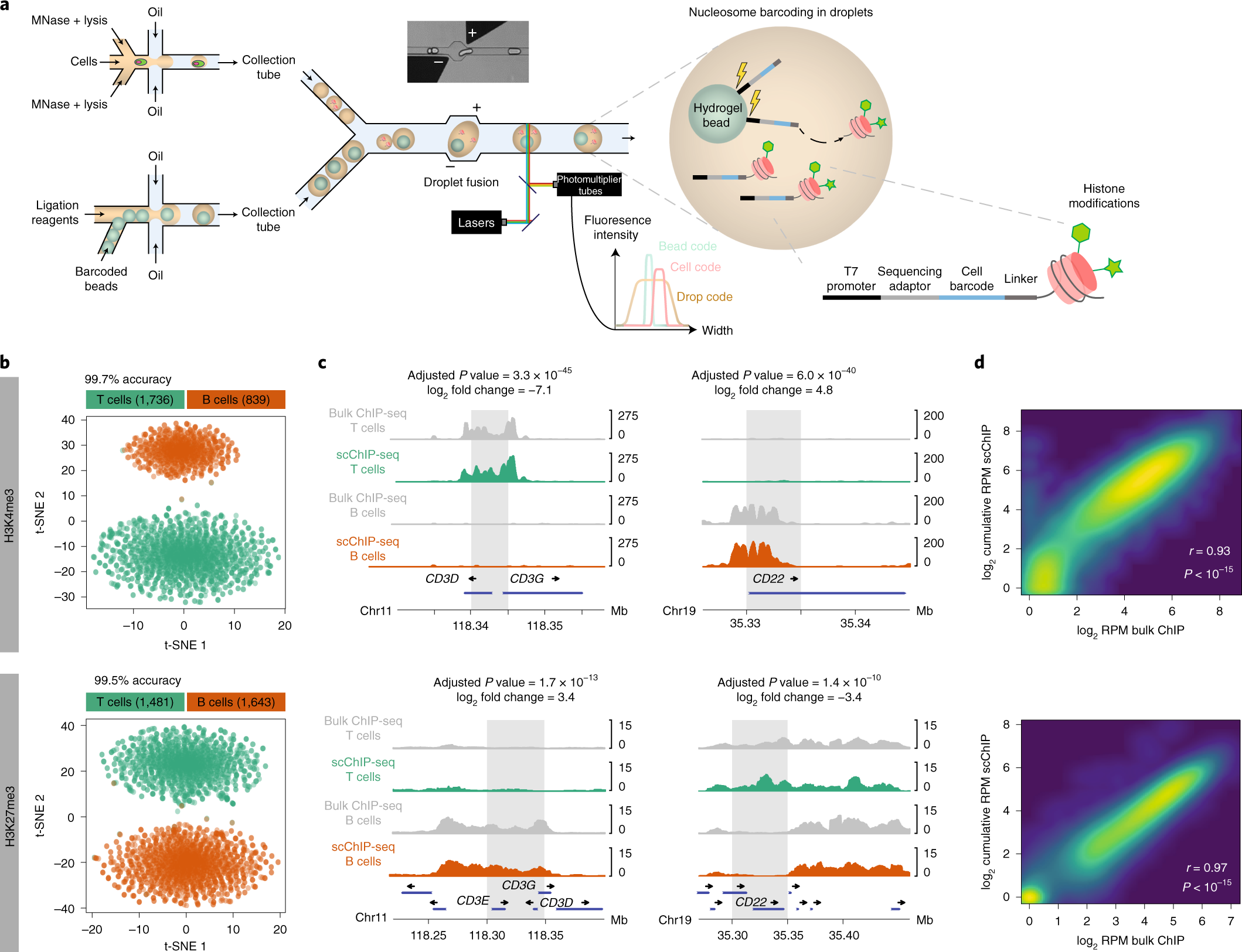
High-throughput single-cell ChIP-seq identifies heterogeneity of chromatin states in breast cancer | Nature Genetics

Chromatin immunoprecipitation-sequencing and RNA-sequencing for complex and low-abundance tree buds | bioRxiv

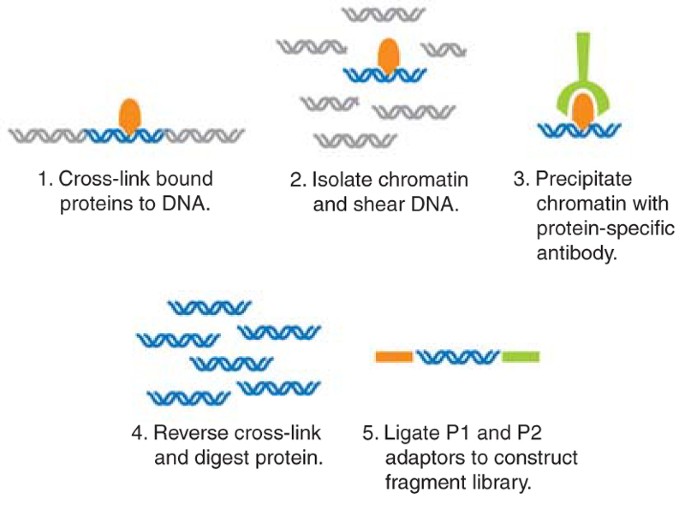
Chromatin Immunoprecipitation Sequencing (ChIP-Seq) on the 5500xl Genetic Analyzer | Thermo Fisher Scientific - DE

