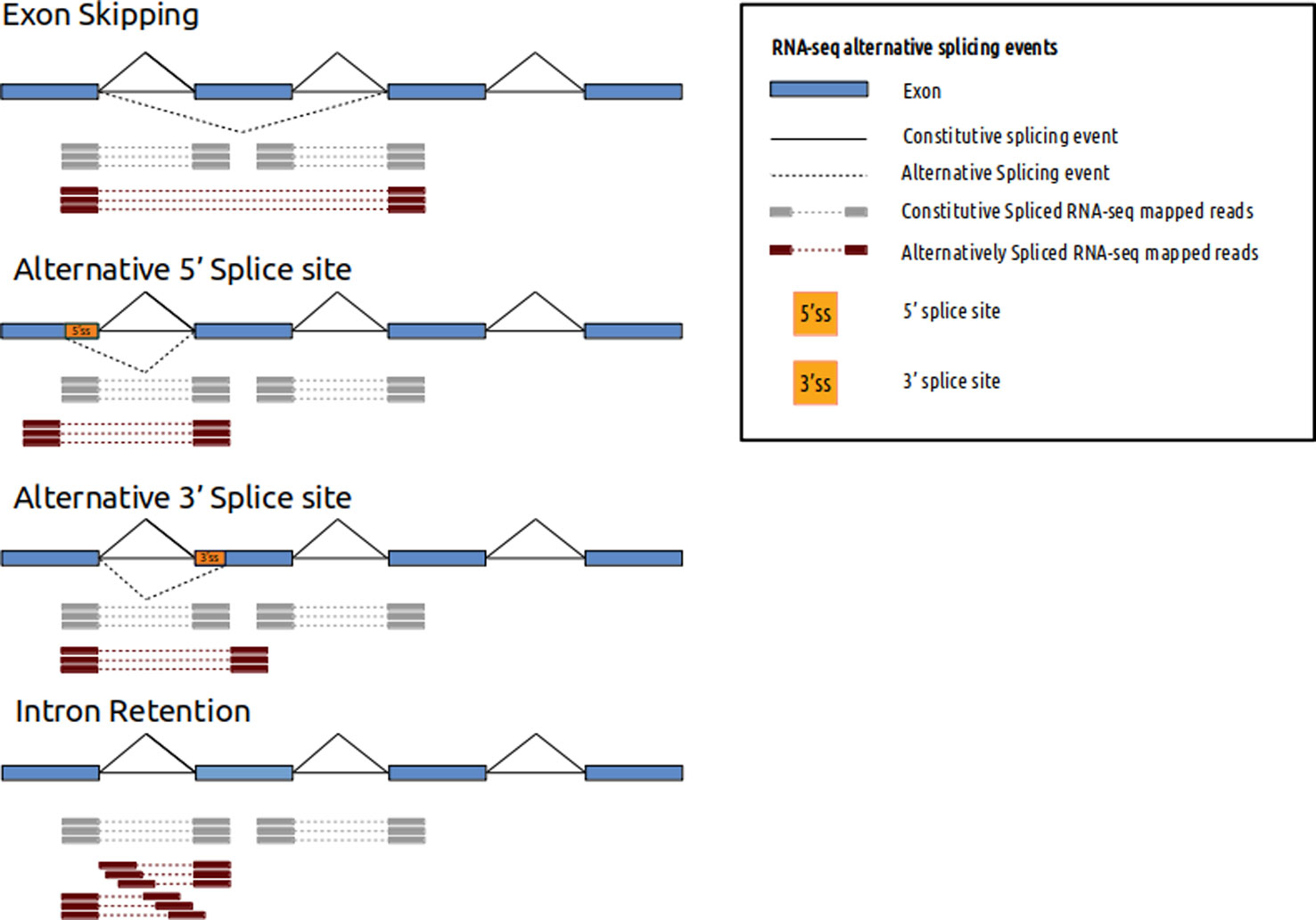
A work flow for 3'end-seq and PacBio Iso-seq. (A) Multiple versions of... | Download Scientific Diagram

