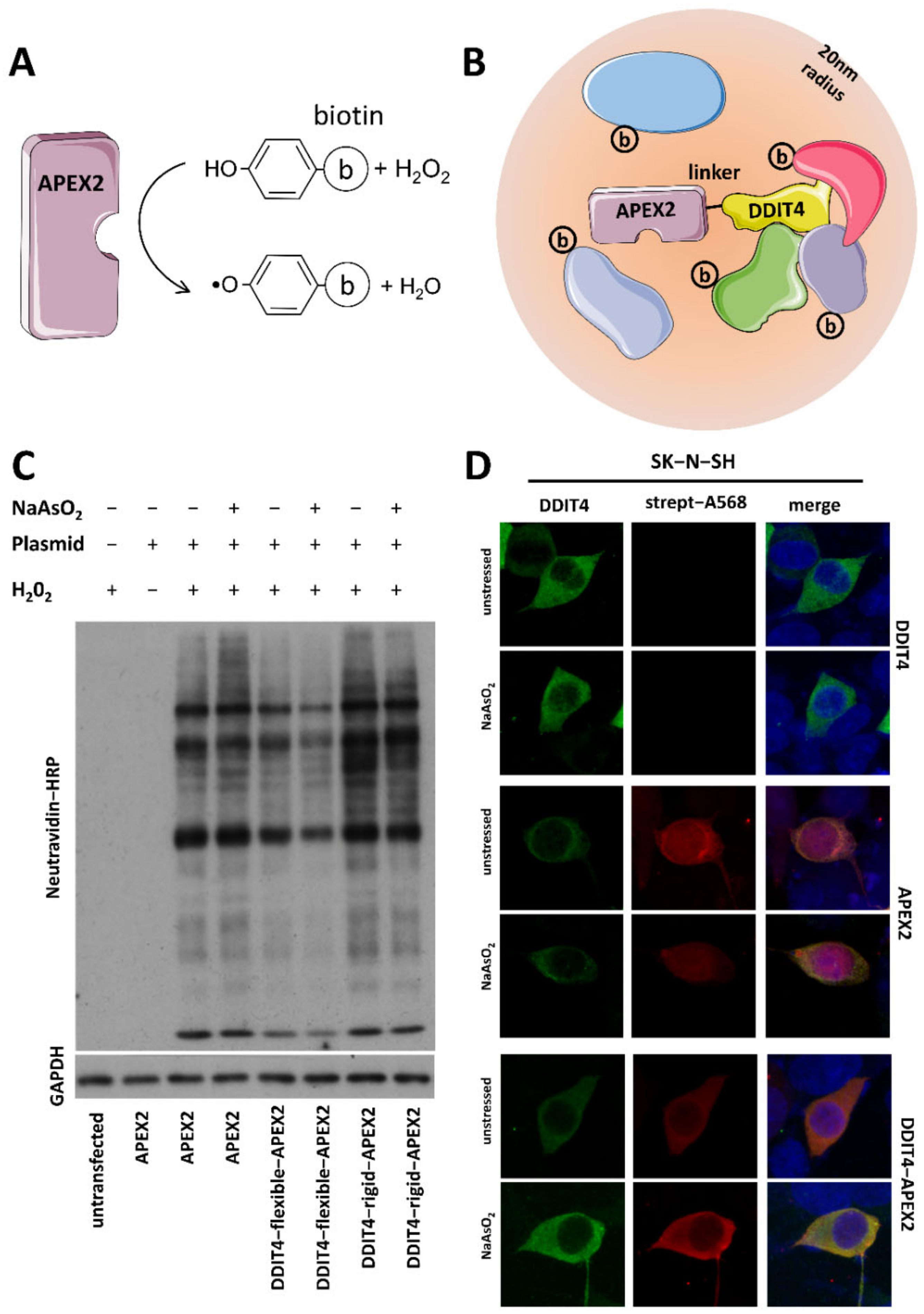
APEX2‐mediated RAB proximity labeling identifies a role for RAB21 in clathrin‐independent cargo sorting | EMBO reports

Targeting APEX2 to human telomerase RNA with CRISPR-Cas13. (A) Design... | Download Scientific Diagram

Expanding APEX2 Substrates for Proximity‐Dependent Labeling of Nucleic Acids