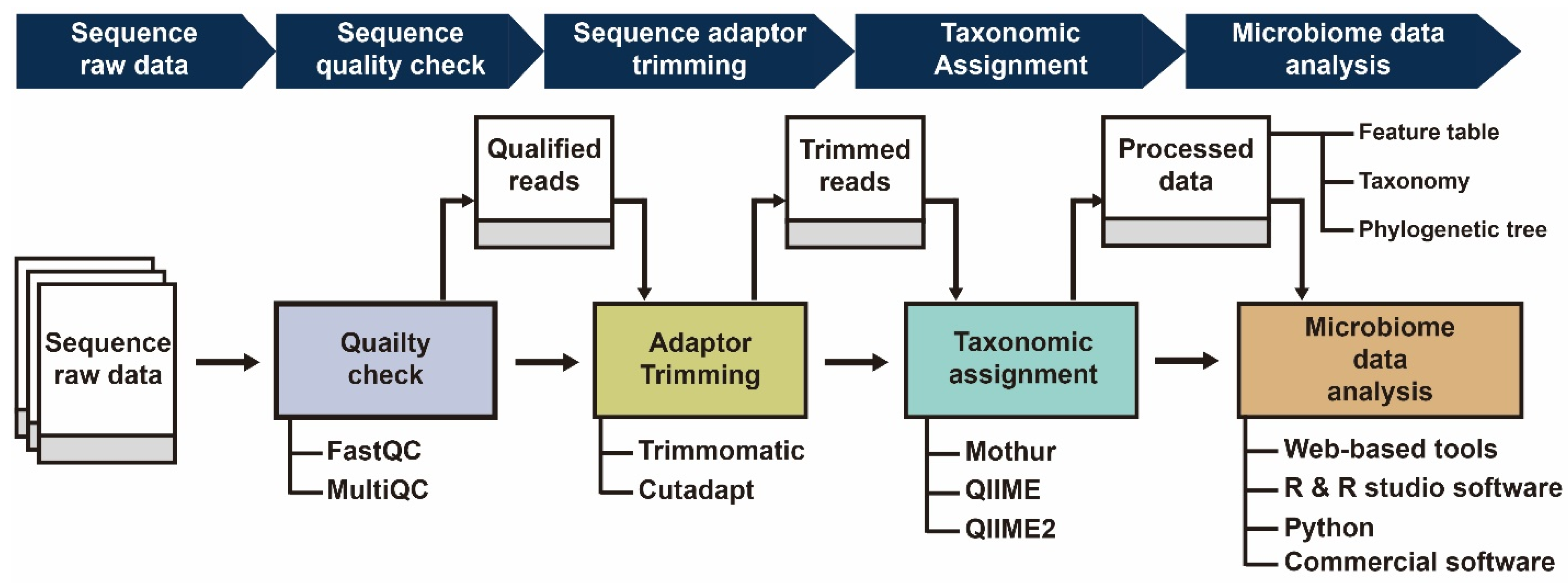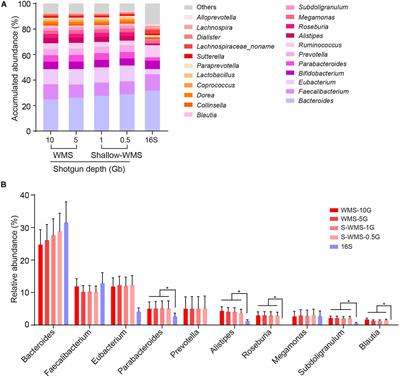
Frontiers | Characterization of Shallow Whole-Metagenome Shotgun Sequencing as a High-Accuracy and Low-Cost Method by Complicated Mock Microbiomes
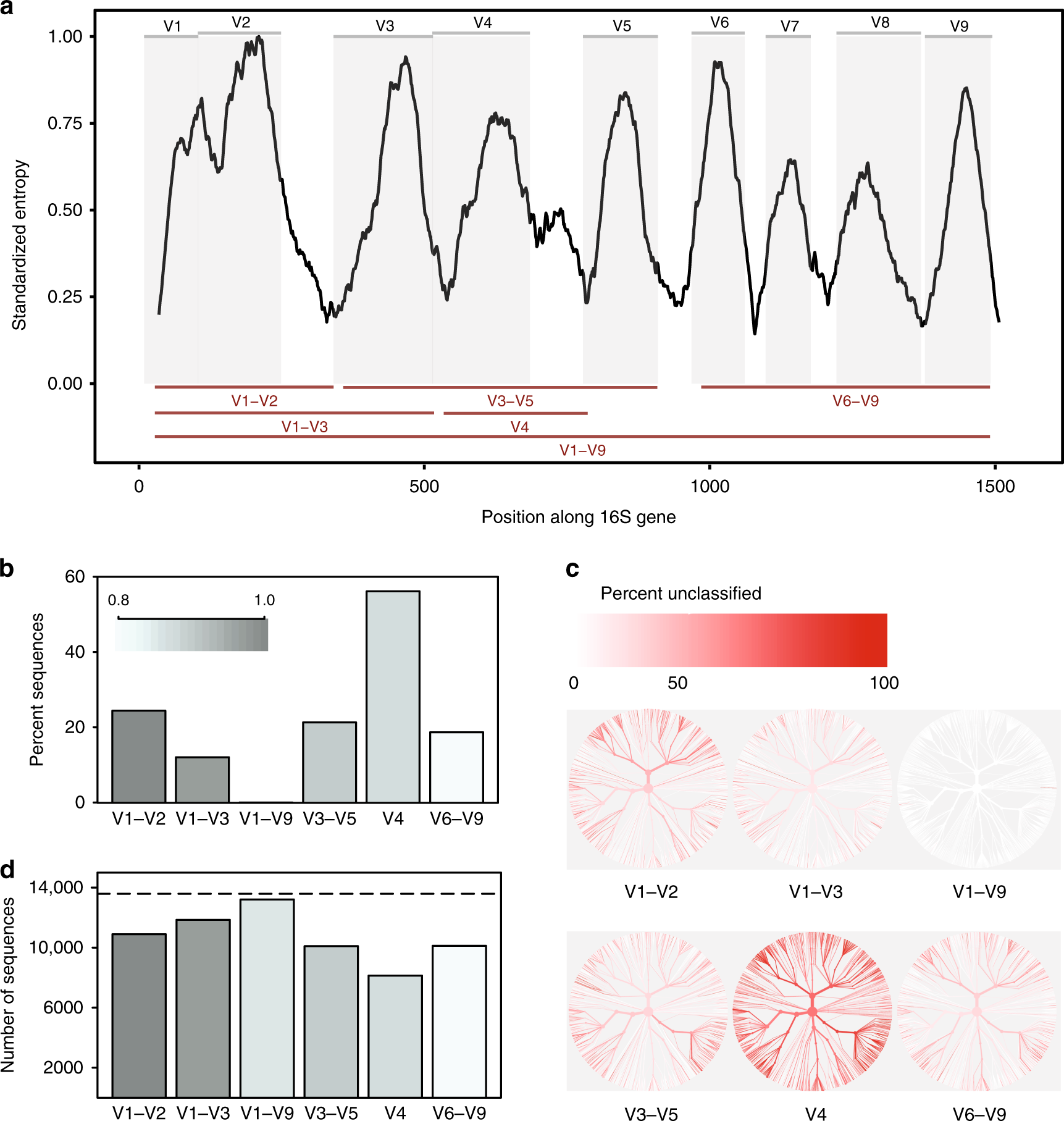
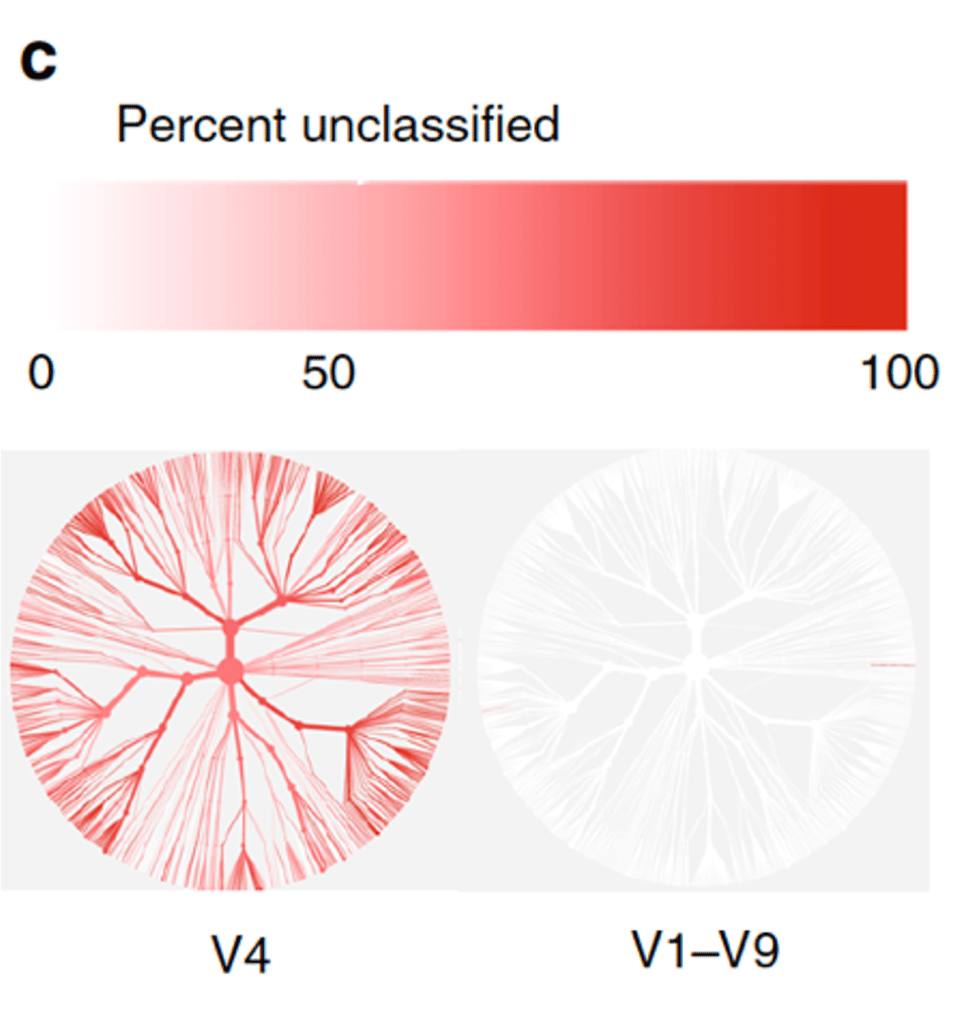
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis | Nature Communications