The 'TranSeq' 3′‐end sequencing method for high‐throughput transcriptomics and gene space refinement in plant genomes - Tzfadia - 2018 - The Plant Journal - Wiley Online Library
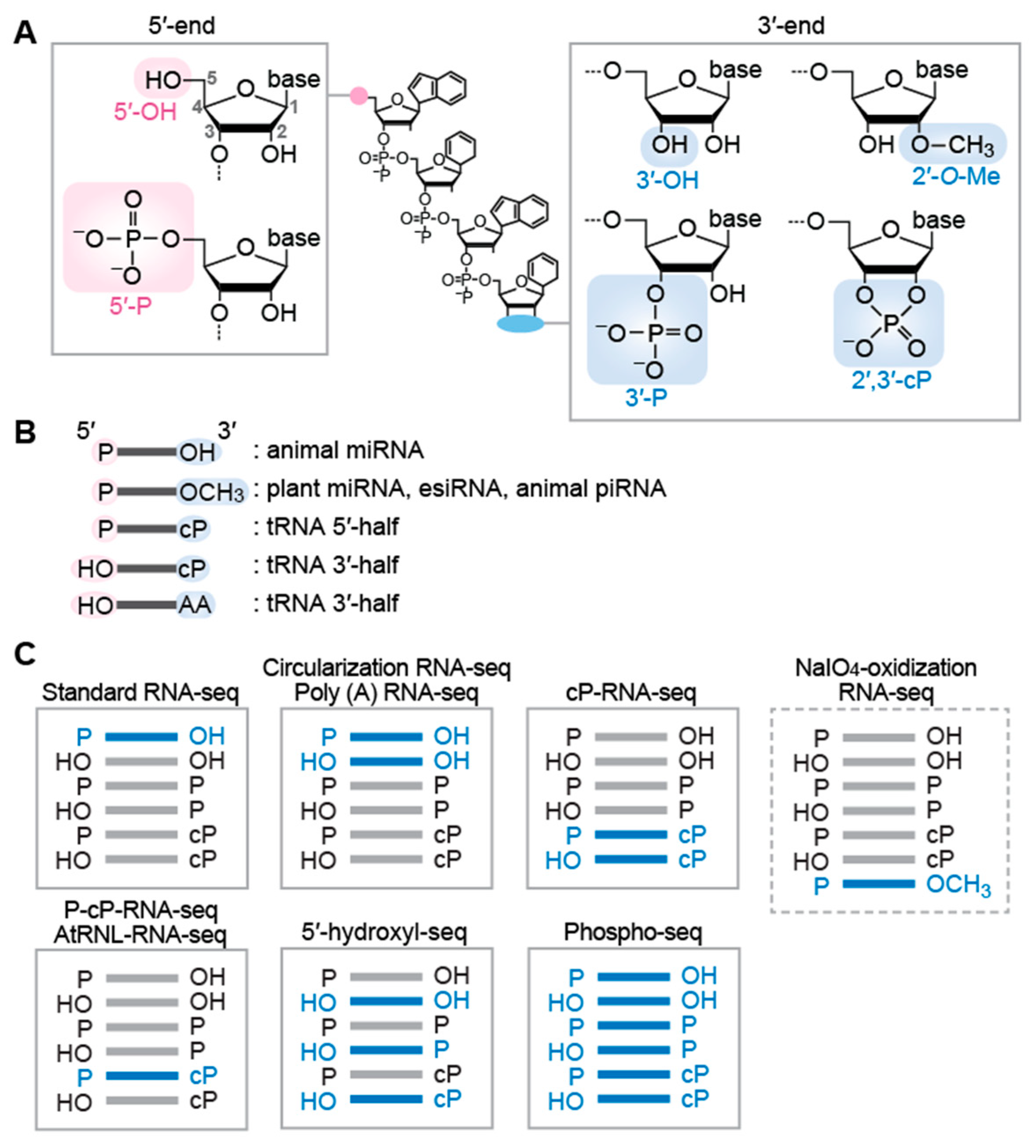
Biomolecules | Free Full-Text | Making Invisible RNA Visible: Discriminative Sequencing Methods for RNA Molecules with Specific Terminal Formations

