
Proteomic Identification of Protease Cleavage Sites Characterizes Prime and Non-prime Specificity of Cysteine Cathepsins B, L, and S | Journal of Proteome Research

Chemical structure and cleavage mechanism by Cathepsin B of (A) HPMA... | Download Scientific Diagram
Workflow to analyze cathepsin B substrate cleavage site preferences for... | Download Scientific Diagram

Trivalent metal complex geometry of the substrate governs cathepsin B enzymatic cleavage rate - Chemical Communications (RSC Publishing)

TMPRSS2 and furin are both essential for proteolytic activation of SARS-CoV-2 in human airway cells | Life Science Alliance
![PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/694d662801bcbb00cd4c5638a9a6d7ac44b518cf/4-Figure2-1.png)
PDF] Fluorogenic peptide substrates for carboxydipeptidase activity of cathepsin B. | Semantic Scholar

Protease cleavage site fingerprinting by label‐free in‐gel degradomics reveals pH‐dependent specificity switch of legumain | The EMBO Journal

Sequence patterns at cathepsin D and E cleavage sites. In the upper and... | Download Scientific Diagram
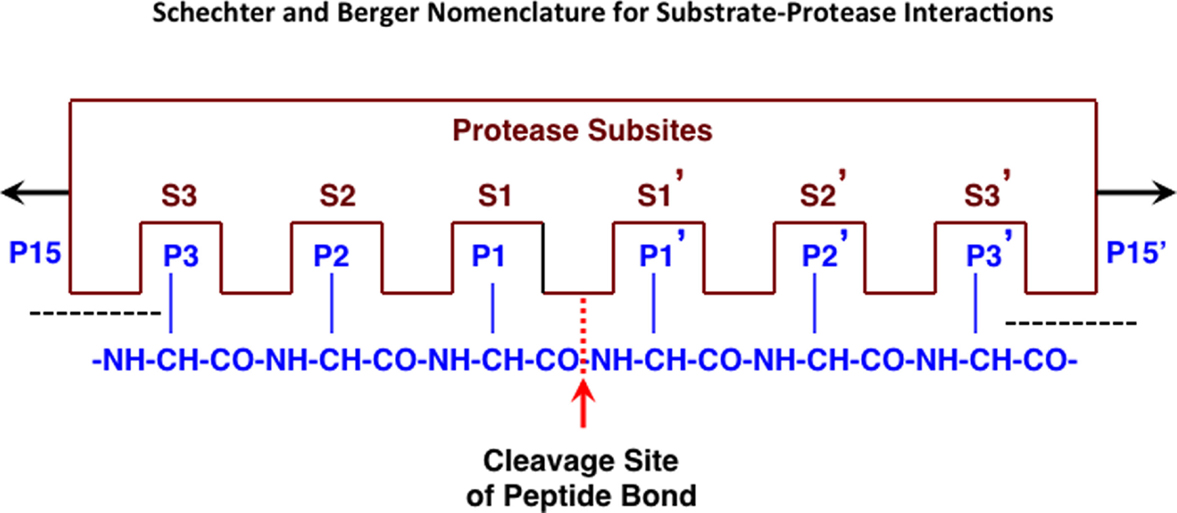
Frontiers | Cathepsin B is a New Drug Target for Traumatic Brain Injury Therapeutics: Evidence for E64d as a Promising Lead Drug Candidate

Selective Neutral pH Inhibitor of Cathepsin B Designed Based on Cleavage Preferences at Cytosolic and Lysosomal pH Conditions | ACS Chemical Biology

