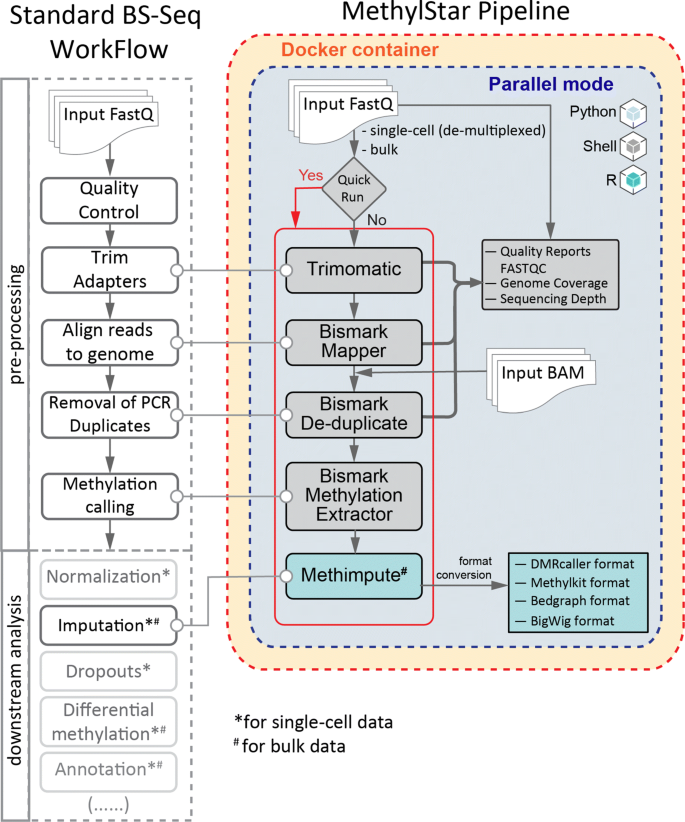
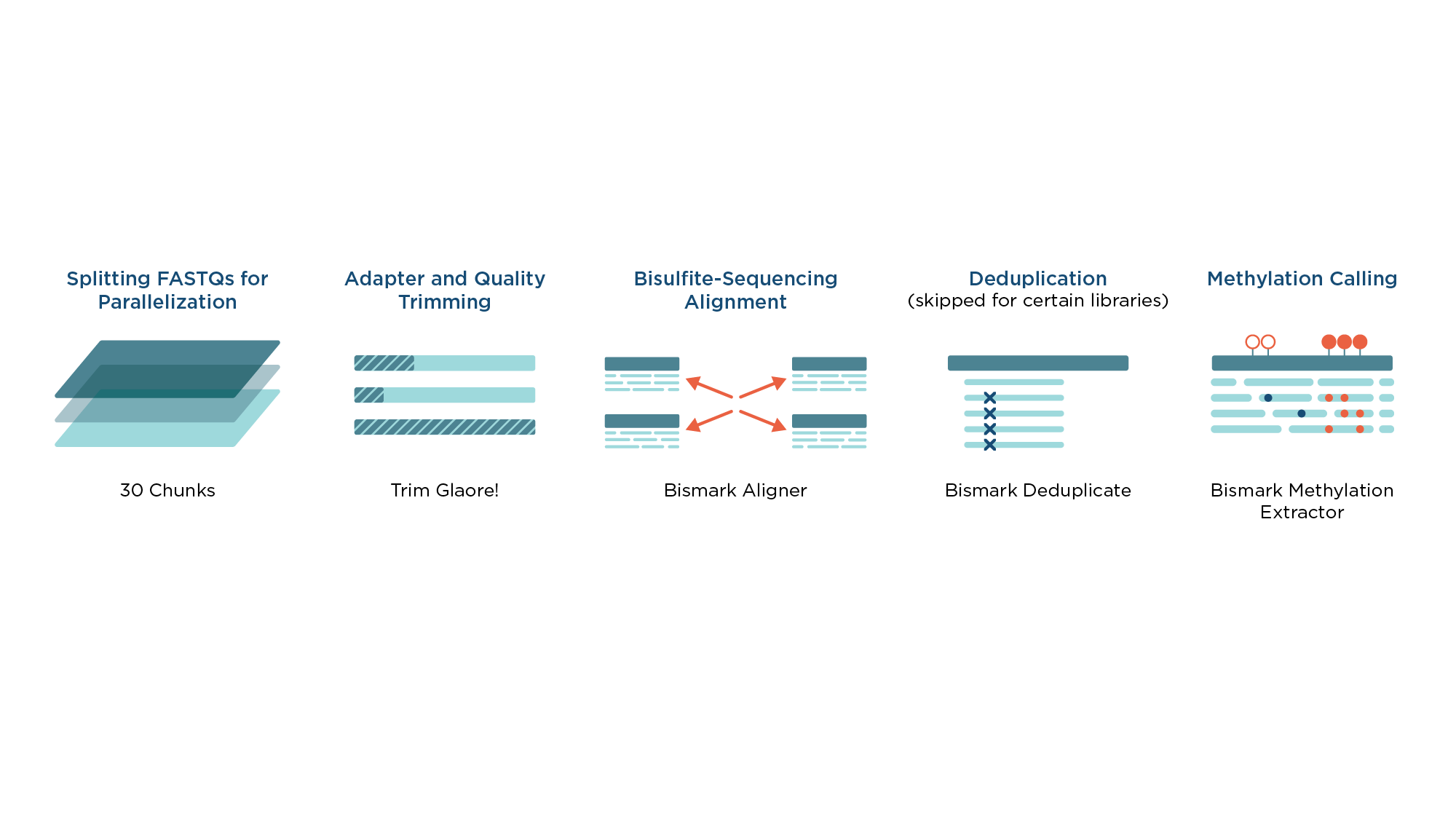
MethylStar: A fast and robust pre-processing pipeline for bulk or single-cell whole-genome bisulfite sequencing data | BMC Genomics | Full Text
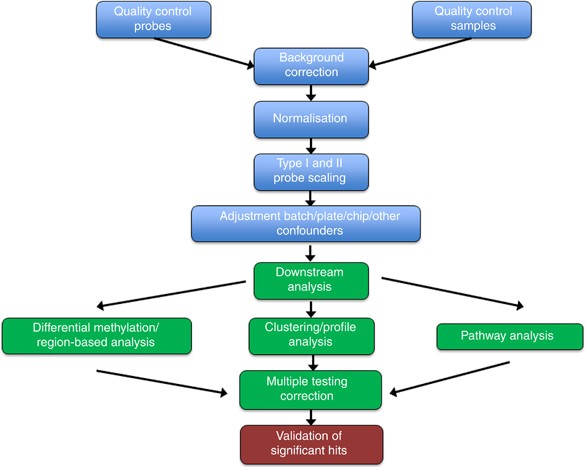