and Proteins in Living Cells - Zhou - 2019 - Angewandte Chemie - Wiley Online Library

In vivo discovery of RNA proximal proteins in human cells via proximity-dependent biotinylation | bioRxiv

APEX2‐based Proximity Labeling of Atox1 Identifies CRIP2 as a Nuclear Copper‐binding Protein that Regulates Autophagy Activation - Chen - 2021 - Angewandte Chemie International Edition - Wiley Online Library
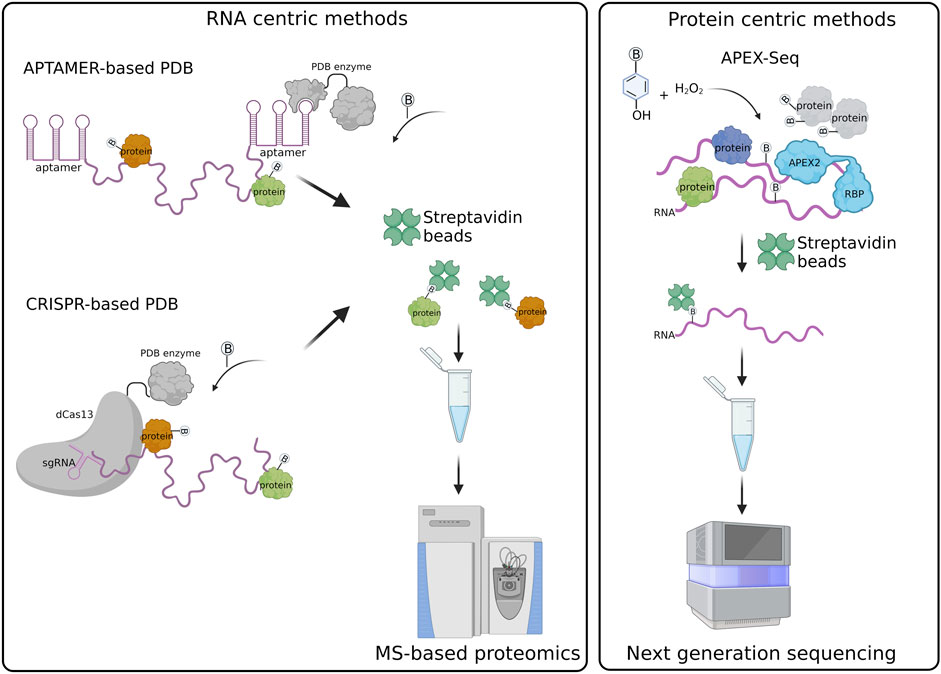
Frontiers | Proximity-dependent biotinylation technologies for mapping RNA-protein interactions in live cells
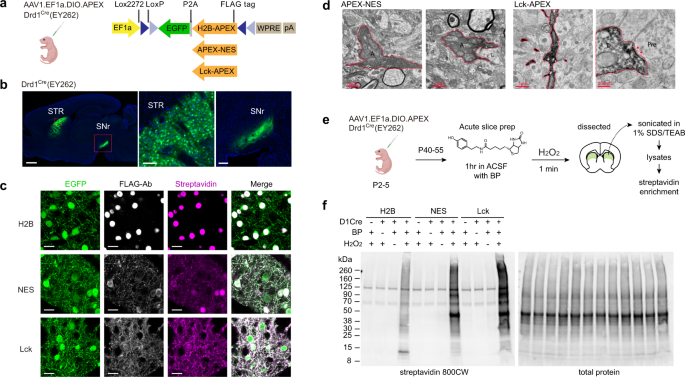
Cell-type and subcellular compartment-specific APEX2 proximity labeling reveals