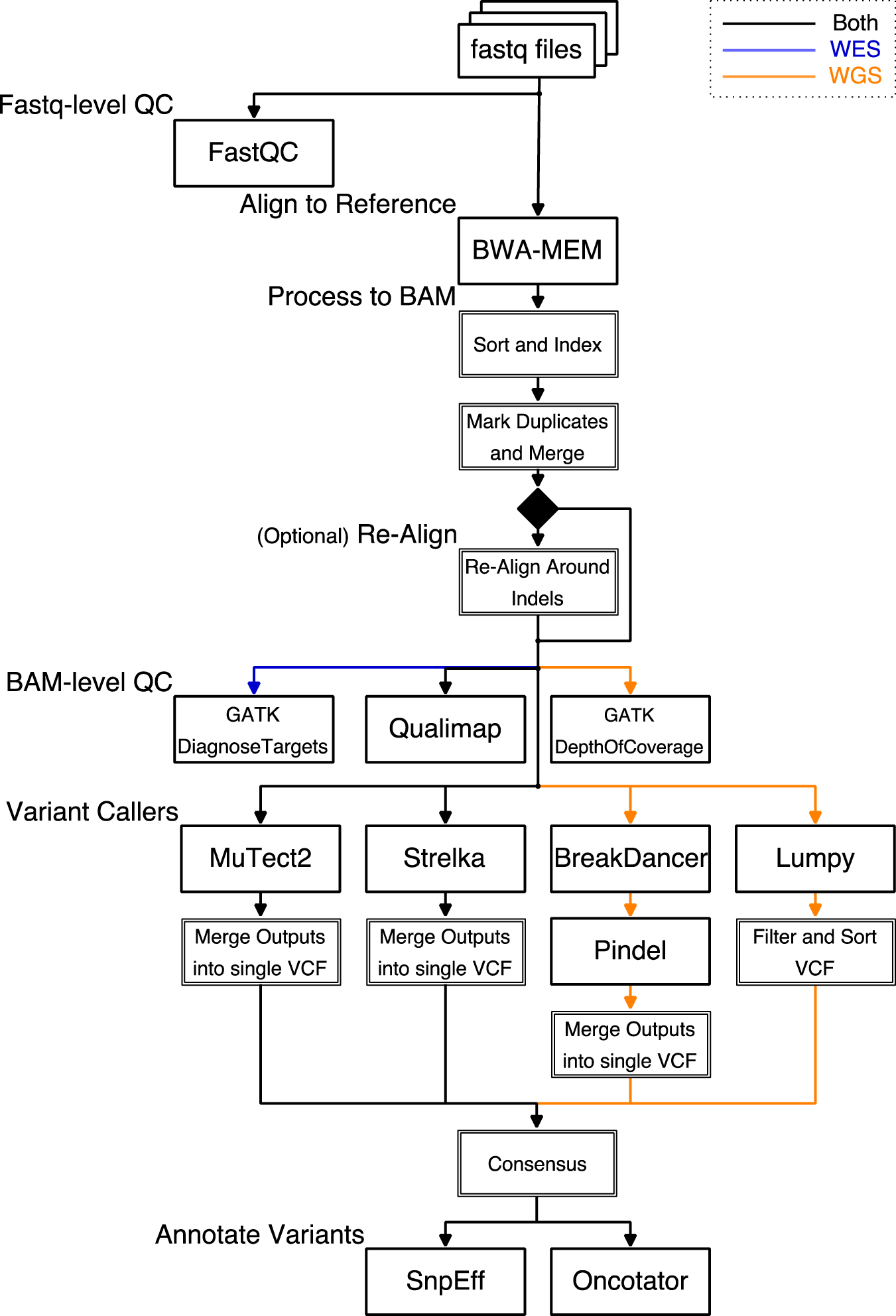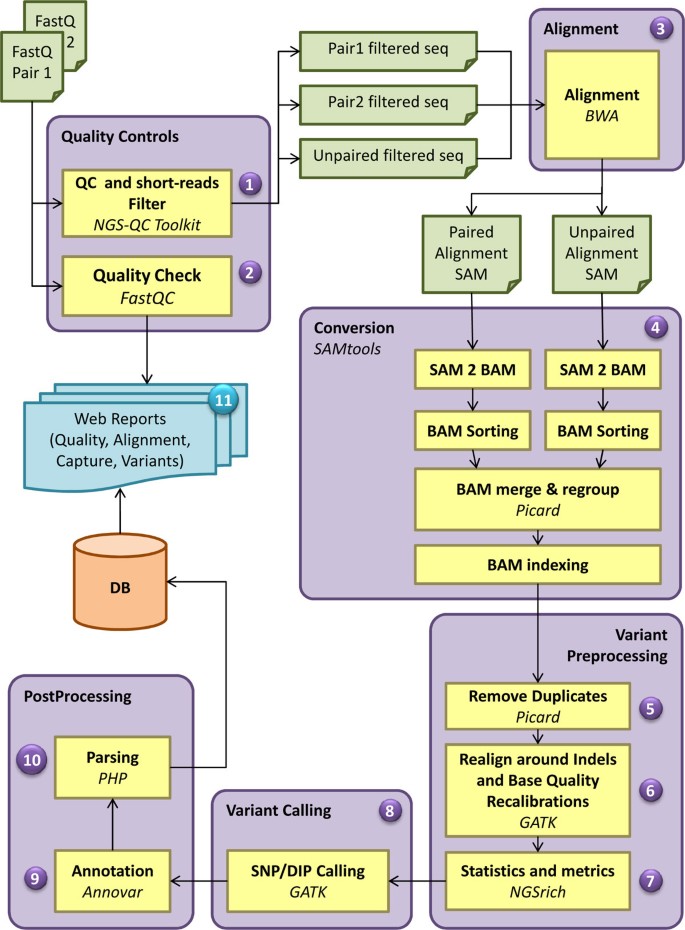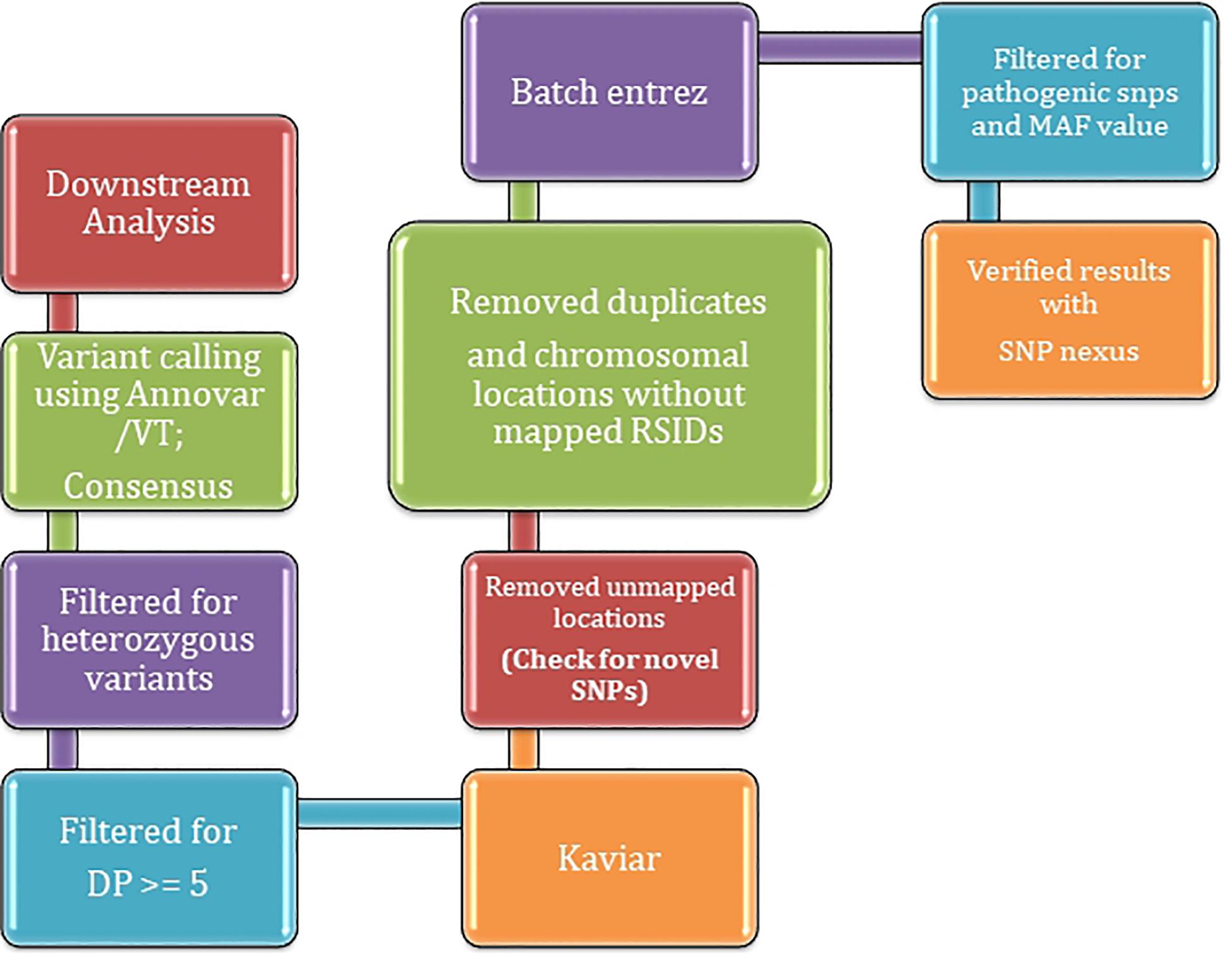
Frontiers | A Pilot Study on the Whole Exome Sequencing of Prostate Cancer in the Indian Phenotype Reveals Distinct Polymorphisms