Review of processing and analysis methods for DNA methylation array data | British Journal of Cancer

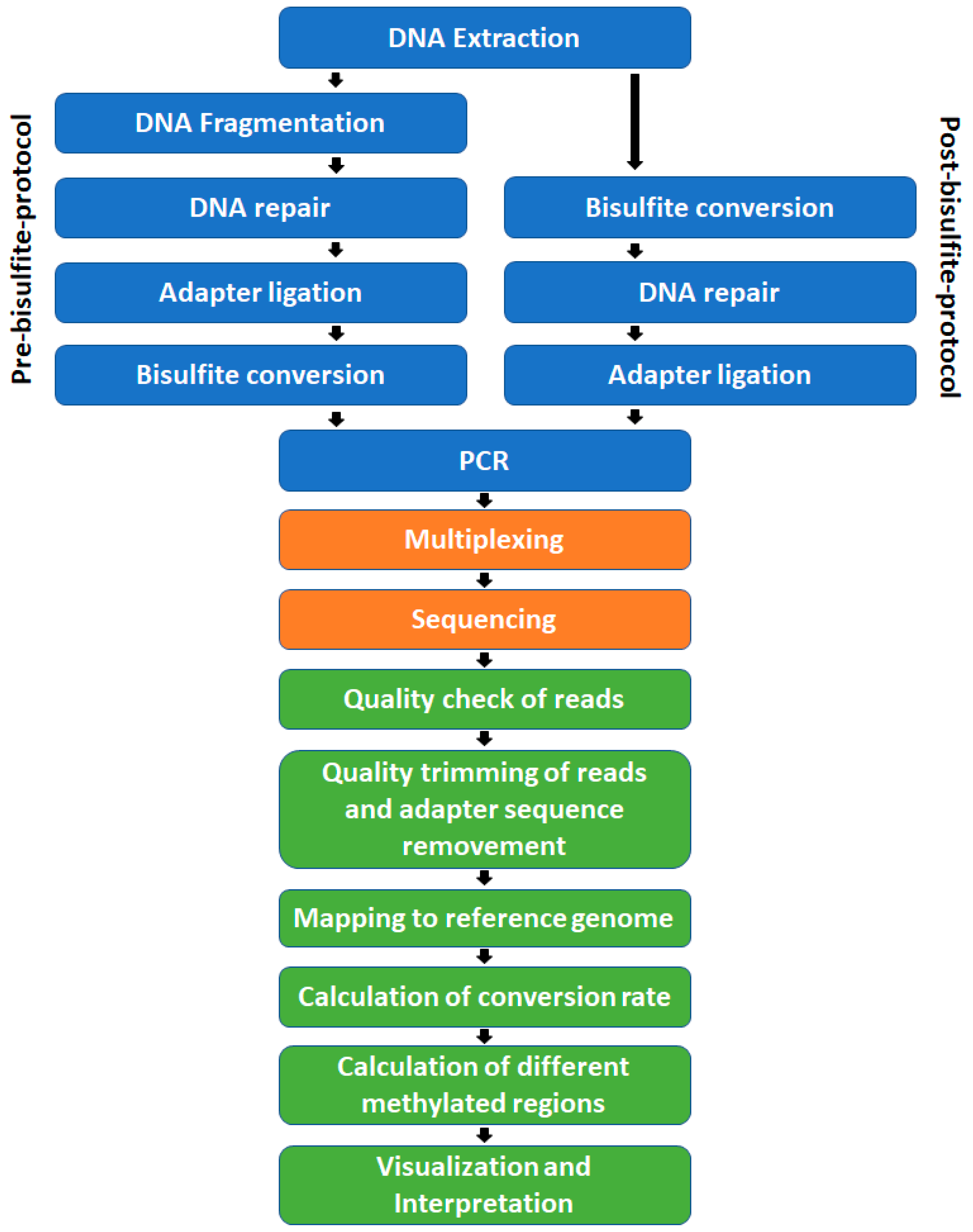
The workflow of analyzing DNA methylation using bisulfite sequencing data. | Download Scientific Diagram
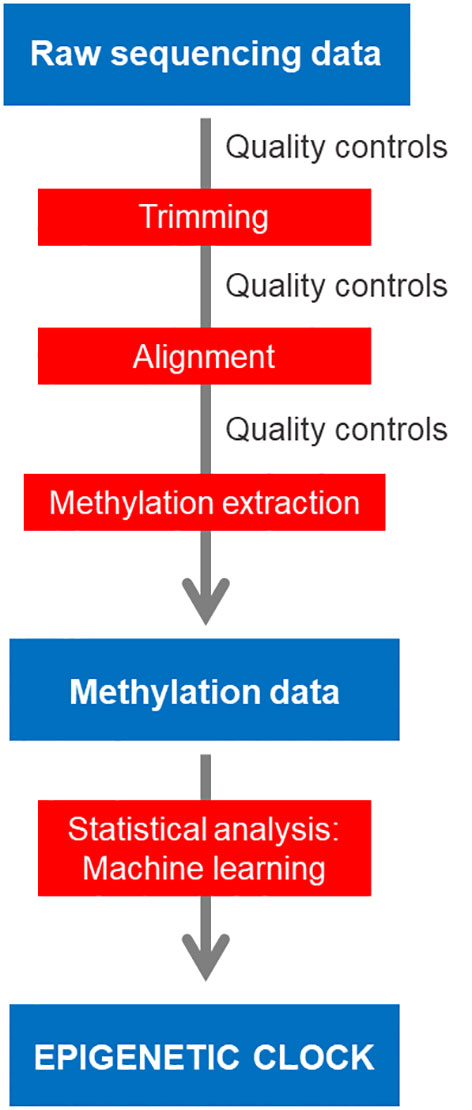
Frontiers | Bioinformatic analysis for age