activity-dependent nuclear proteome dynamics in the striatum | Nature Communications

Three-Dimensional Visualization of APEX2-Tagged Erg11 in Saccharomyces cerevisiae Using Focused Ion Beam Scanning Electron Microscopy | mSphere

Proximity RNA Labeling by APEX-Seq Reveals the Organization of Translation Initiation Complexes and Repressive RNA Granules - ScienceDirect
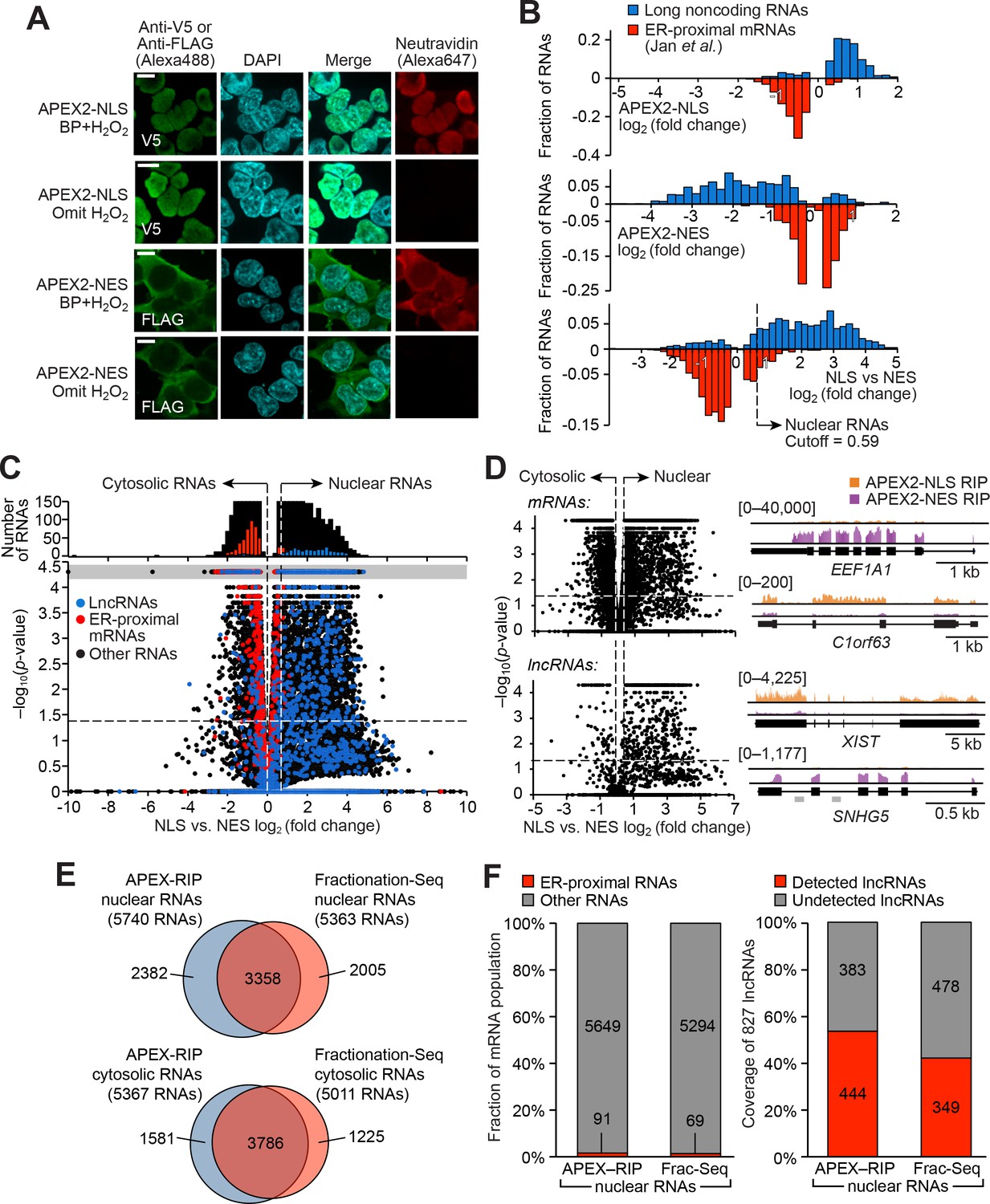
Figures and data in Live-cell mapping of organelle-associated RNAs via proximity biotinylation combined with protein-RNA crosslinking | eLife
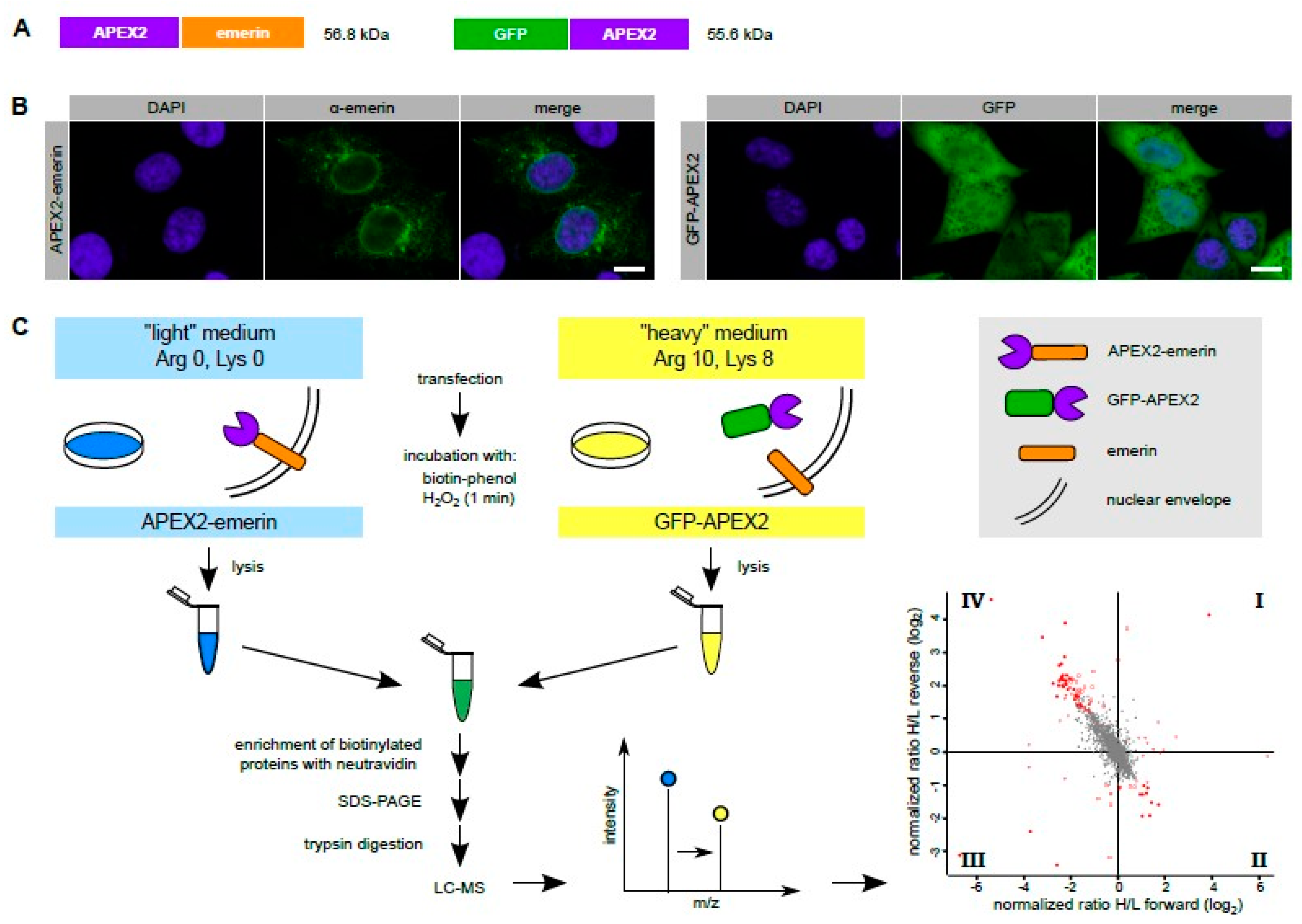
Cells | Free Full-Text | Probing the Environment of Emerin by Enhanced Ascorbate Peroxidase 2 (APEX2)-Mediated Proximity Labeling