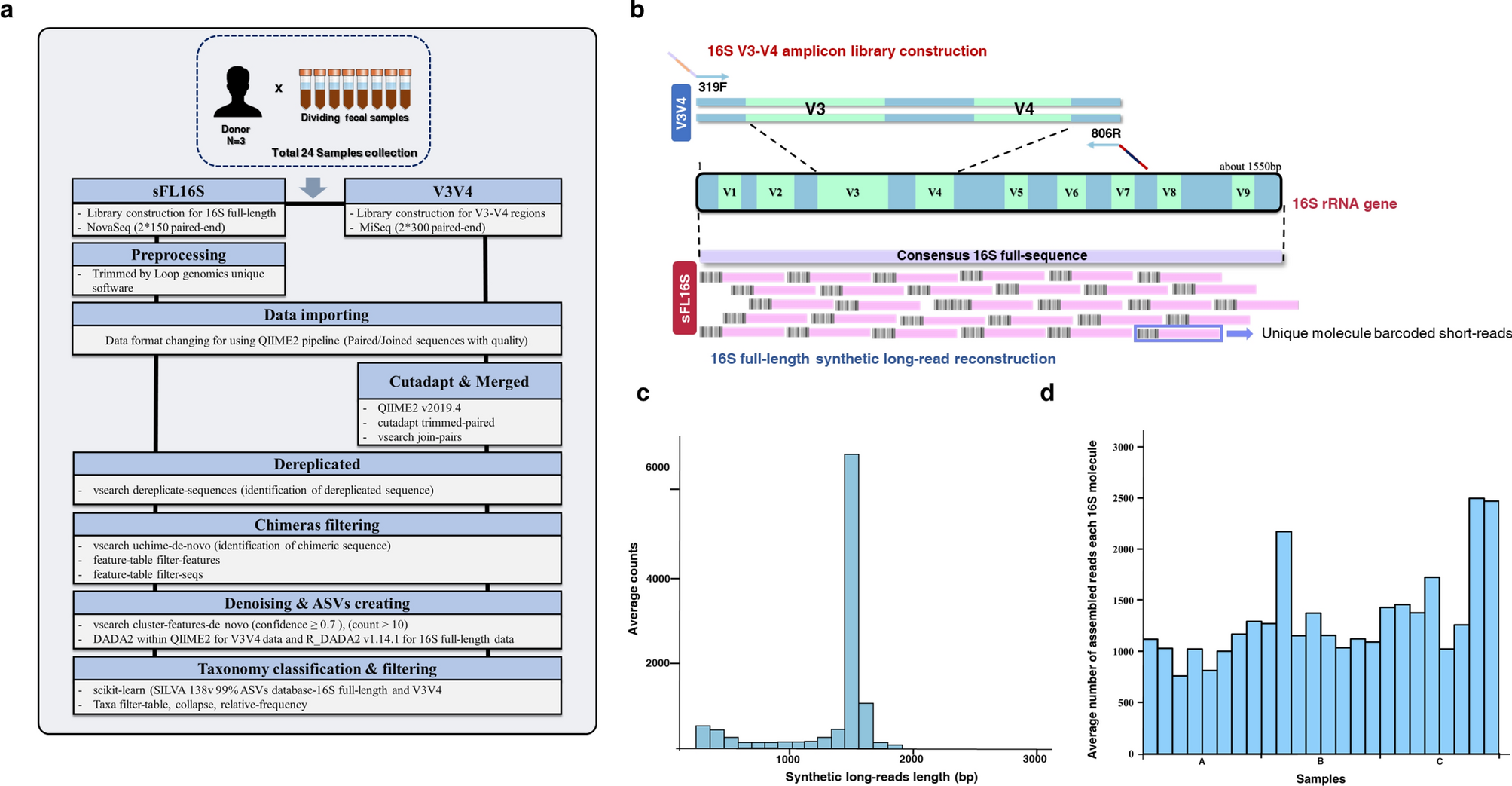
The effect of taxonomic classification by full-length 16S rRNA sequencing with a synthetic long-read technology | Scientific Reports

evSeq: Cost-Effective Amplicon Sequencing of Every Variant in a Protein Library | ACS Synthetic Biology
![Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ] Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/10778/1/fig-1-full.png)
Evaluation of full-length nanopore 16S sequencing for detection of pathogens in microbial keratitis [PeerJ]

Optimization of the 16S rRNA sequencing analysis pipeline for studying in vitro communities of gut commensals - ScienceDirect

Comparison between 16S rRNA and shotgun sequencing data for the taxonomic characterization of the gut microbiota | Scientific Reports

Reproducibility and repeatability of six high-throughput 16S rDNA sequencing protocols for microbiota profiling - ScienceDirect

Evaluation of metagenomic, 16S rRNA gene and ultra-plexed PCR-based sequencing approaches for profiling antimicrobial resistance gene and bacterial taxonomic composition of polymicrobial samples | bioRxiv