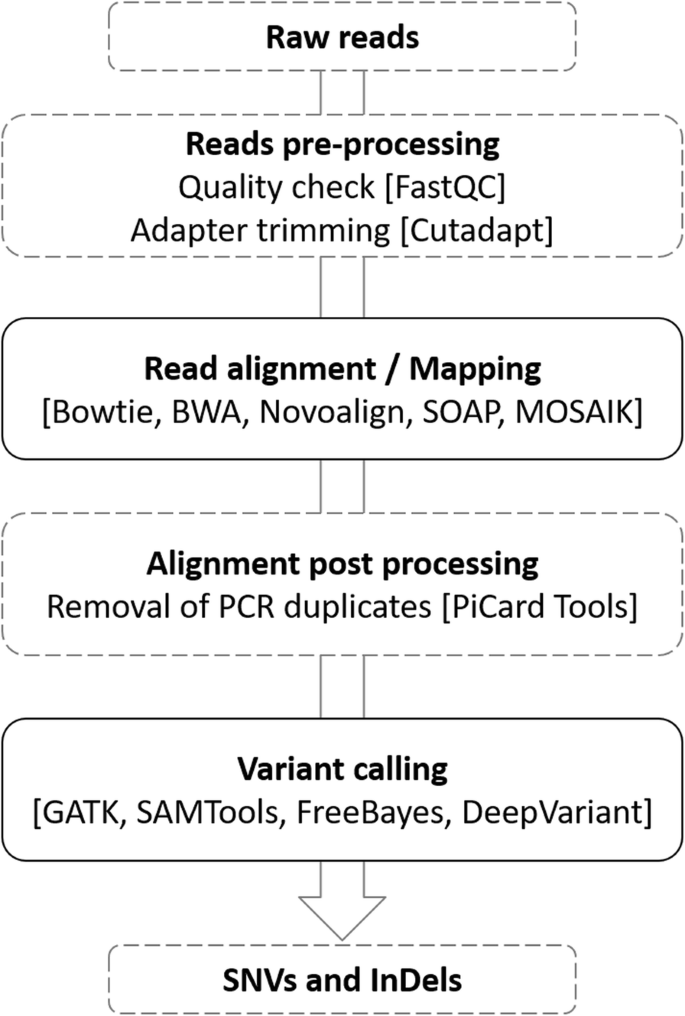
Performance assessment of variant calling pipelines using human whole exome sequencing and simulated data | BMC Bioinformatics | Full Text
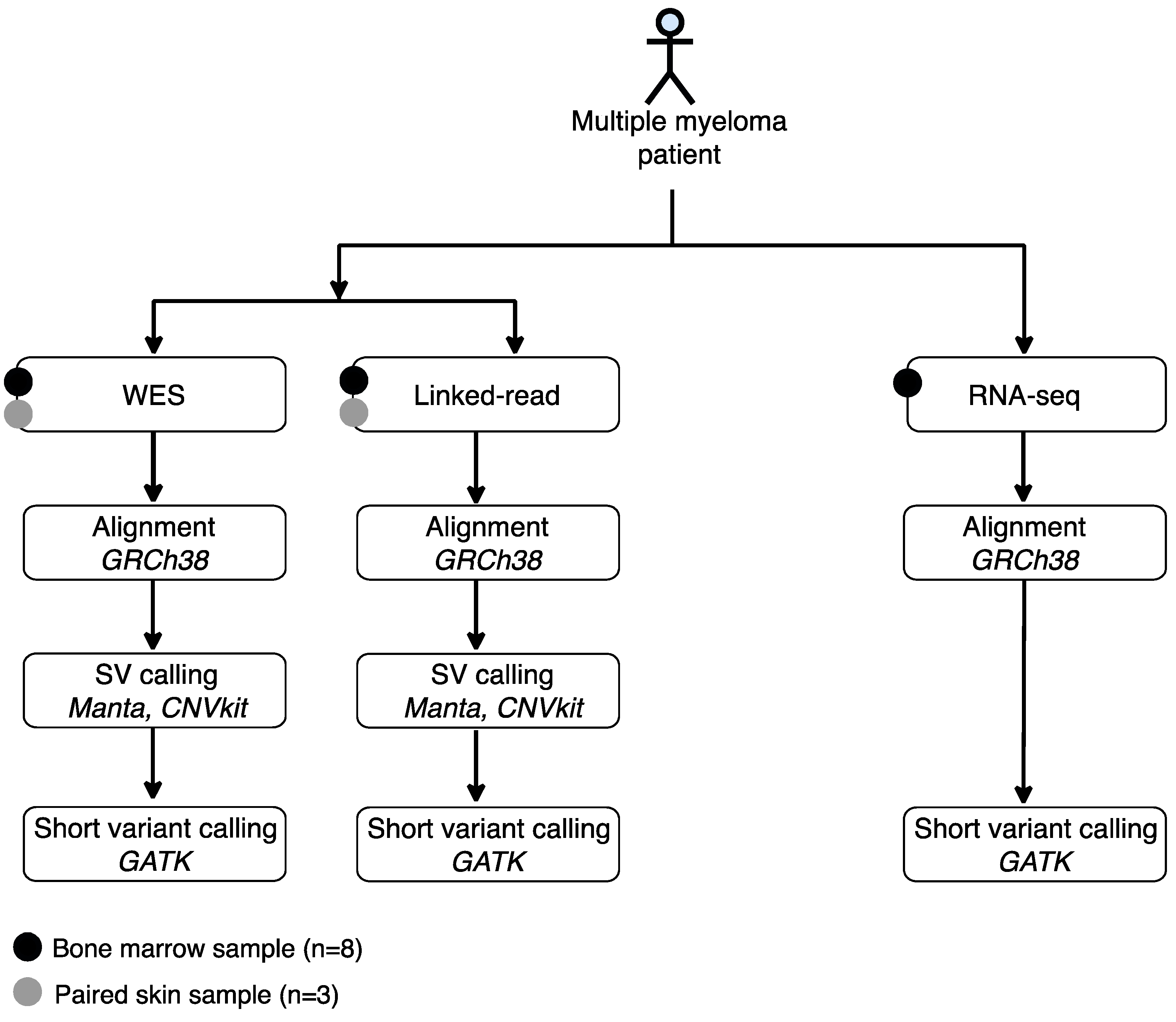
Cancers | Free Full-Text | Comparison of