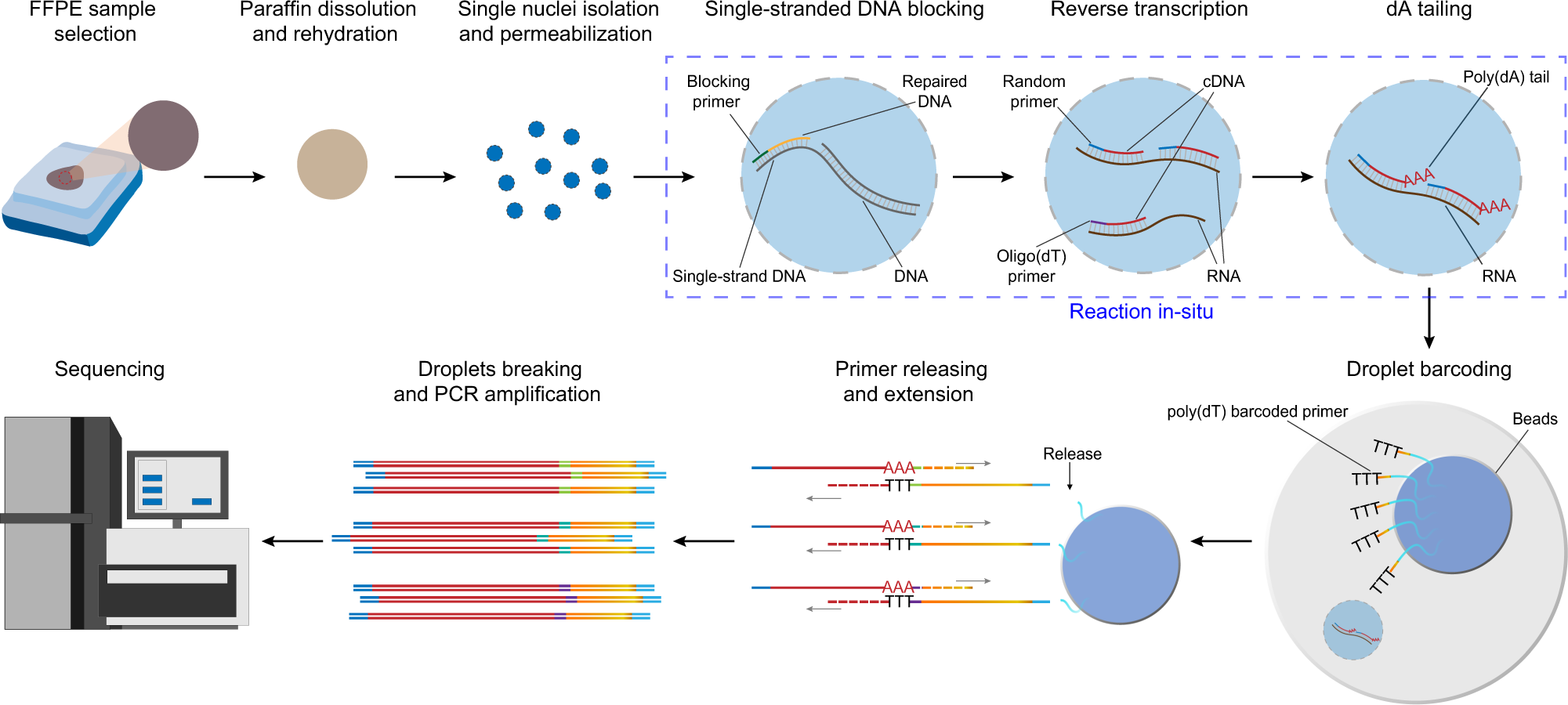
High-throughput single nucleus total RNA sequencing of formalin-fixed paraffin-embedded tissues by snRandom-seq | Nature Communications