Sequence patterns at cathepsin D and E cleavage sites. In the upper and... | Download Scientific Diagram
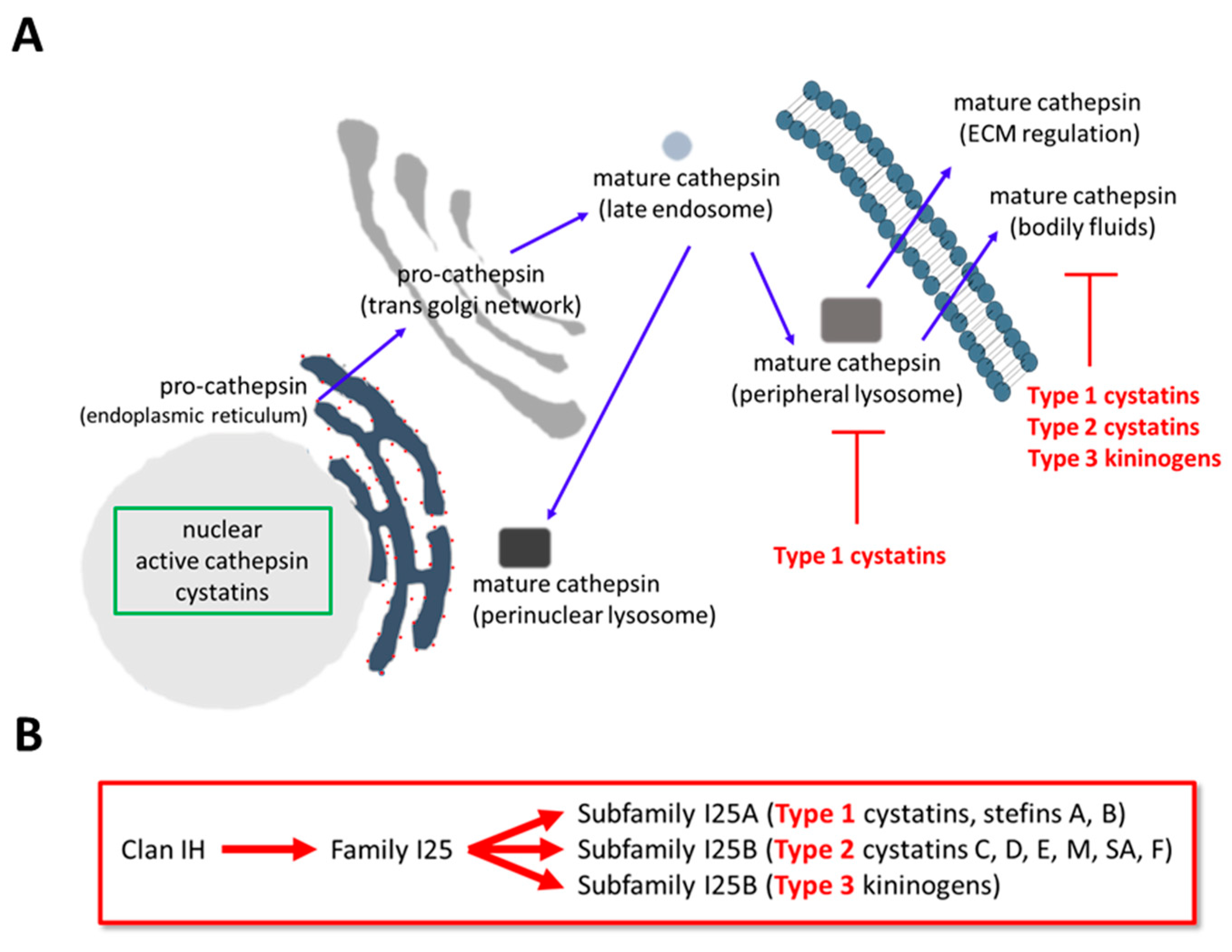
Pharmaceuticals | Free Full-Text | Cysteine Cathepsin Protease Inhibition: An update on its Diagnostic, Prognostic and Therapeutic Potential in Cancer

Proteomic Identification of Protease Cleavage Sites Characterizes Prime and Non-prime Specificity of Cysteine Cathepsins B, L, and S | Journal of Proteome Research

Computational predictions of cysteine cathepsin‐mediated fibrinogen proteolysis - Ferrall‐Fairbanks - 2018 - Protein Science - Wiley Online Library

Multiple Sites on SARS-CoV-2 Spike Protein are Susceptible to Proteolysis by Cathepsins B, K, L, S, and V | bioRxiv

The two cathepsin B-like proteases of Arabidopsis thaliana are closely related enzymes with discrete endopeptidase and carboxydipeptidase activities

Computational predictions of cysteine cathepsin‐mediated fibrinogen proteolysis - Ferrall‐Fairbanks - 2018 - Protein Science - Wiley Online Library