Structural and Short Variants Detected by Linked-Read and Whole-Exome Sequencing in Multiple Myeloma

iCOMIC: a graphical interface-driven bioinformatics pipeline for analyzing cancer omics data | bioRxiv
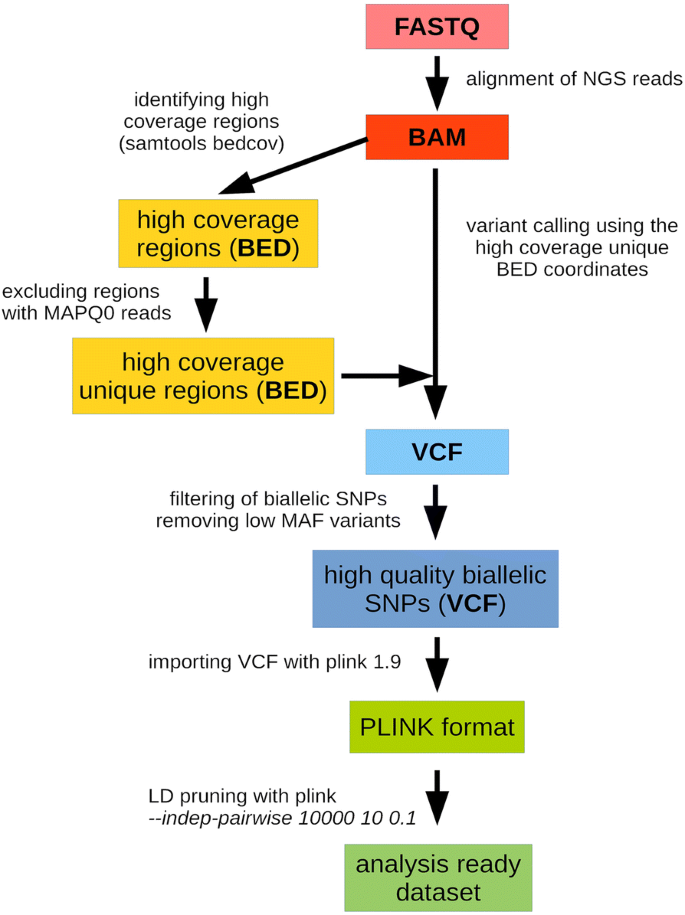