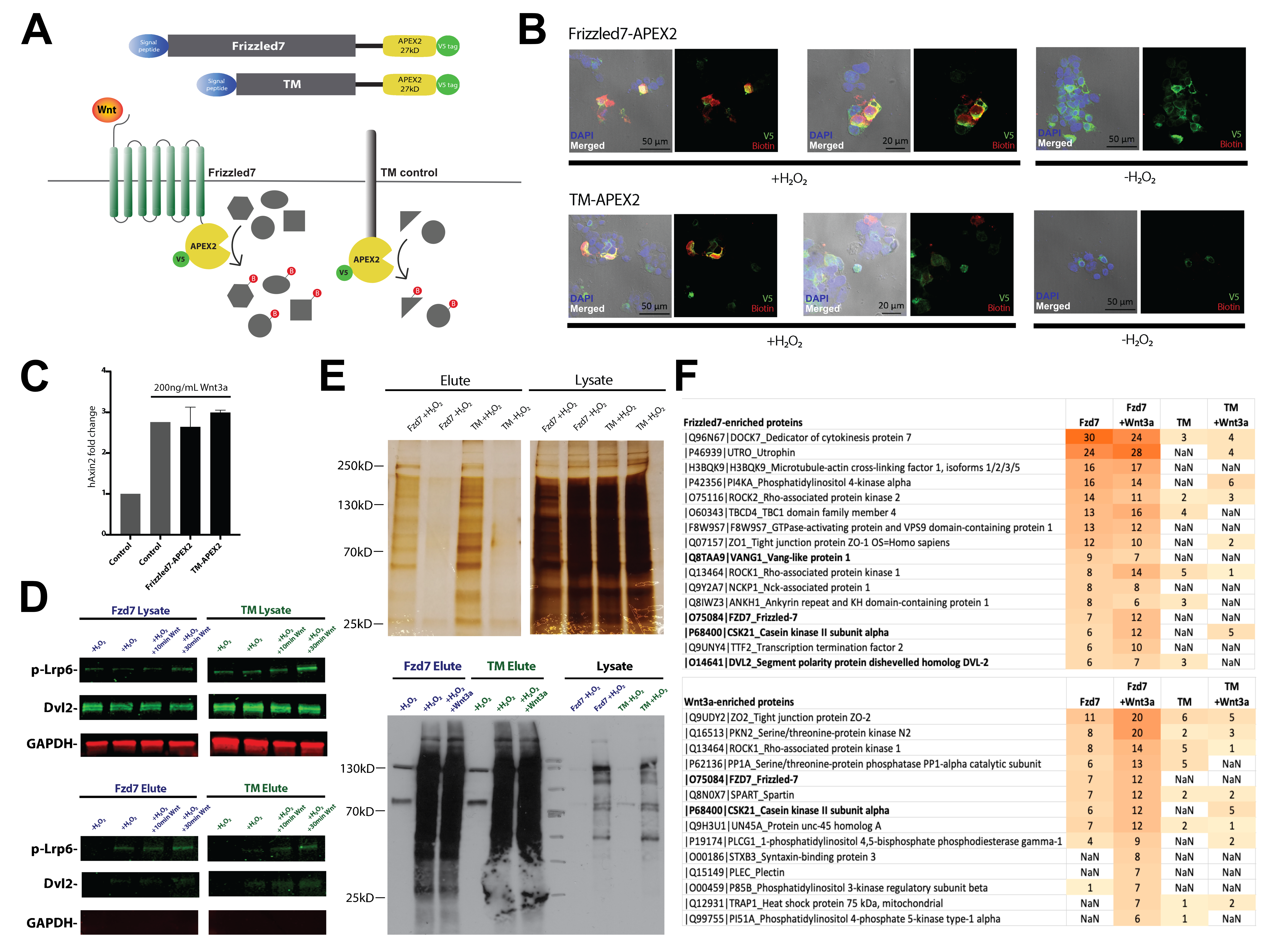
APEX2-Mediated Proximity Labeling of Wnt Receptor Interactors Upon Pathway Activation | microPublication
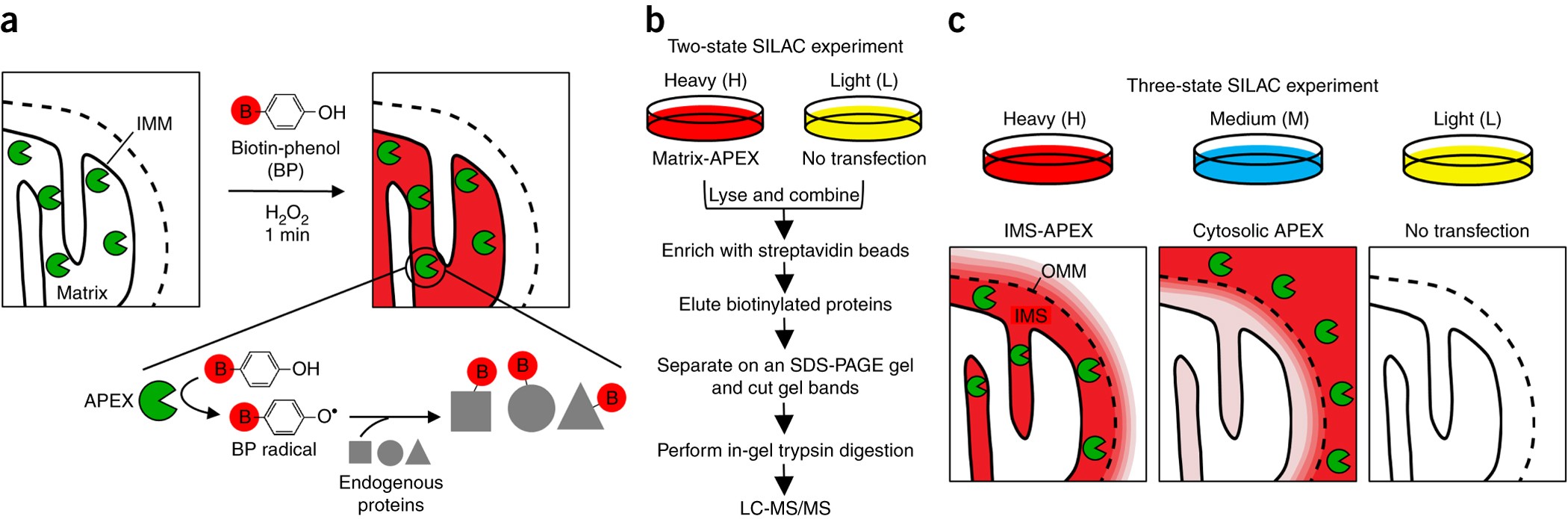
Spatially resolved proteomic mapping in living cells with the engineered peroxidase APEX2 | Nature Protocols