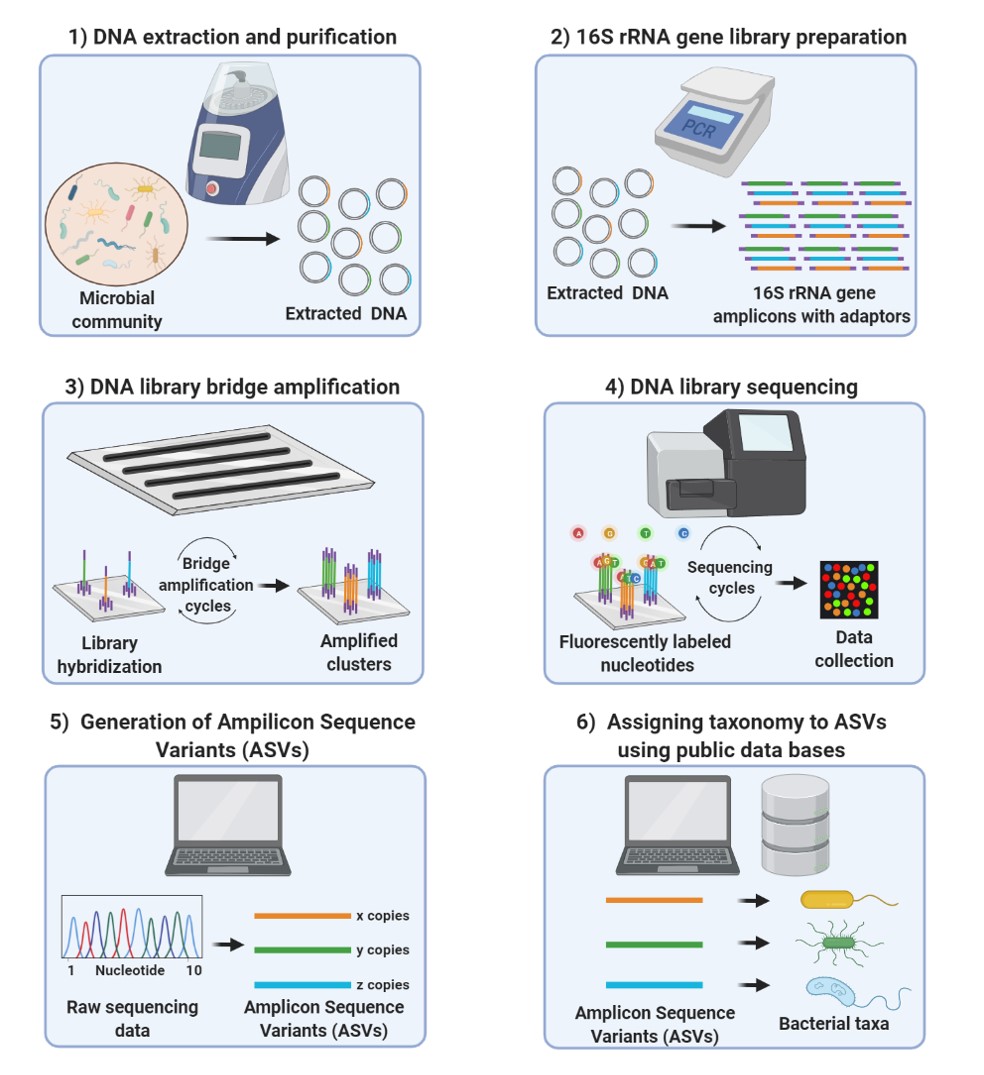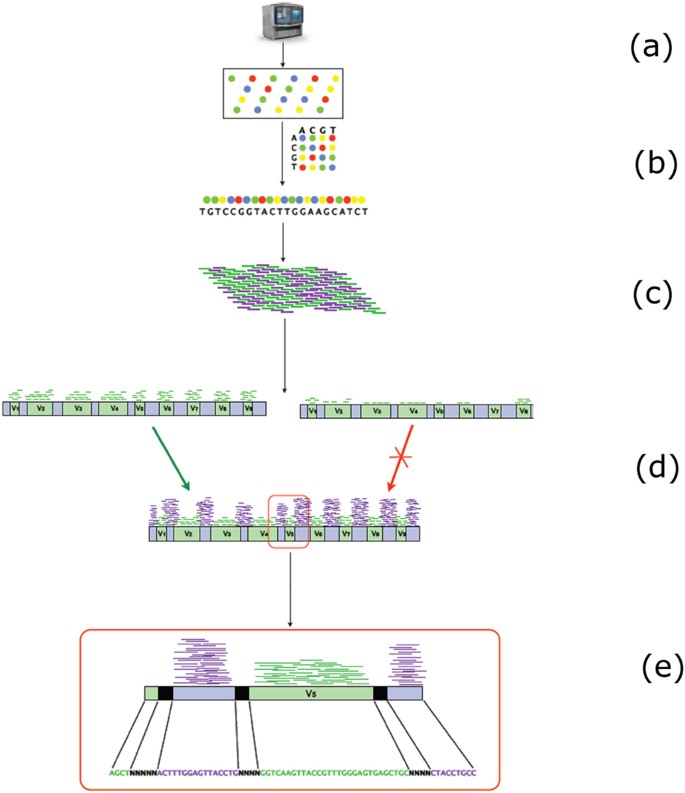
Direct 16S rRNA-seq from bacterial communities: a PCR-independent approach to simultaneously assess microbial diversity and functional activity potential of each taxon | Scientific Reports

Elisabeth Bik on Twitter: "16S rRNA amplicon sequencing for epidemiological surveys of bacteria in wildlife - M Galan https://t.co/A4GpyL1qdI https://t.co/wcqLpN3lrI" / Twitter