Evaluation of whole exome sequencing as an alternative to BeadChip and whole genome sequencing in human population genetic analysis | BMC Genomics | Full Text

Exome Sequencing | Whole Exome Sequencing Cost | SNP Genotyping | Quality NGS Bioinformatics Data Analysis Services
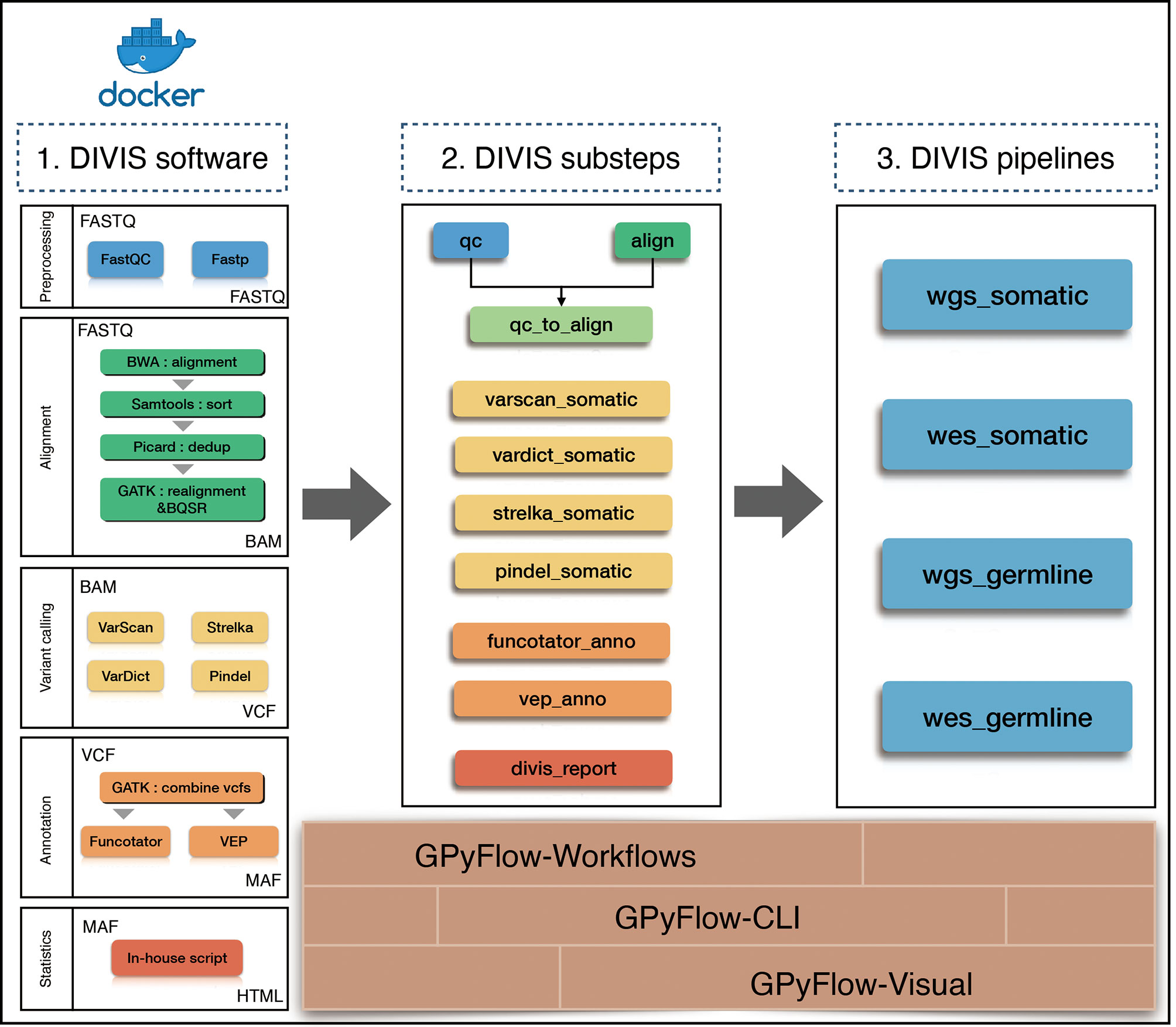
Frontiers | DIVIS: Integrated and Customizable Pipeline for Cancer Genome Sequencing Analysis and Interpretation
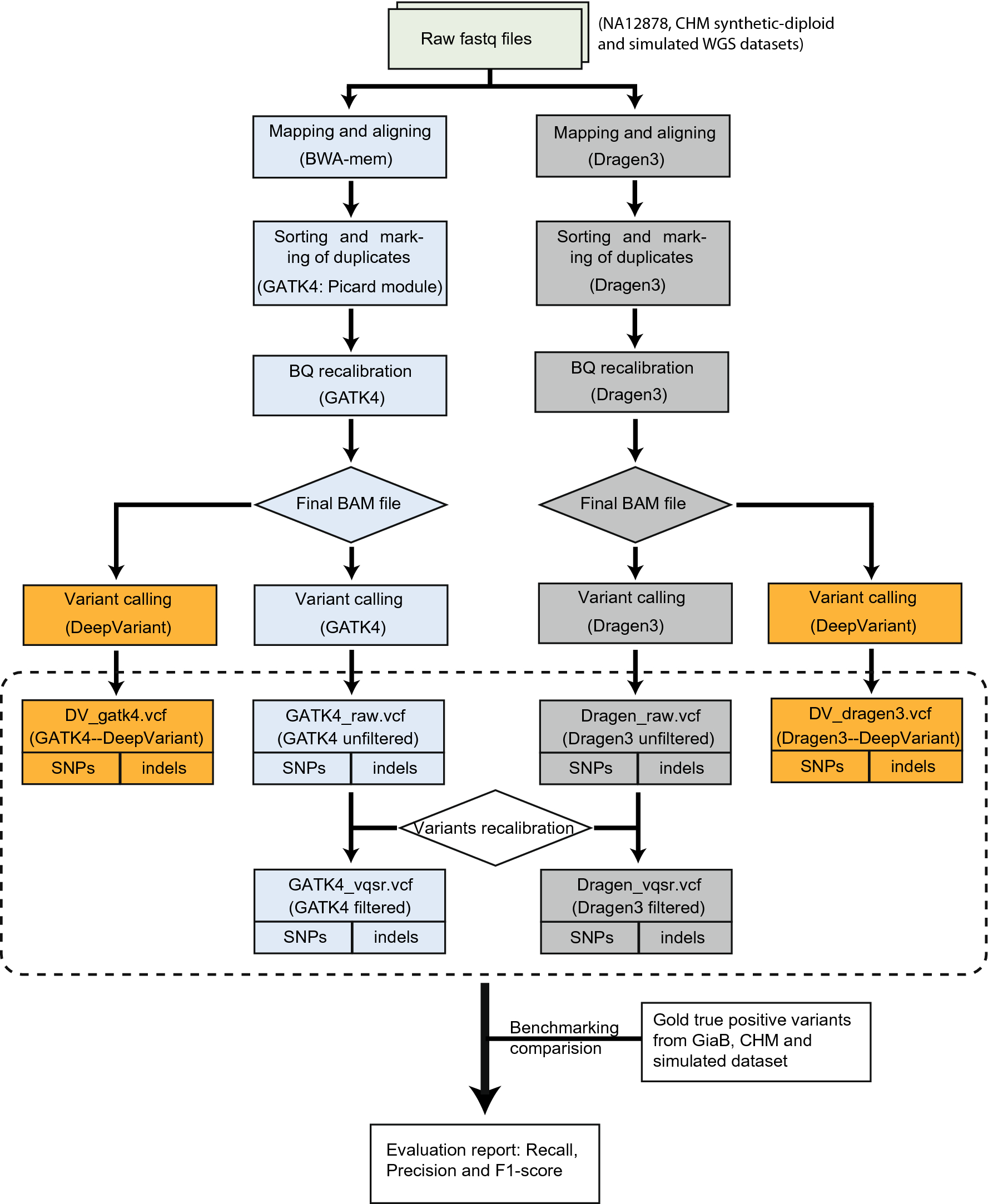
Accuracy and efficiency of germline variant calling