Quantitative Sequencing of 5-Methylcytosine and 5-Hydroxymethylcytosine at Single-Base Resolution | Science

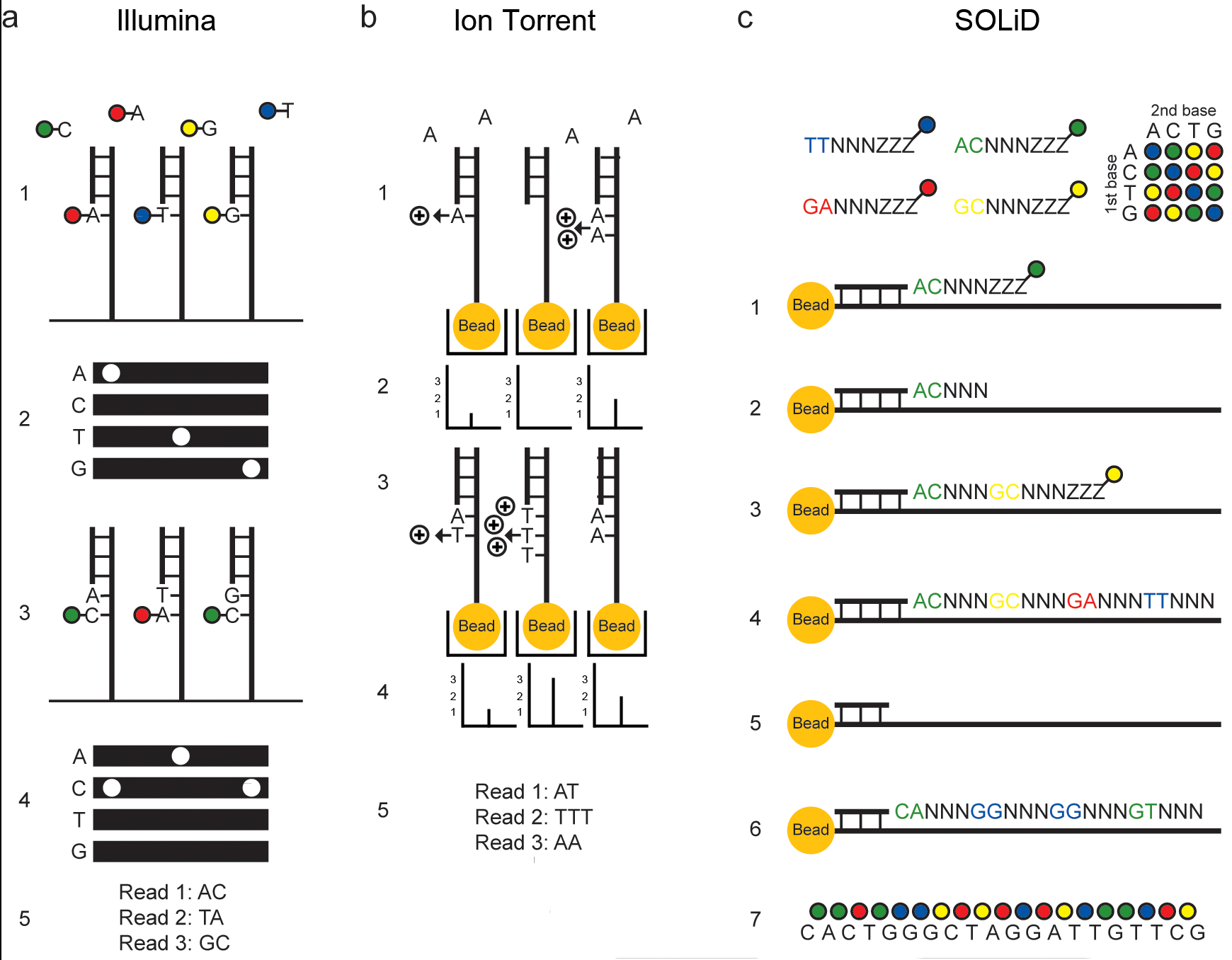
Figure 5. Applied Biosystems (Life Technologies) SOLiD sequencing by ligation. - Applications of Clinical Microbial Next-Generation Sequencing - NCBI Bookshelf
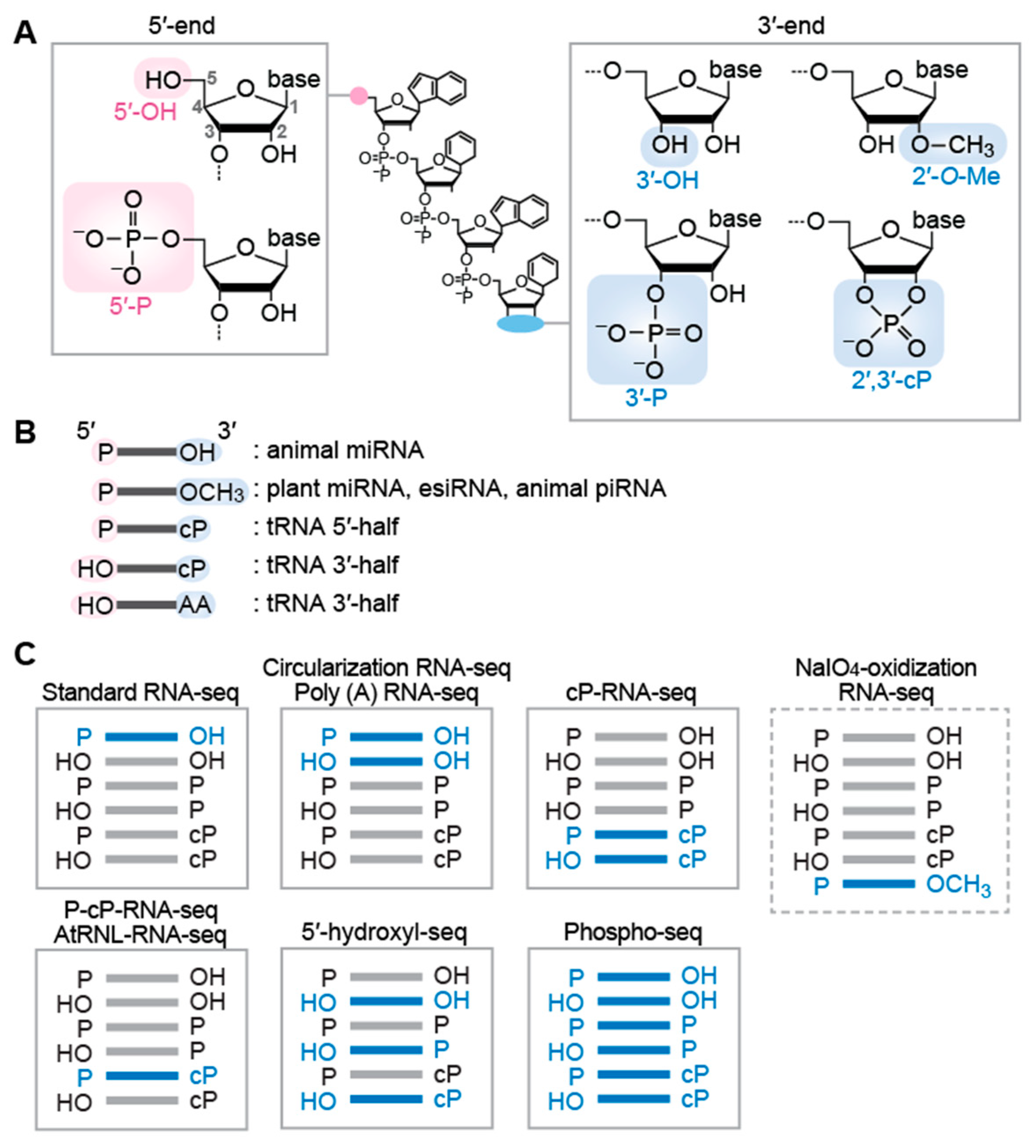
Biomolecules | Free Full-Text | Making Invisible RNA Visible: Discriminative Sequencing Methods for RNA Molecules with Specific Terminal Formations

5'XP sRNA-seq: A simple sequencing method to identify small RNA with and without 5' phosphorylation in a single library from low input samples | bioRxiv

High-throughput 5′P sequencing enables the study of degradation-associated ribosome stalls - ScienceDirect

Schematic overview of RACE-seq.: Standard 5′ and 3′ RACE primers are... | Download Scientific Diagram