General outline of oligo(dT)-based 3′-end sequencing protocols (e.g.,... | Download Scientific Diagram
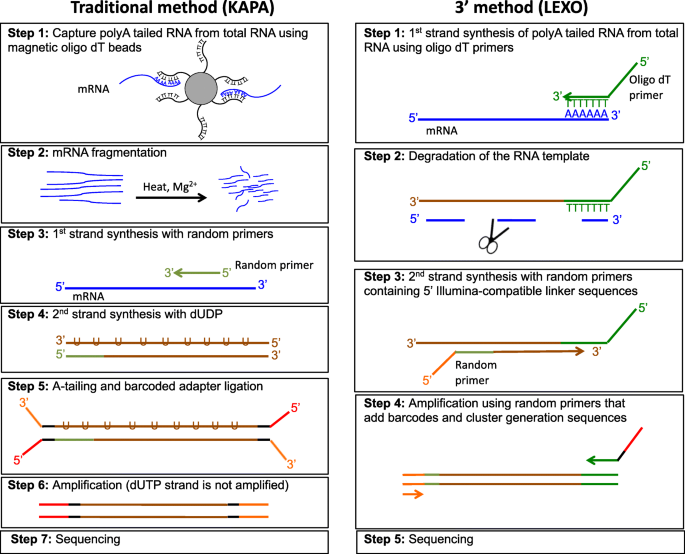
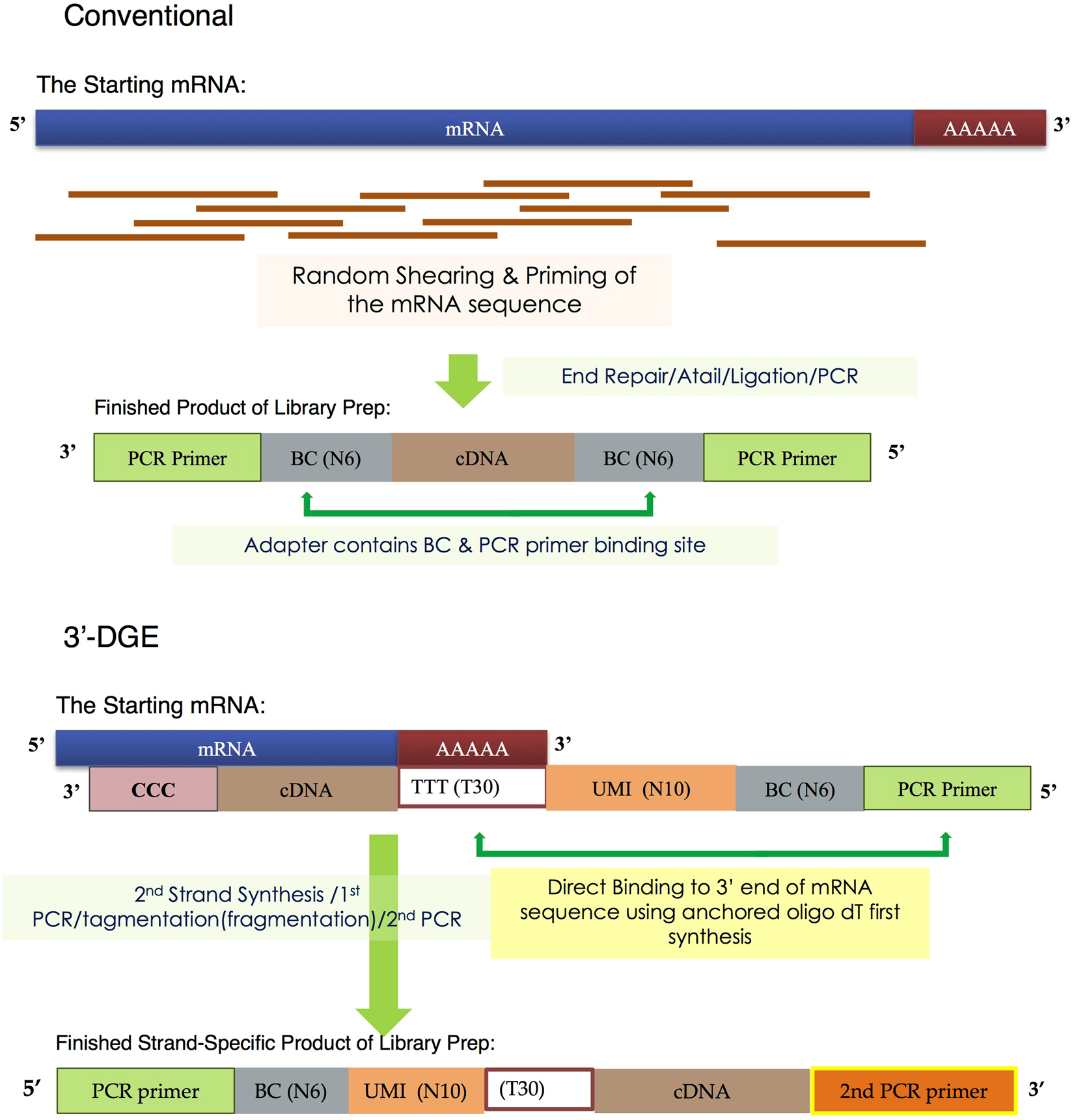
A comparison between whole transcript and 3' RNA sequencing methods using Kapa and Lexogen library preparation methods | BMC Genomics | Full Text