pipelines for human genome data | Scientific Reports
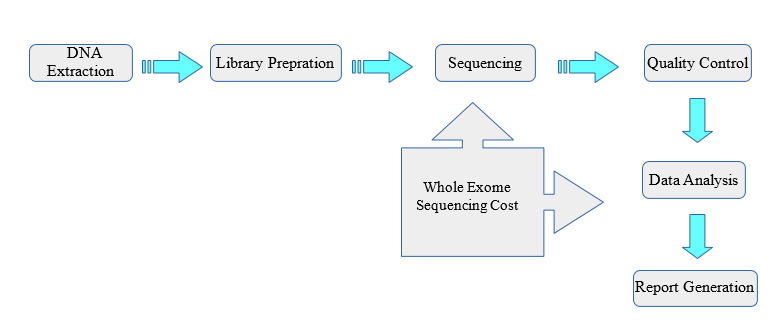
1010Genome Presents Next Generation Sequencing and Whole Exome Sequencing Services | Quality NGS Bioinformatics Data Analysis Services
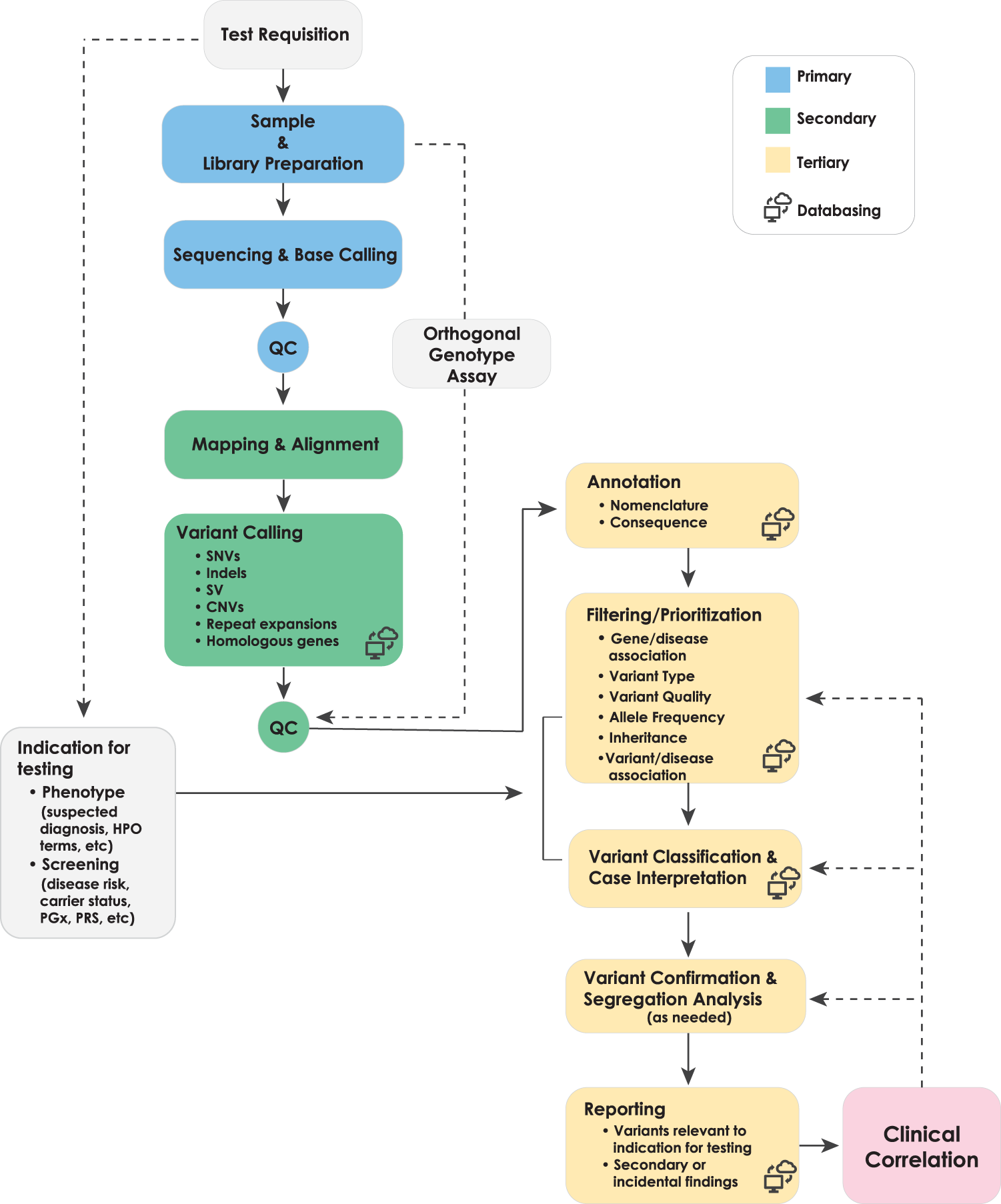