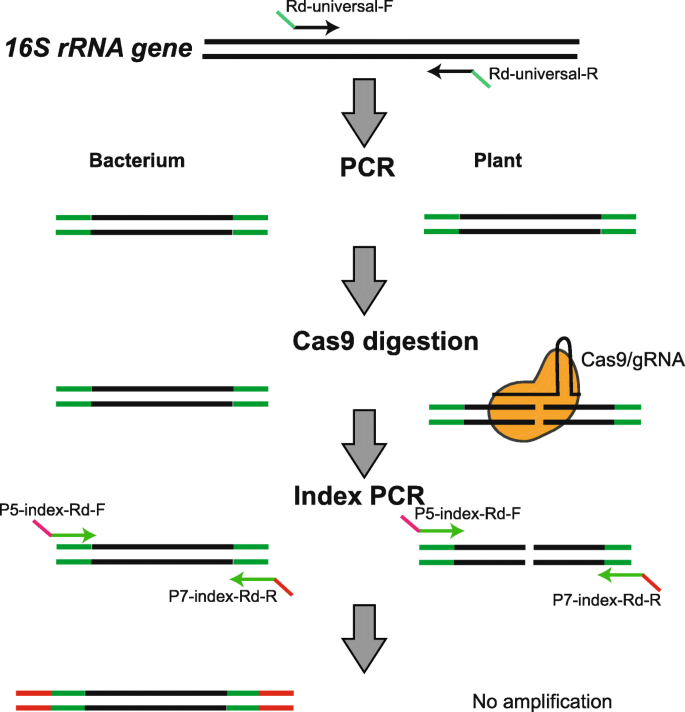
Engineering CRISPR/Cas9 to mitigate abundant host contamination for 16S rRNA gene-based amplicon sequencing | Microbiome | Full Text

Quantifying dominant bacterial genera detected in metagenomic data from fish eggs and larvae using genus‐specific primers - Najafpour - 2022 - MicrobiologyOpen - Wiley Online Library