3′ end sequencing workflow with oligo(dT) 19 V. a) Overview of the 3′... | Download Scientific Diagram

Nano3P-seq: transcriptome-wide analysis of gene expression and tail dynamics using end-capture nanopore cDNA sequencing | Nature Methods

Multiplex structural variant detection by whole-genome mapping and nanopore sequencing | Scientific Reports

End-seq – map both 5' and 3' transcript ends genome-wide at single nucleotide resolution in 3 days | RNA-Seq Blog

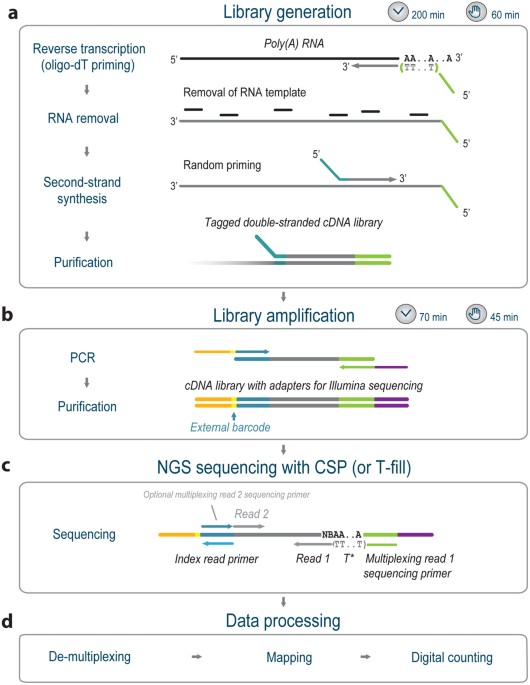
Figure 1 from A multiplex RNA-seq strategy to profile poly(A+) RNA: application to analysis of transcription response and 3' end formation. | Semantic Scholar





![3′ end sequencing methods. a In DeepSAGe [50] poly(A) + RNAs are... | Download Scientific Diagram 3′ end sequencing methods. a In DeepSAGe [50] poly(A) + RNAs are... | Download Scientific Diagram](https://www.researchgate.net/publication/262339133/figure/fig4/AS:667031334572034@1536044071441/end-sequencing-methods-a-In-DeepSAGe-50-polyA-RNAs-are-captured-by-oligo-dT.png)