
Benchmarking machine learning robustness in Covid-19 genome sequence classification | Scientific Reports
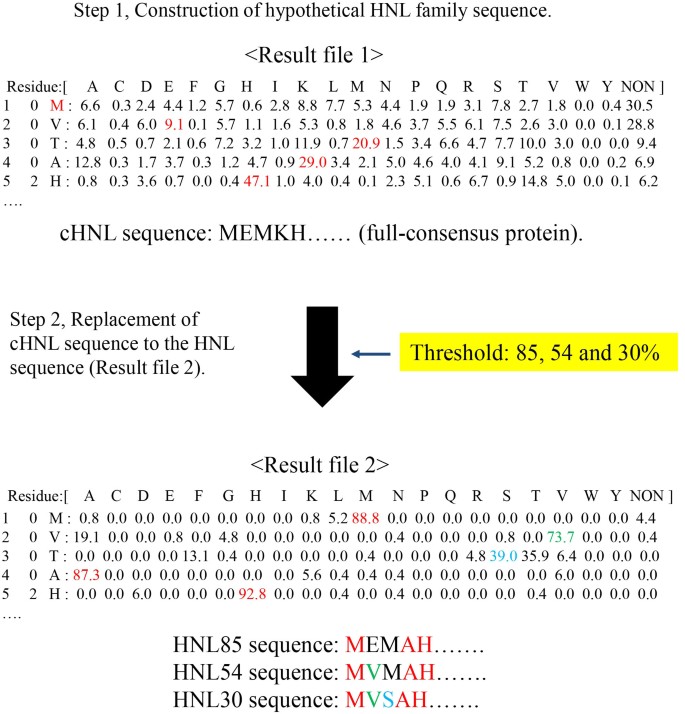
Protein evolution analysis of S-hydroxynitrile lyase by complete sequence design utilizing the INTMSAlign software | Scientific Reports

The use of consensus sequence information to engineer stability and activity in proteins - ScienceDirect
GitHub - msternke/protein-consensus-sequence: A set a basic Python scripts to edit a multiple sequence alignment and generate a protein consensus sequence

The use of consensus sequence information to engineer stability and activity in proteins - ScienceDirect
GitHub - josephhughes/Sequence-manipulation: A range of different perl scripts for manipulating sequences, conducting alignments, consensus sequences, changing formats
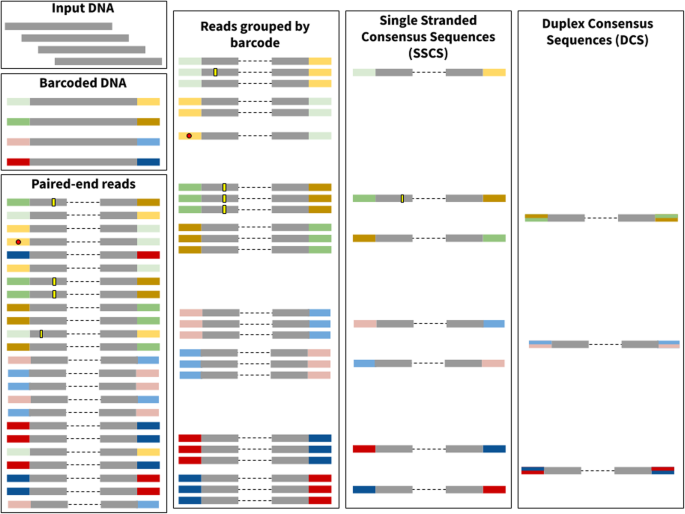

Family reunion via error correction: an efficient analysis of duplex sequencing data | BMC Bioinformatics | Full Text