Research Techniques Made Simple: Bacterial 16S Ribosomal RNA Gene Sequencing in Cutaneous Research - ScienceDirect
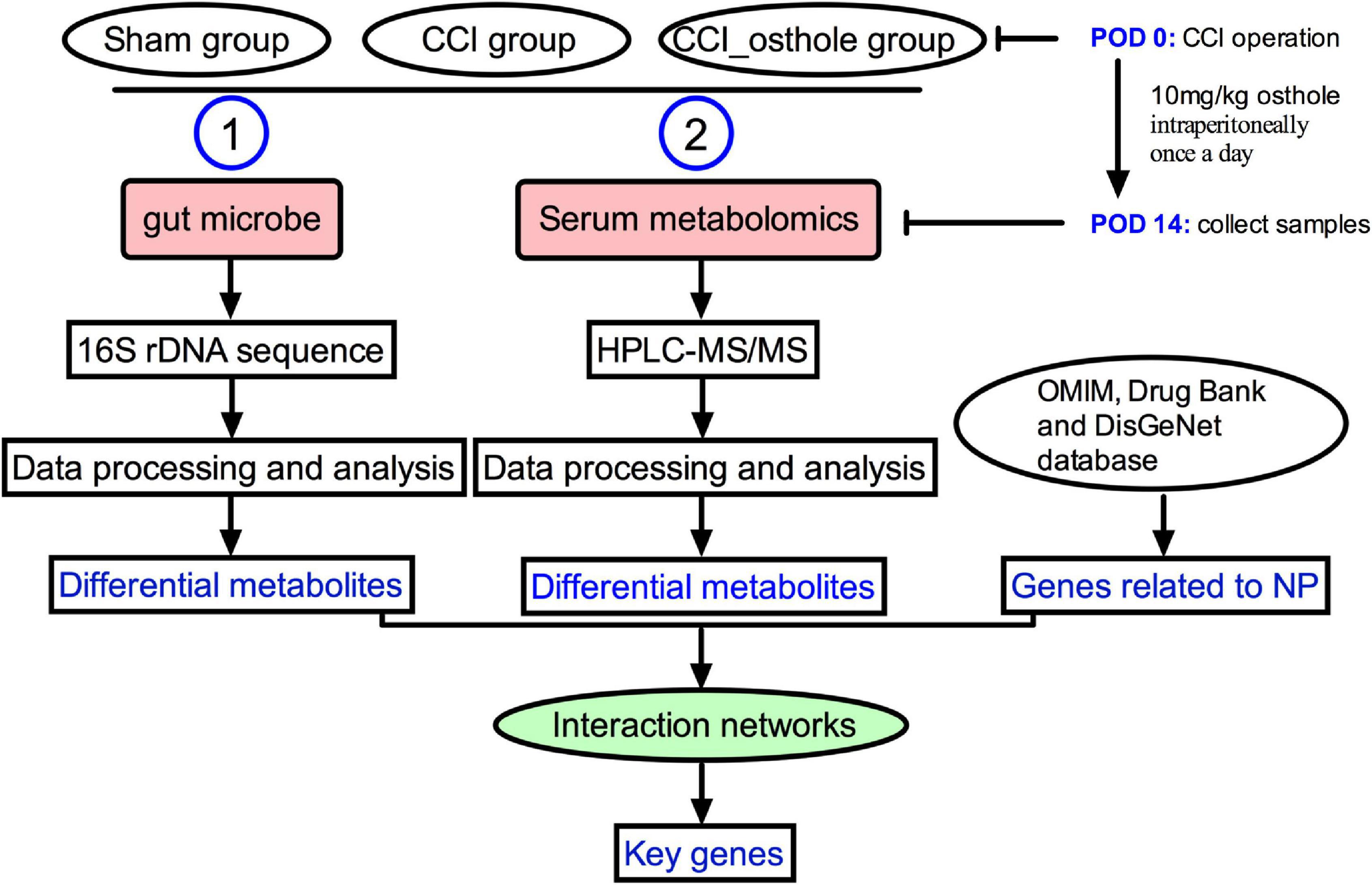
Frontiers | Integrated 16S rRNA Gene Sequencing and Metabolomics Analysis to Investigate the Important Role of Osthole on Gut Microbiota and Serum Metabolites in Neuropathic Pain Mice
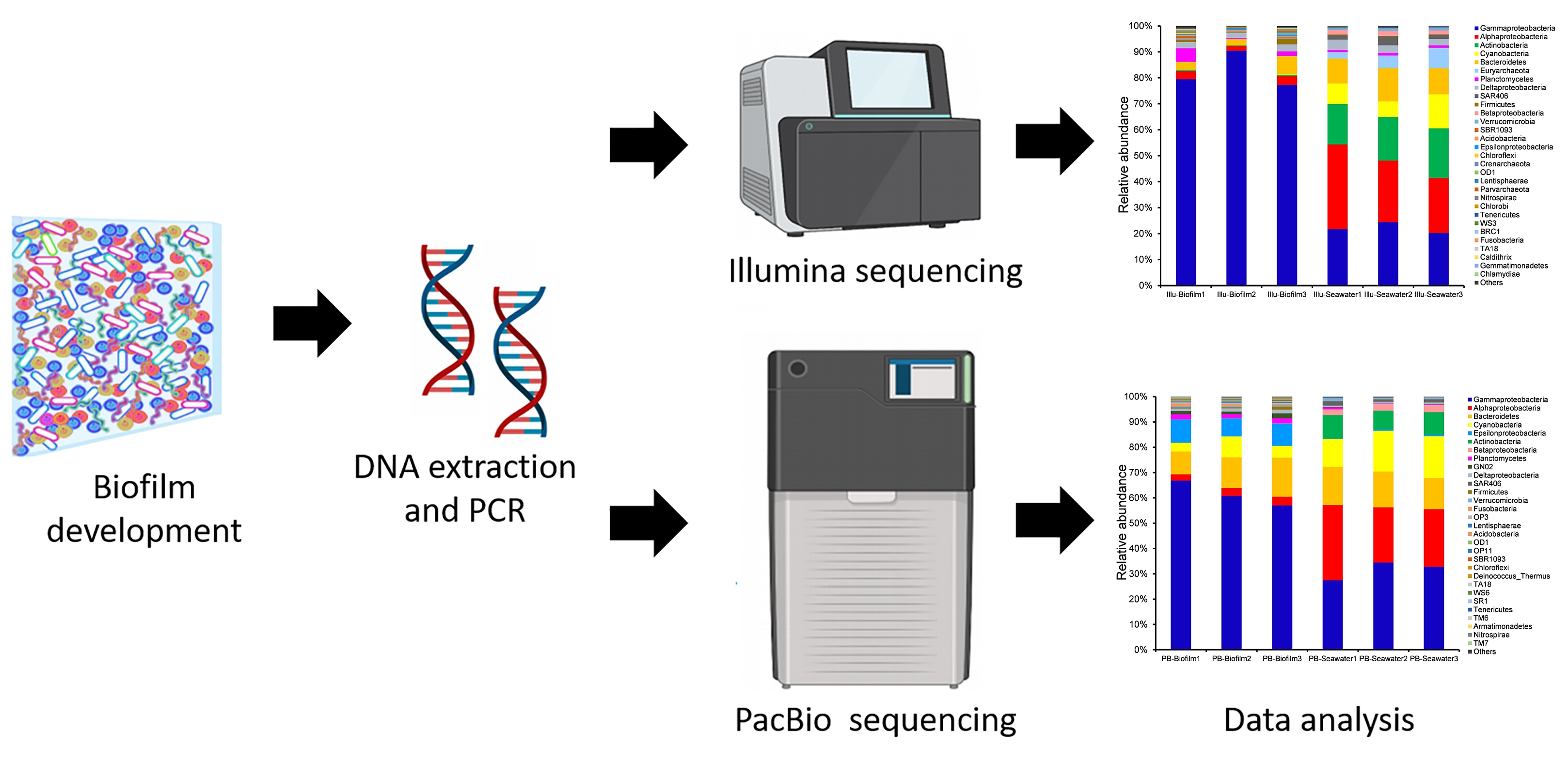