Best practices for the interpretation and reporting of clinical whole genome sequencing | npj Genomic Medicine

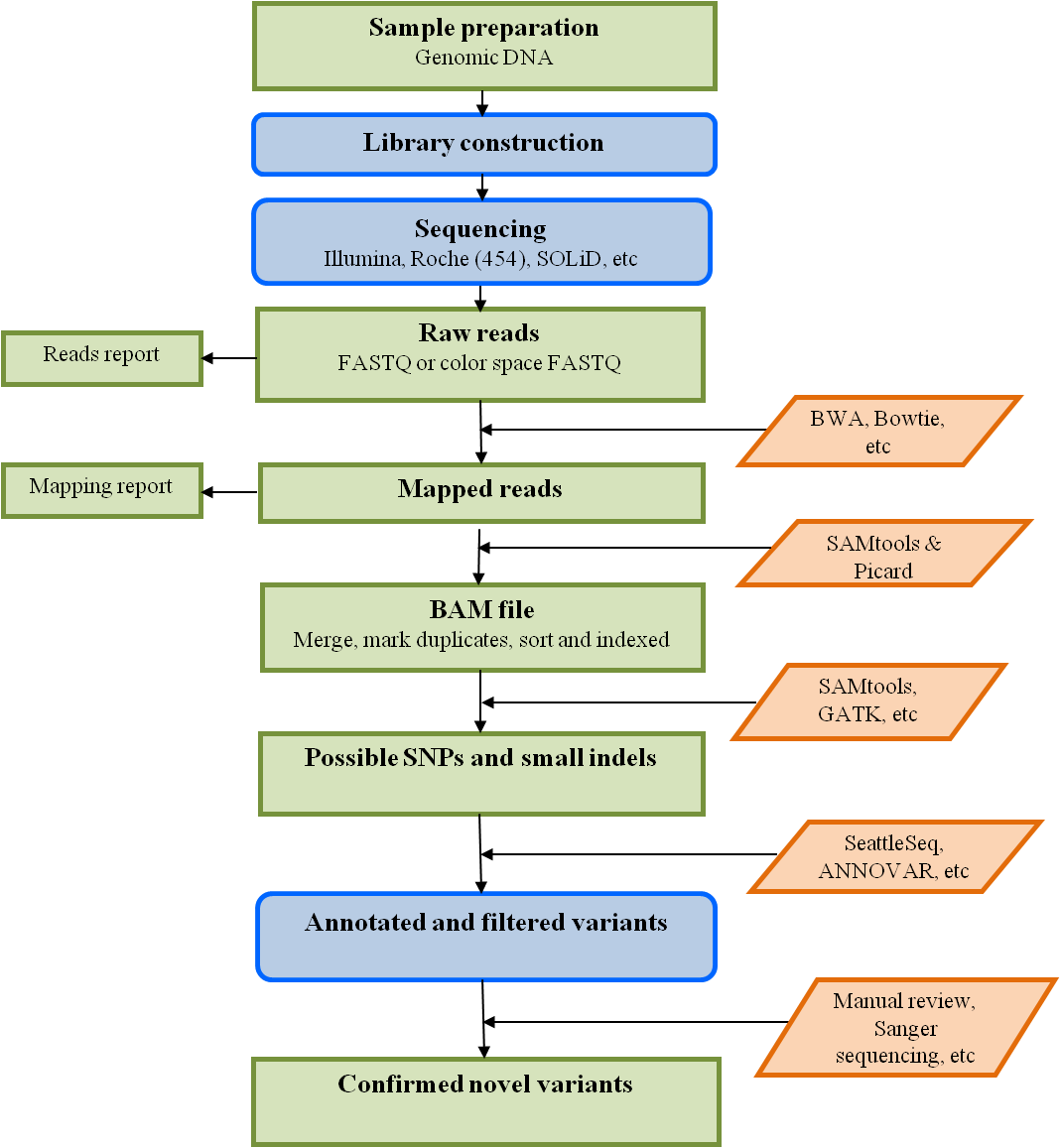
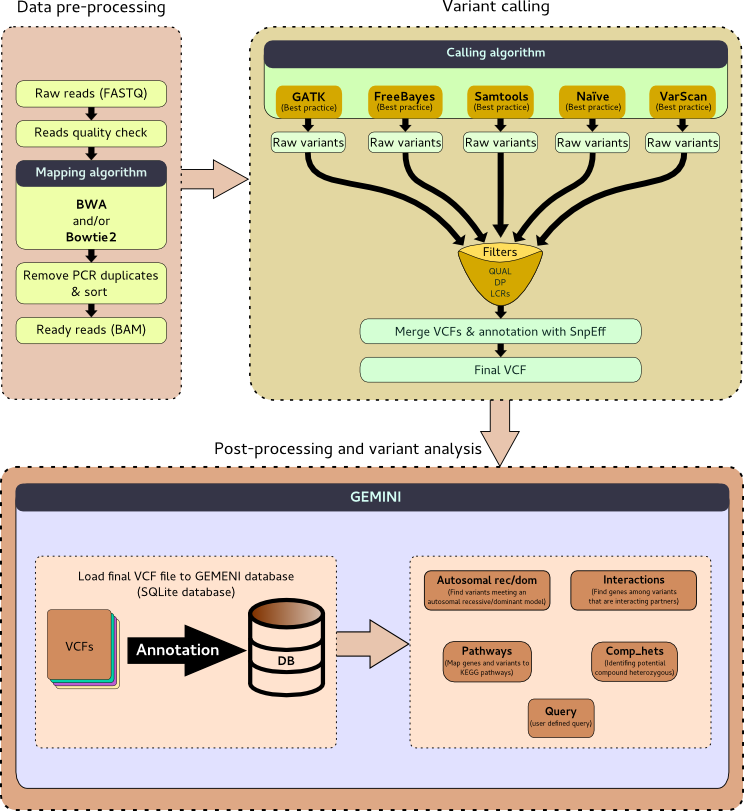
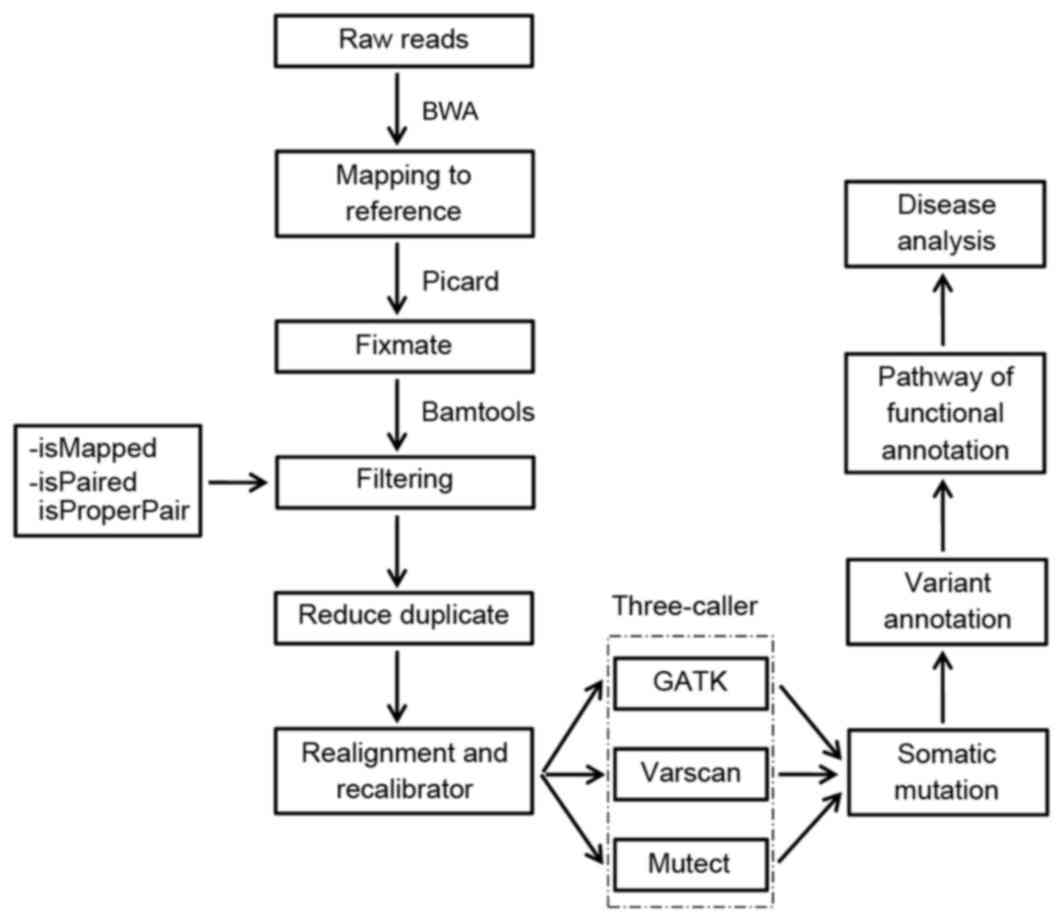