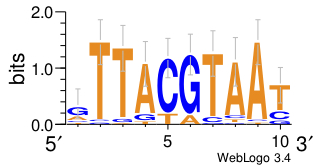
Alignment of σ54 binding sites. Weblogos show the consensus sequence... | Download Scientific Diagram
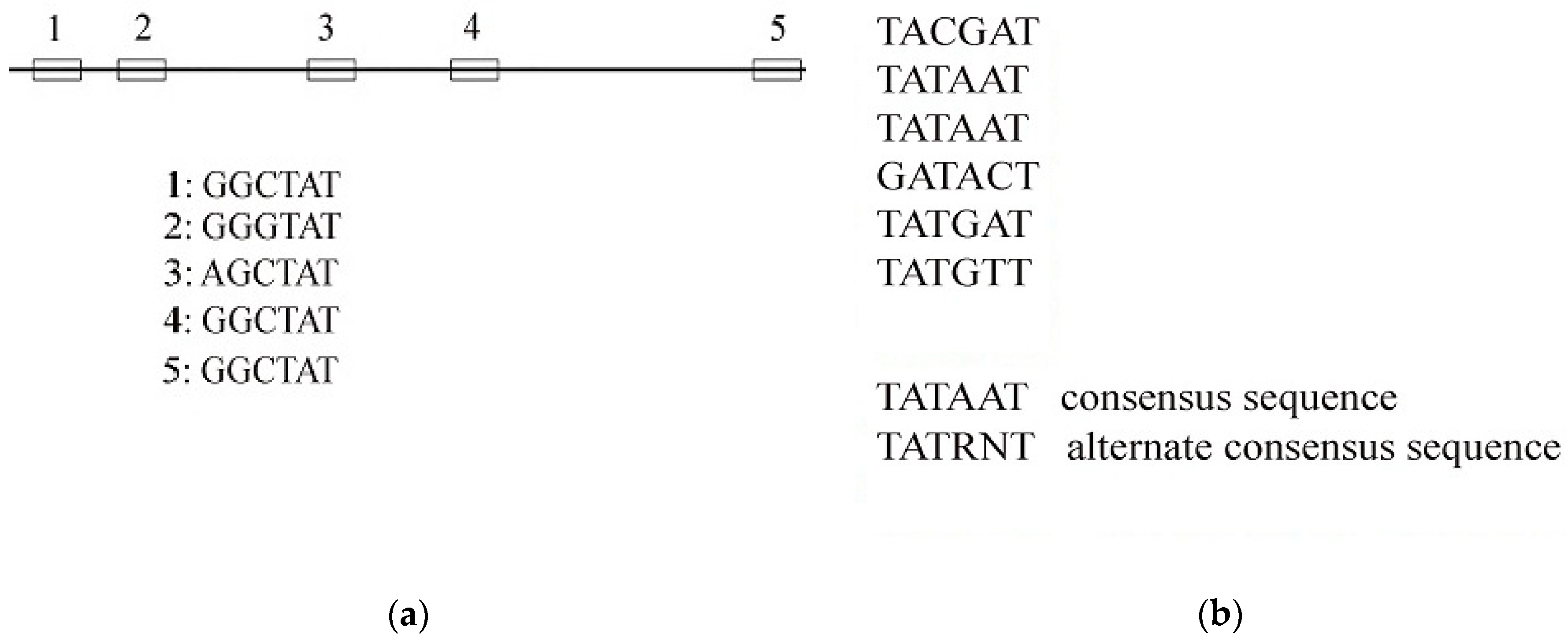
Symmetry | Free Full-Text | A Modified Median String Algorithm for Gene Regulatory Motif Classification
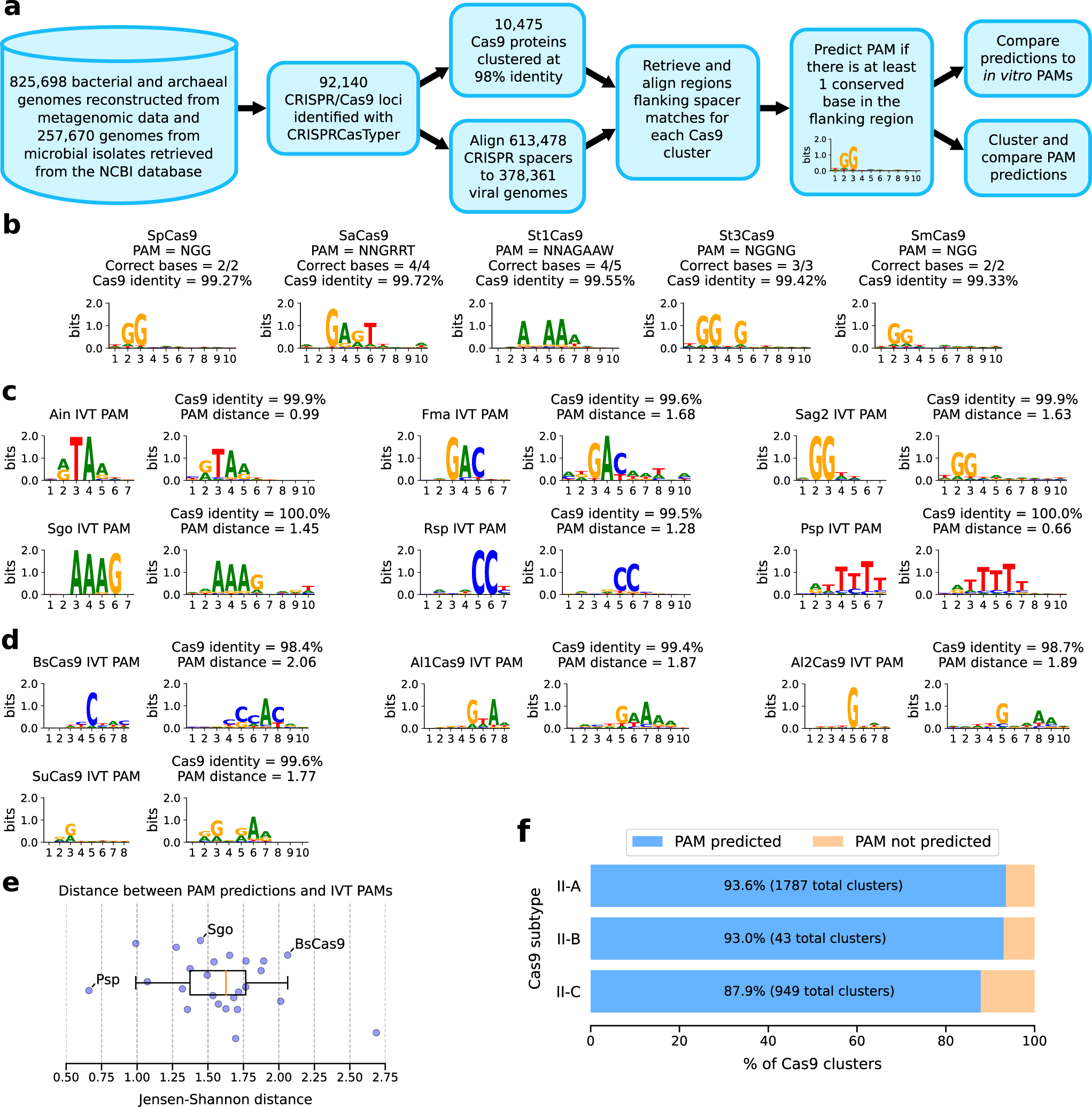
Automated identification of sequence-tailored Cas9 proteins using massive metagenomic data | Nature Communications