prediction using epigenetic clocks: Application to fisheries management and conservation biology

MethylScore, a pipeline for accurate and context-aware identification of differentially methylated regions from population-scale plant whole-genome bisulfite sequencing data | Quantitative Plant Biology | Cambridge Core
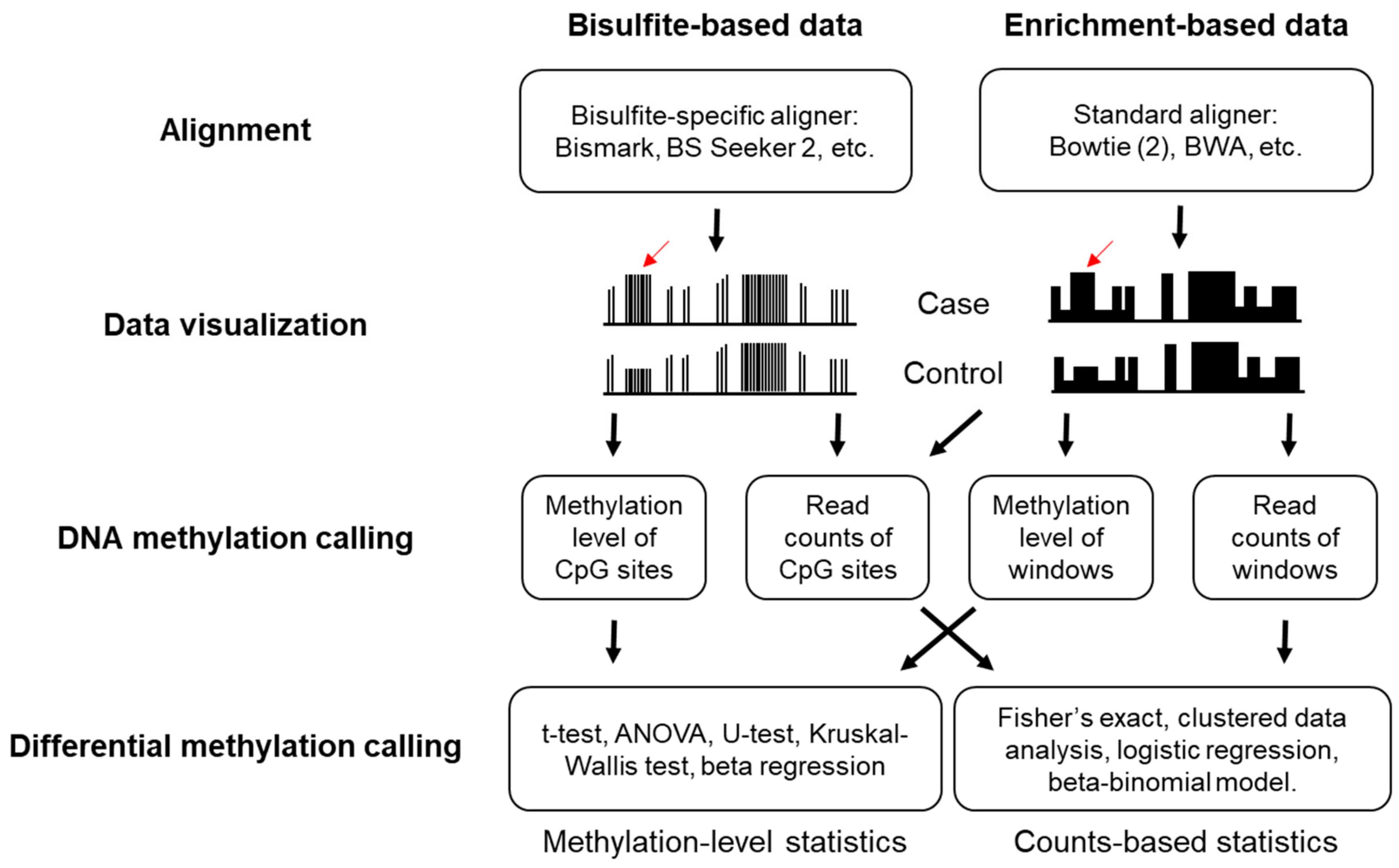
Cancers | Free Full-Text | Cell-Free DNA Methylation Profiling Analysis—Technologies and Bioinformatics

Webinar: Epigenetics Part I – Bisulfite Sequence Analysis and Adenosine to Inosine Modifications - YouTube
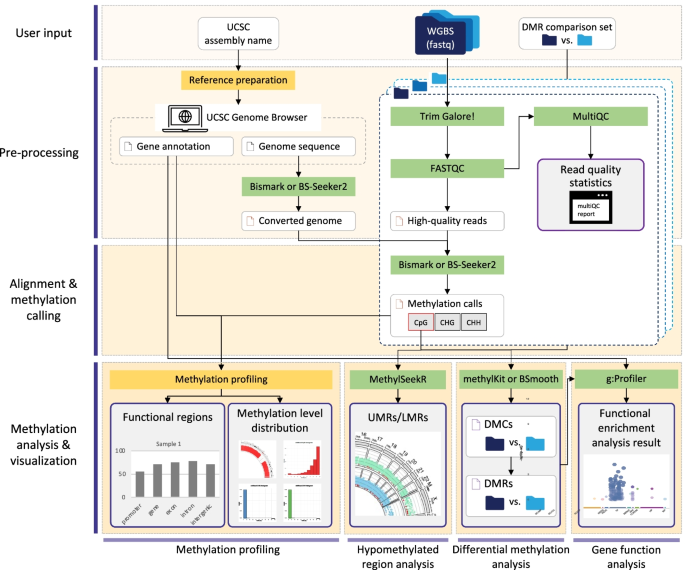