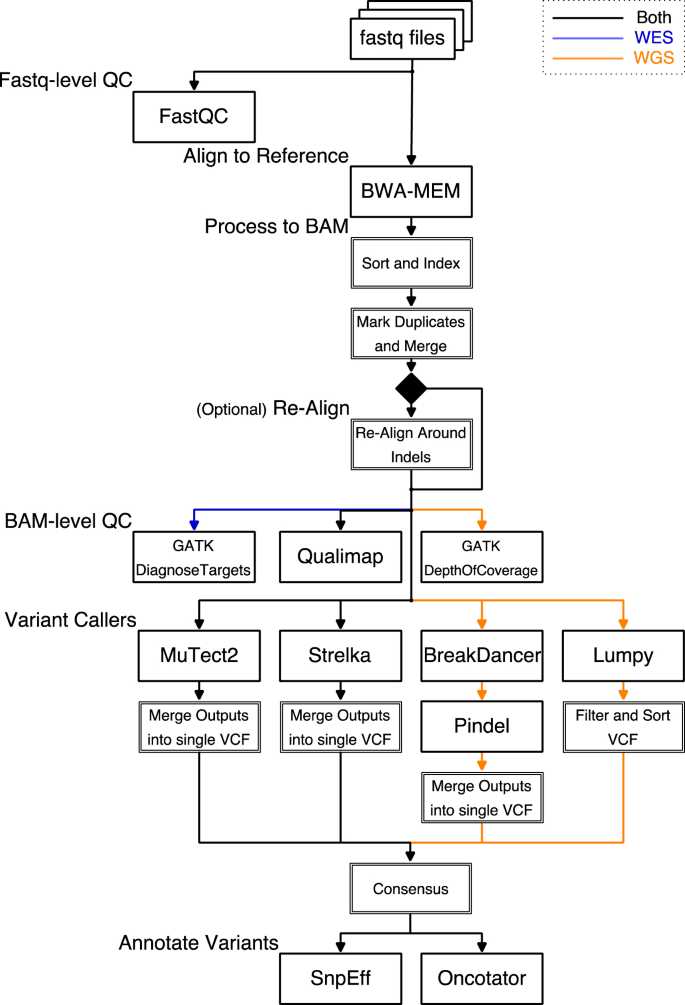
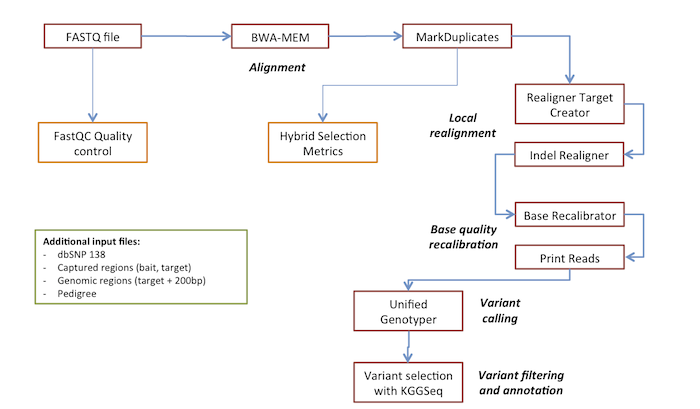
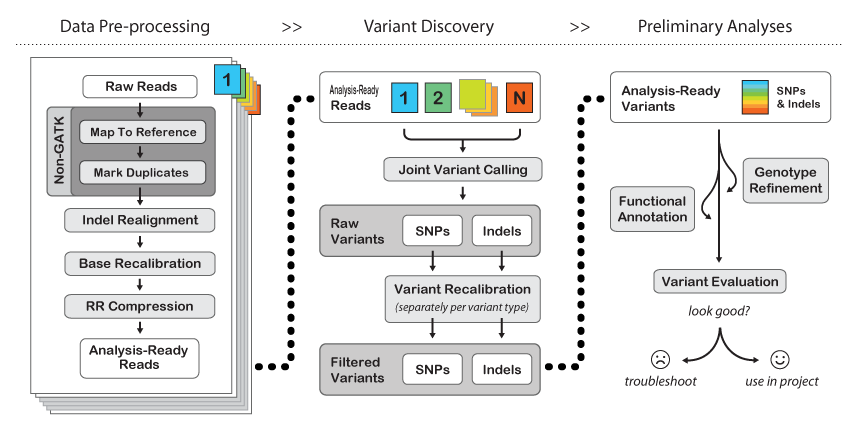
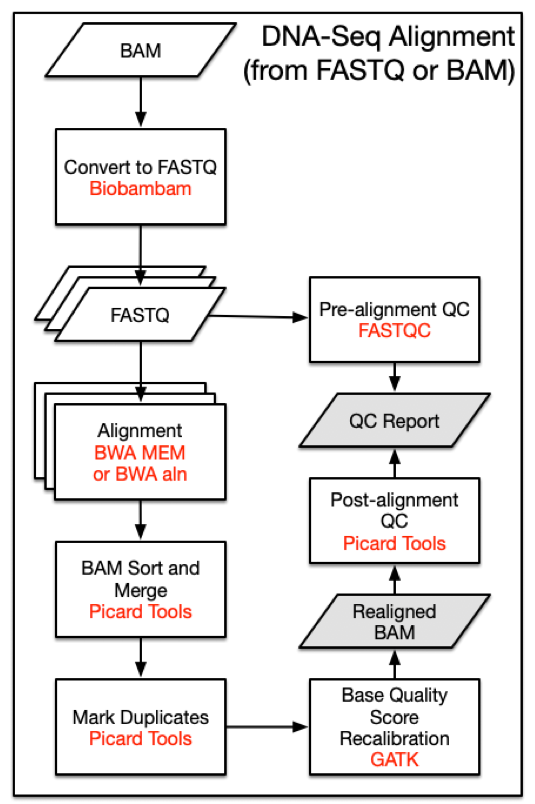
Overview of whole exome sequencing pipeline. SNV, single nucleotide... | Download Scientific Diagram