Multiple sequence alignment showing global consensus sequence of HCV E2... | Download Scientific Diagram


![Rosalind Consensus and Profile in Bioinformatics with Python [Finding common ancestor] - YouTube Rosalind Consensus and Profile in Bioinformatics with Python [Finding common ancestor] - YouTube](https://i.ytimg.com/vi/lZGCj1ovRwc/maxresdefault.jpg)
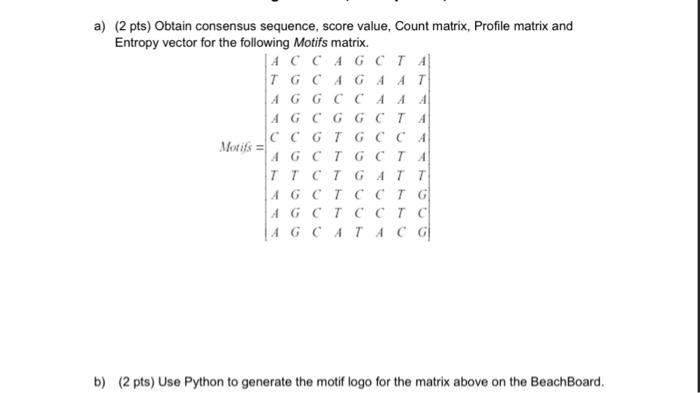