msPIPE: a pipeline for the analysis and visualization of whole-genome bisulfite sequencing data | BMC Bioinformatics | Full Text

Benchmarking DNA methylation analysis of 14 alignment algorithms for whole genome bisulfite sequencing in mammals - ScienceDirect
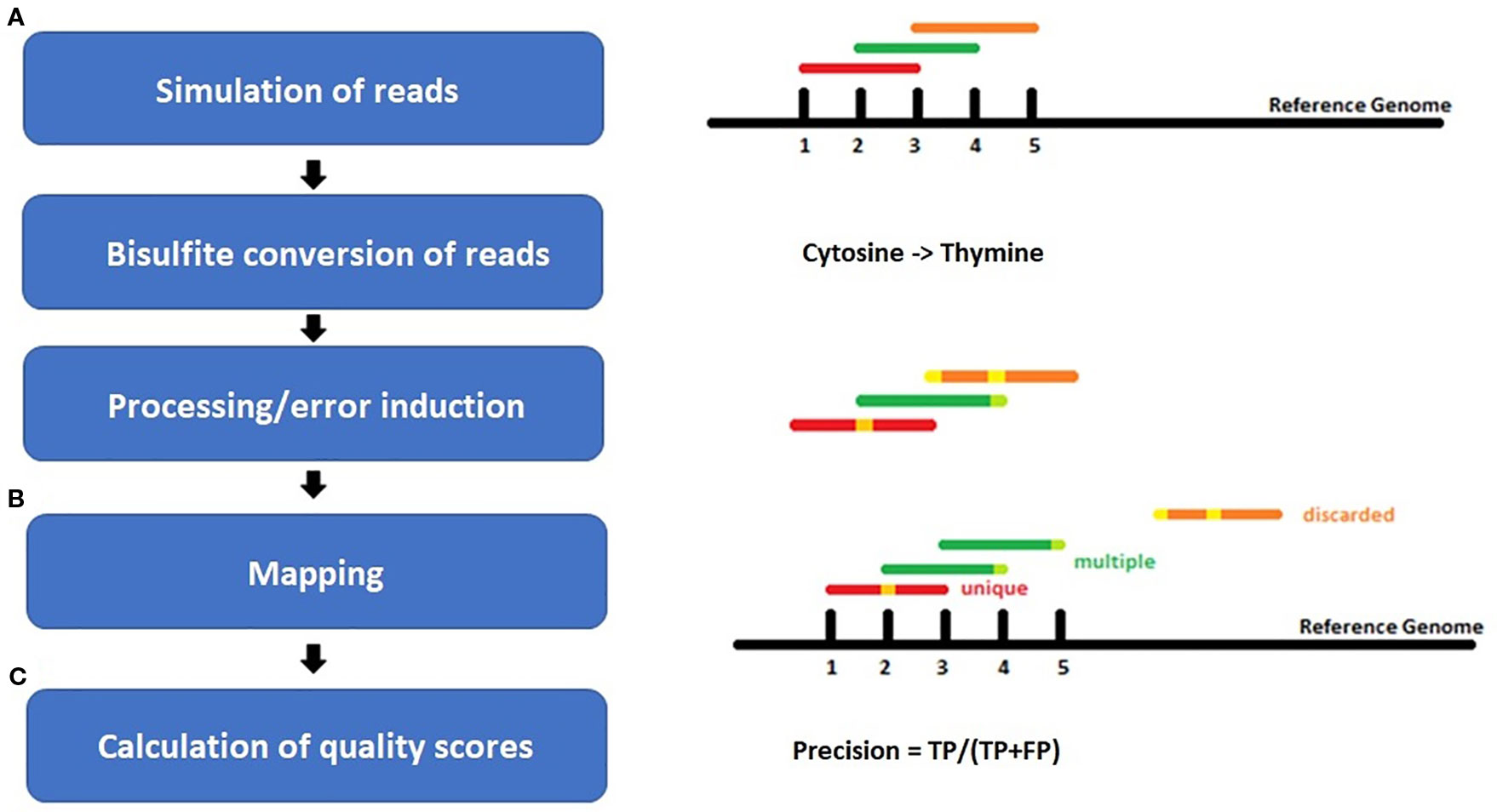