TAF-ChIP: an ultra-low input approach for genome-wide chromatin immunoprecipitation assay | Life Science Alliance
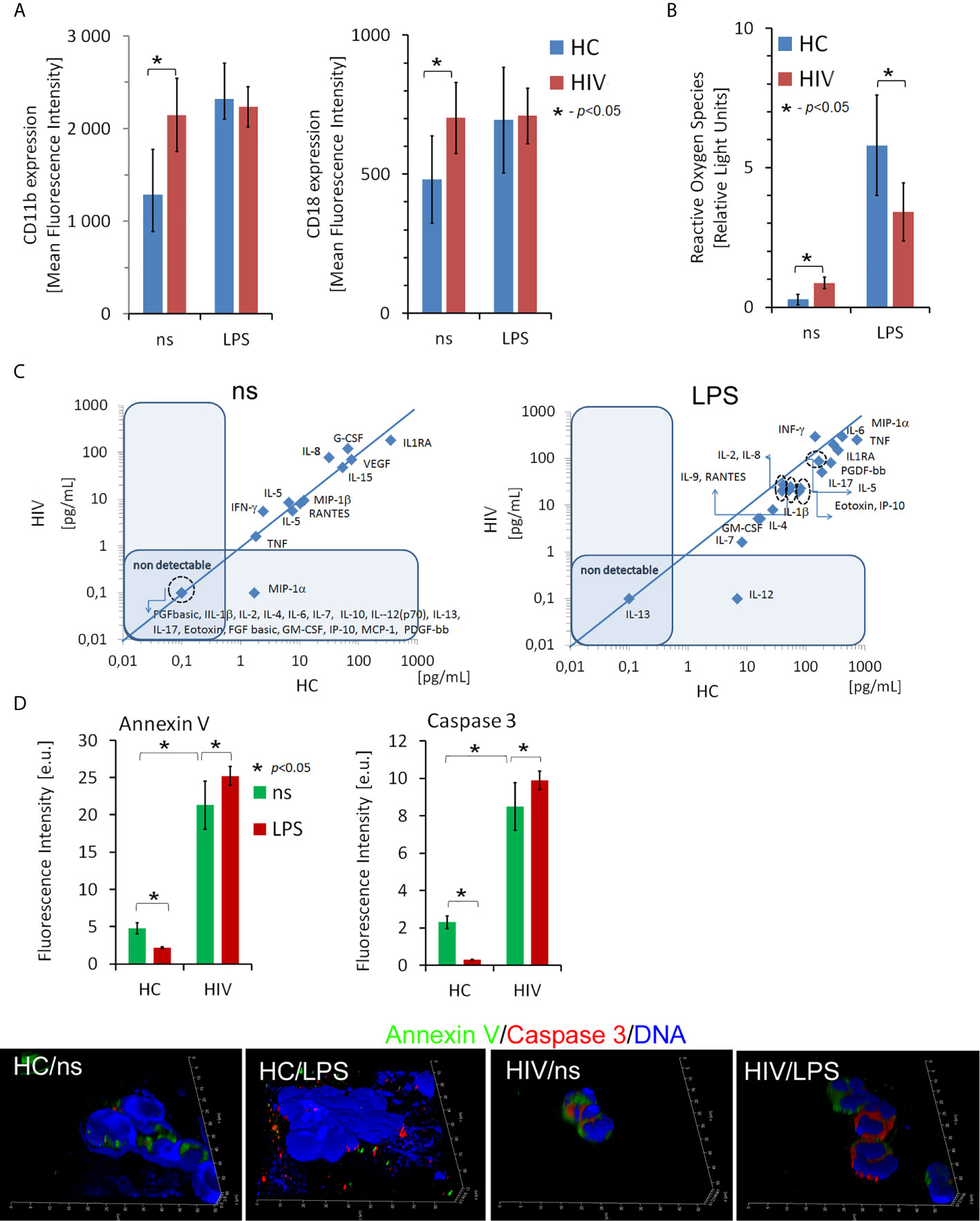
Frontiers | H3K4me3 Histone ChIP-Seq Analysis Reveals Molecular Mechanisms Responsible for Neutrophil Dysfunction in HIV-Infected Individuals