Genes | Free Full-Text | Microbial Richness of Marine Biofilms Revealed by Sequencing Full-Length 16S rRNA Genes
Demonstration of the workflow for the 16S rRNA and ITS amplicon sequencing. | Download Scientific Diagram
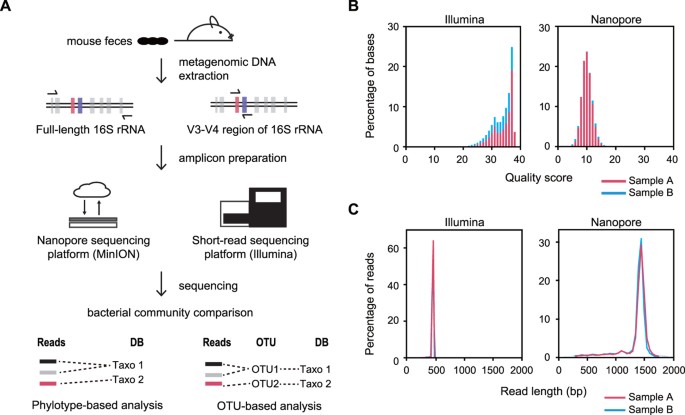
Analysis of the mouse gut microbiome using full-length 16S rRNA amplicon sequencing | Scientific Reports
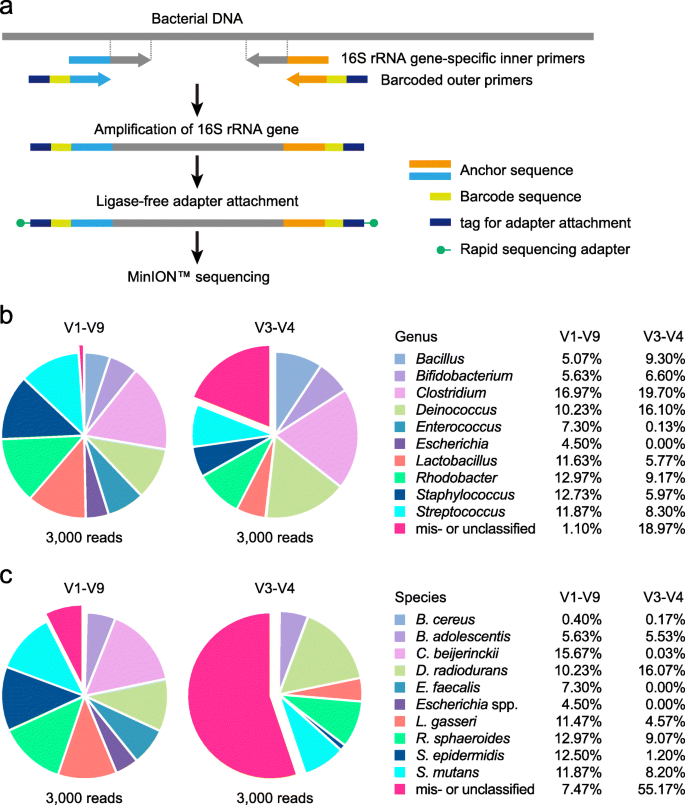