![PDF] MethylSig: a whole genome DNA methylation analysis pipeline | Semantic Scholar PDF] MethylSig: a whole genome DNA methylation analysis pipeline | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/ced5bb274877737921b0e71c311e9043d480021c/3-Figure1-1.png)



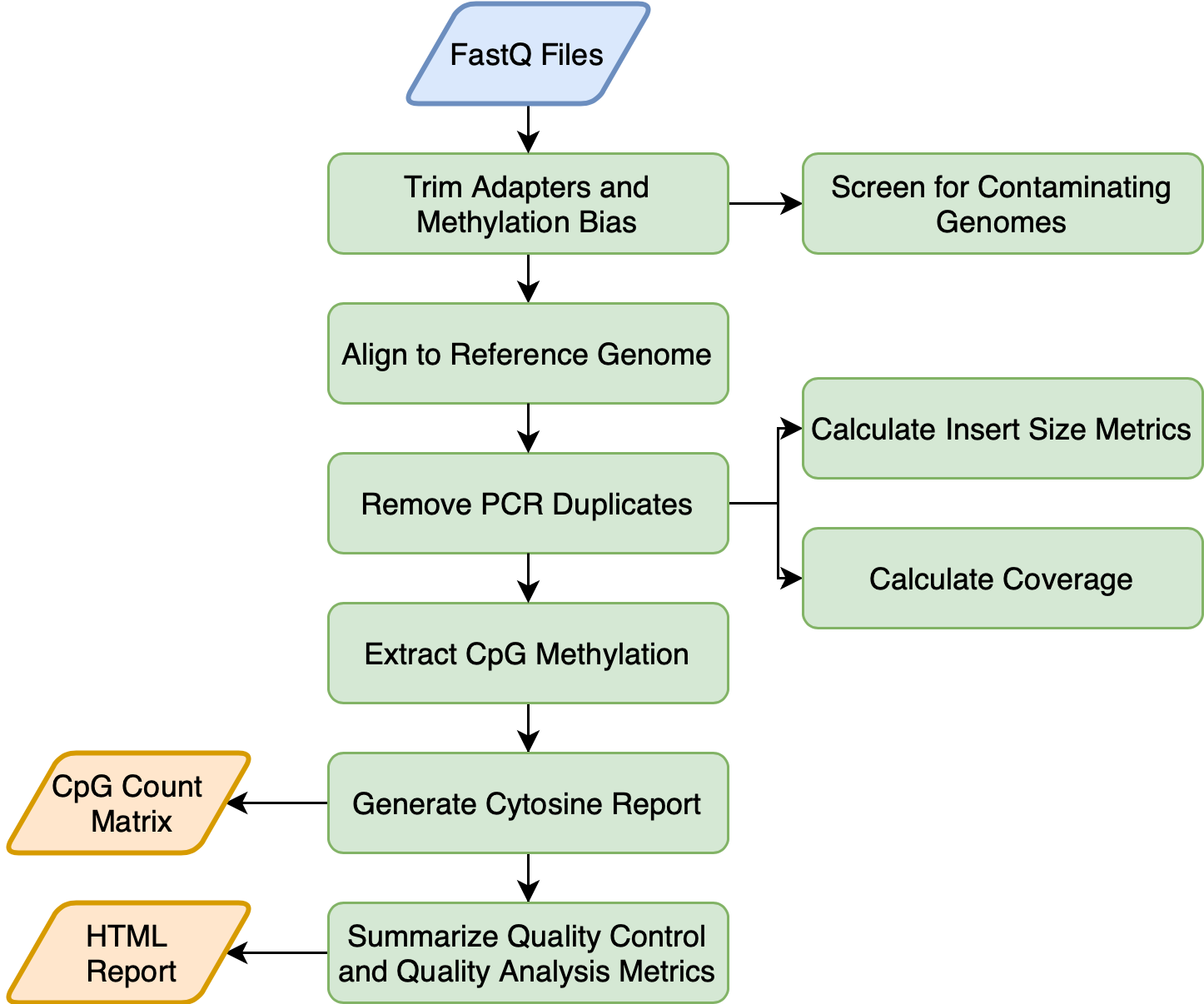



![PDF] Bicycle: a bioinformatics pipeline to analyze bisulfite sequencing data | Semantic Scholar PDF] Bicycle: a bioinformatics pipeline to analyze bisulfite sequencing data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/3fce655ad9994e0081aaf428ba74adf45682a11a/2-Figure1-1.png)