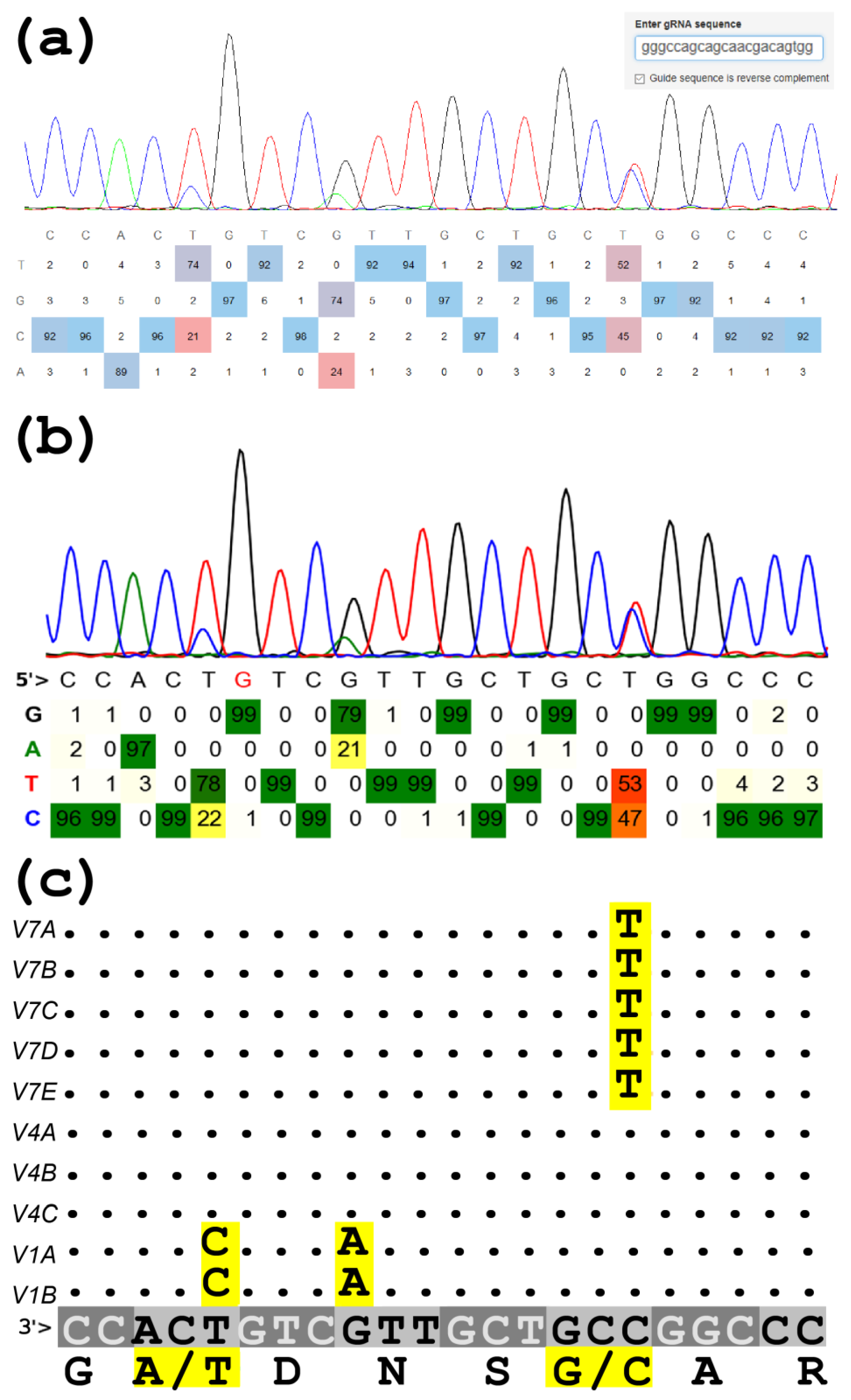




![A Python script to merge Sanger sequences [PeerJ] A Python script to merge Sanger sequences [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11354/1/fig-4-full.png)