Full-length 16S rRNA gene amplicon analysis of human gut microbiota using MinION™ nanopore sequencing confers species-level resolution | BMC Microbiology | Full Text
Strengths and Limitations of 16S rRNA Gene Amplicon Sequencing in Revealing Temporal Microbial Community Dynamics | PLOS ONE
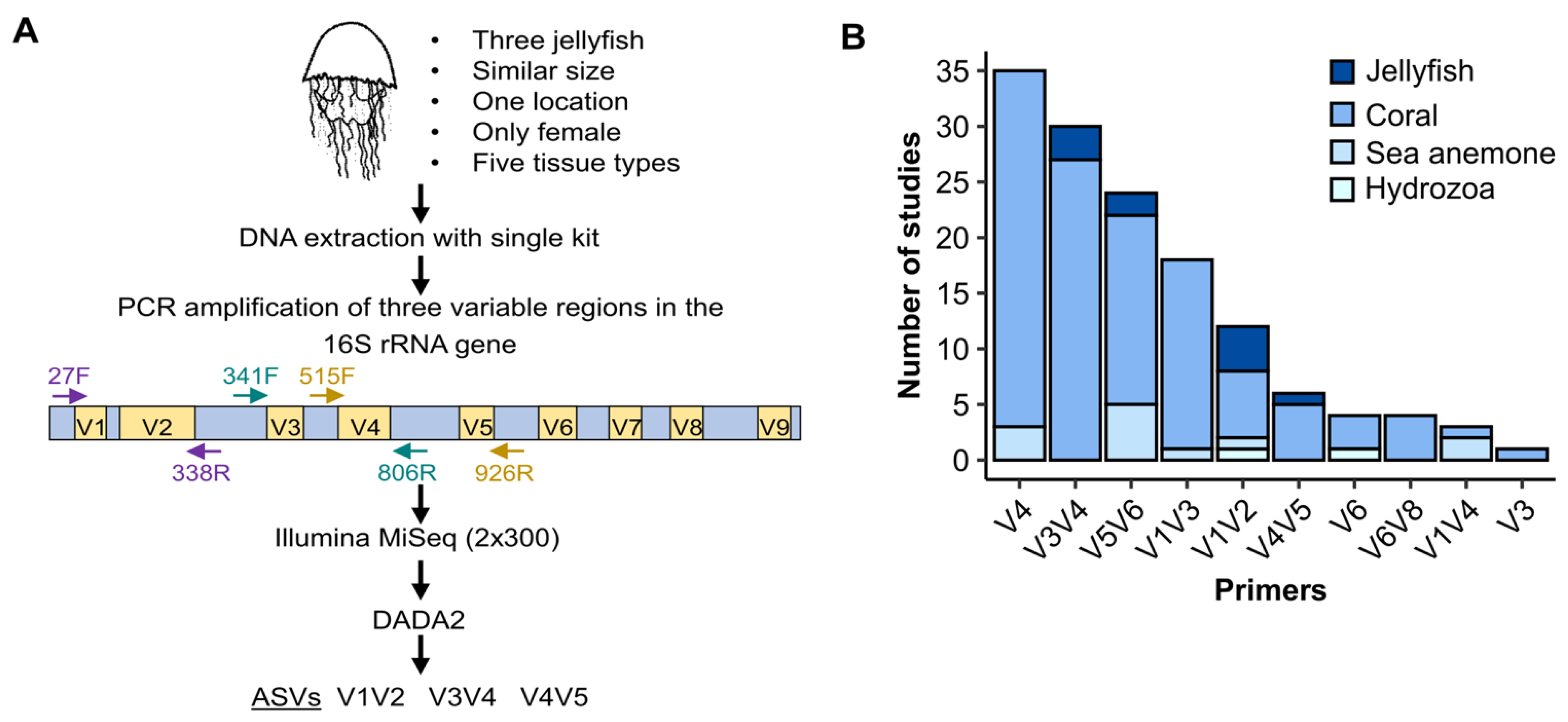




![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)